Inductively coupled plasma mass spectrometry (ICPMS) is one of the most commonly used analytical instruments for metal detection in various fields owing to its noteworthy features, including high sensitivity to trace level elements, good linear dynamic range, multiplex detection, and low detection limits.1 Furthermore, it can be used in conjunction with other equipment, such as highperformance liquid chromatography (HPLC),2 liquid chromatography (LC),3 gas chromatography (GC),4 size exclusion chromatography (SEC),5,6 and so on. The hyphenated techniques extends its application to various fields and helps to address the limitations of ICP-MS. Furthermore, some hyphenated methods have had a profound impact on clinical and biological analysis because of their high sensitivity and selectivity with multiplex information from analytical processes. Generally, ICP-MS can easily lose specific biological information when biomolecules are mixed with various matrices and certain metals, which resulted in low sensitivity, poor reliability, and high backgrounds due to the complicated matrix. For those reasons, separation techniques such as LC, HPLC, GC, and gel electrophoresis have been used to extract or separate certain target molecules from the matrix. Essentially, most researches using ICP-MS in bio analysis have focused on metallic functions or mechanisms through direct determination of metal. Recently, clinical fields have been strongly addressed because metallic pollutants from environment, stress, changes in diet, and other reasons that affect human life and lead to an increased chance of disease are being issued. For instance, a human has almost 3.72 × 1013 cells,7 and each of them contains numerous biomolecules. Some cells participate in chemical reactions in body, which are important for living things to survive; therefore, these reactions are strictly controlled by complicated or crosslinked checking systems.7 However, although reactions are checked or revised with various correction systems, the unsuspected involvement of metallic ions sometimes creates abnormal or malfunctioning cells in a body, and the metal controls are not always complete.7 Therefore, sensitive and selective detection is almost a necessity for the investigation of those reactions.
For clinical purpose, many researchers try to discover and treat cancer cells at an early stage, before they progress to a late stage or metastasizes to other organs.8 For this, various immunoassay of electrochemistry, chemiluminescence, and ELISA have been studied.9,10,11,12,13 However, some of those techniques have limitations in sensitivity and selectivity to be applied for diagnosis. In case of ICP-MS, metal tagging must be required if no metallic element were in the target molecule. In this article, various ICP-MS techniques for clinical and bio analysis will be discussed.
Analytical Techniques using ICP-MS
Current analytical techniques using ICP-MS for bio and clinical analysis can be classified into two categories, direct and indirect analysis. As explained, the biggest difference between them is the use of tagging materials. For example, direct analysis identifies analytes directly by detecting metallic elements involved in the target molecules. On the other hand, indirect analysis requires tagging materials for quantification, so that the targets are determined from the tagged elements.
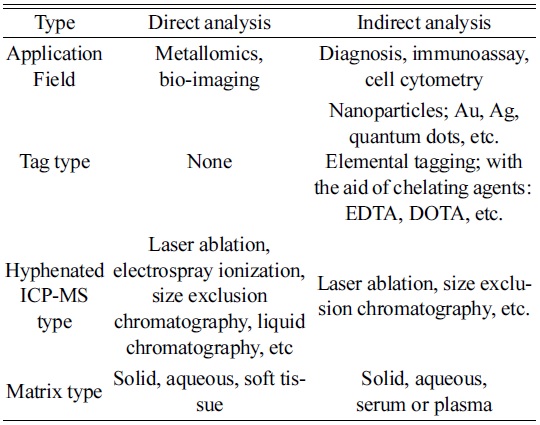
As listed in Table 1, one of the typical direct methods has been known as metallomics, proposed by Hiroki Haraguchi in a study of metal-containing biomolecules or metalloproteins of organisms.14 Even though metals are not the main elements in a body, they play important roles in the functioning of biological systems. For example, zinc acts as a “zinc finger” that is crucial for gene regulation in the cell.15,16 It also participates in various actions in a body, such as immune function, cell growth, hormone activity, and protein metabolism.17
[Table 1.] Classification of possible ICP-MS techniques for bio analysis.
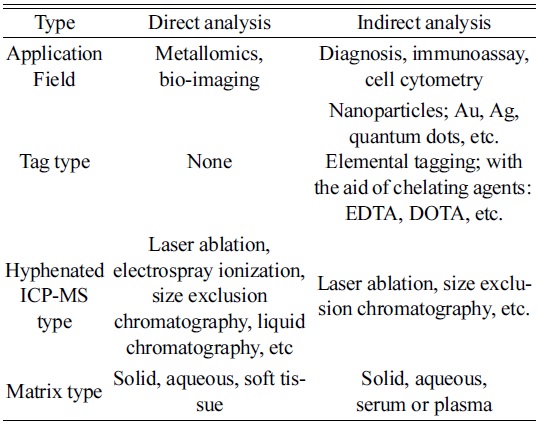
Classification of possible ICP-MS techniques for bio analysis.
The sample introduction technique for bio analysis has been remarkably advanced, i.e., from solution nebulization to laser ablation. Among them, laser ablation is a unique technique for bio samples and significantly contributes to the imaging of metal distribution in tissues. For example, Becker et al. reviewed bioimaging of metals using laser ablation inductively coupled plasma-mass spectrometry (LA-ICP-MS).18 Recently, LA-ICP-MS was combined with microarray chip sampling for the determination of holo-ceruloplasmin (holo-CP) in serums, of which each molecule contains six Cu ions.19 The use of polydimethylsiloxane (PDMS) microarray chips with 150 ìm-diameter pillars for sampling minimized the sample consumption to about 126 pL loaded on each pillar. The target CP was extracted from the surface of each pillar through immunoreaction for high selectivity.19 Several review articles on direct analysis have been published as well. For example, Parson and Barbosa discuss diagnostic methods using mass spectroscopy,20 and J. Sabine Becker et al. review imaging techniques, including a LA-ICPMS.21,22 Sanz-Medel et al. review protein analysis by elemental tagging.23 Specifically, metalloproteins in brain research were in the focus of targeted therapeutic approaches of neurodegenerative diseases, such as Alzheimer’s and Parkinson’s disease.24 The spatial resolution of bioimaging has also been studied by developing online laser microdissection.25
Indirect analysis has been employed when certain target molecules contain low levels of metals, even lower than quantification limit. In order to achieve the purpose, it requires more sample treatment steps than direct analysis, indicating long analysis times and toilsome efforts. However, it has the advantages of multiplex detection and high sensitivity owing to various tagging elements with no interference in ICP-MS. As a tagging moiety carrying metal ions, various ligands, such as ethylenediaminetetraacetic acid (EDTA), 1,4,7,10- tetraazacyclododecane- 1,4,7,10- tetraacetic acid (DOTA), diethylene-triaminetetraacetic acid (DTTA), and diethylene triamine pentaacetic acid (DTTA), have been proposed although no signal amplification was expected in this case. Recently, attention has been paid to nanoparticles because each nanoparticle contains many metallic atoms, depending on size and density.26,27,28 In addition, high selectivity can be achieved for indirect analysis through immunoreactions, i.e., the target selectively reacts with antibodies immobilized on probe nanoparticles. In the next section, indirect analysis using ICP-MS will be discussed in the field of clinical diagnosis.
Indirect ICP-MS techniques for clinical analysis
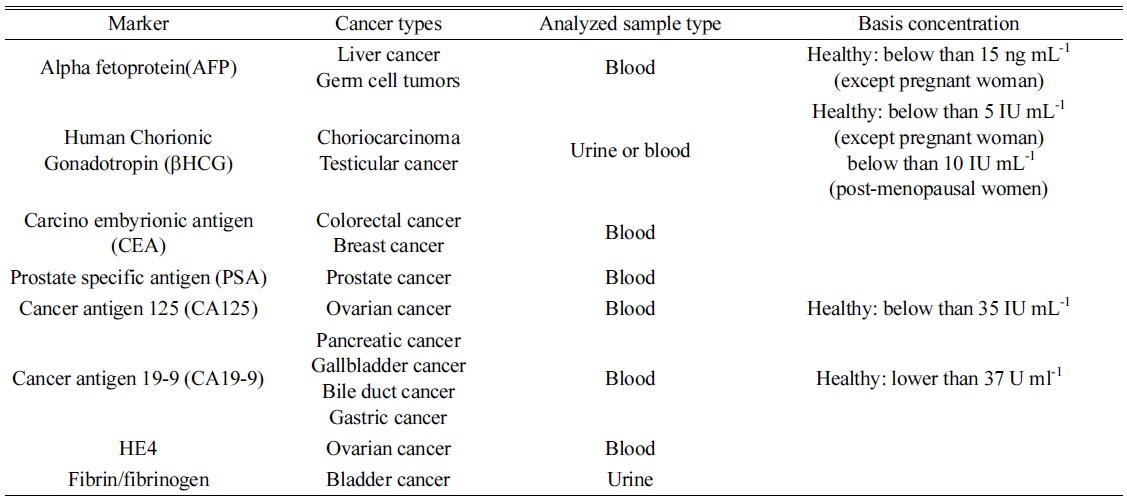
As described above, indirect analysis utilized probe materials to detect a certain biomolecule since doctors desire a diagnostic method with high sensitivity and accuracy with multiplex detection. In the past, they determined a disease by observing their own patients because diseases were not as complicated and diverse as they are now. Presently, various diseases and biomarkers have been discovered, and finding them is one of the barriers for complete recovery at an early stage. Multiplex detection is also needed for improving diagnostic accuracy. In order to satisfy such requirements, various probe nanomaterials have been synthesized for signal amplification, high selectivity, and multiplex detection in indirect analysis. For quantifications, nanoparticles have been modified and immobilized with antibodies to extract target and tag probes through immunoreactions. The diagnosis can be based on a relationship between a specific disease and biomolecule reported as its marker. Table 2 shows some tumor markers extensively discussed and frequently used for diagnosis, of which the data was compiled from the National Cancer Institute of USA and related articles.29,30,31
[Table 2.] List of tumor markers frequently applied for diagnosis.
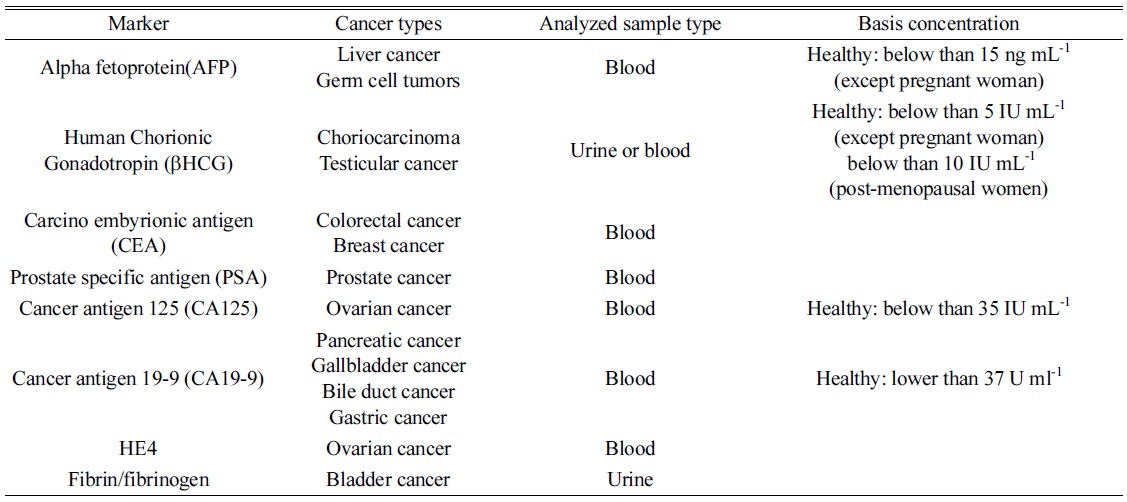
List of tumor markers frequently applied for diagnosis.
Many other researchers have detected tumor markers using elemental tagging. Although they used the same type of tags, various procedures were applied to specific antigens. For example, Sichun et al. and Shenghong Hu et al. explore immunoreactions on a solid plate and determine AFP and hCGß.32 In this method, antigens were captured by monoclonal antibodies immobilized on the microtiter plate. After the target extraction, Eu3+ and Sm3+ labeled antibodies reacted with AFP and hCGß, respectively, and the tagged elements were determined by ICP-MS for quantification.32
Similarly, Hu et al. used a similar solid microtiter plate to simplify sample pretreatment steps and a hyphenated ICP-MS with laser ablation for fast detection. Three antigens of AFP, CEA, and human IgG were prepared and captured by antibodies on a microarray plate, followed by a second immunoreaction with Sm3+-labeled AFP antibodies, Eu3+-labeled CEA antibodies, and Au-labeled goat-antihuman IgG (GAH). After the immunoreaction, the plate was ablated using a laser, and the generated particles were directly introduced into a plasma source using a carrier gas of He.33
Another hyphenated ICP-MS technique has been suggested by Terenghi et al. to improve sensitivity and to simplify a separation step.34 According to their strategy, five cancer biomarker proteins (r-fetoprotein (AFP), human chorionic gonadotropin (hCG), CEA, ovarian tumor antigen (CA125/MUC16), and gastrointestinal tumor antigen (CA19-9)) were determined using five different lanthanide tagged antibodies with Pr3+, Eu3+, Gd3+, Ho3+, and Tb3+. After tagging the elements, unbounded free metal ions and antibodies were selectively separated by SEC, and the results were compared to ELISA. The demonstrated method showed a similar limit of detection (LOD) with ELISA, and the amount of sample consumption was three times smaller than that of ELISA.34 As noticed, elemental tagging generally has shown reasonable analytical results with multiplex detection, but sensitivity was not satisfied for a specific purpose.
Recently, nanoparticle tagging has been extensively discussed because of several reasons: easy sample concentration and separation; the ability to improve sensitivity and accuracy through signal amplification; and the ability to provide a narrow surface area for tagging antigens, antibodies, or other biomolecules. Generally, Au nanoparticles (NPs) and quantum dots (QDs) are excellent candidates because of their high signal-to-noise ratio and low interference in ICP-MS. Notably, Au NP is the most discussed nanomaterial in analytical fields since it has not only good properties for spectroscopic analysis but also high sensitivity for mass spectrometry. For example, Liu and Lu use Au NPs to determine IgG and recombinant human erythropoietin, respectively, on the solid plate.35 The authors also found a detectable particle size of around 15 nm (0.03 fg gold colloids) from their experimental conditions. Furthermore, they compared the single particle mode with the conventional mode for human IgG. The method produced a relatively good linear correlation coefficient (R2) of 0.9816 and the quantitative result well agreed with that of ELISA. Lu et al. also used Au NPs to detect recombinant human erythropoietin (rhEPO), a hormone stimulating the proliferation of red cells.36 In the work, rhEPO was immobilized on the solid plate and distinguished from Au NP labeled anti-erythropoietin (EPO) antibodies.
Liu, et al. demonstrated a separation technique for sample treatment to detect albumin in aqueous conditions.37 They used ICP-MS hyphenated with capillary electrophoresis (CE) for matrix removal. From their strategy, human urinary protein, which was excreted with urine during renal processing of the blood, was separated and monitored for patients with diabetes and hypertension. Experimentally, they used Au NPs excessively, which produced Au NP-albumin adducts that presented more than 2000 atoms attached to each albumin molecule. Therefore, the LOD became extremely low: 0.5 pM when 280 nL of sample was consumed.
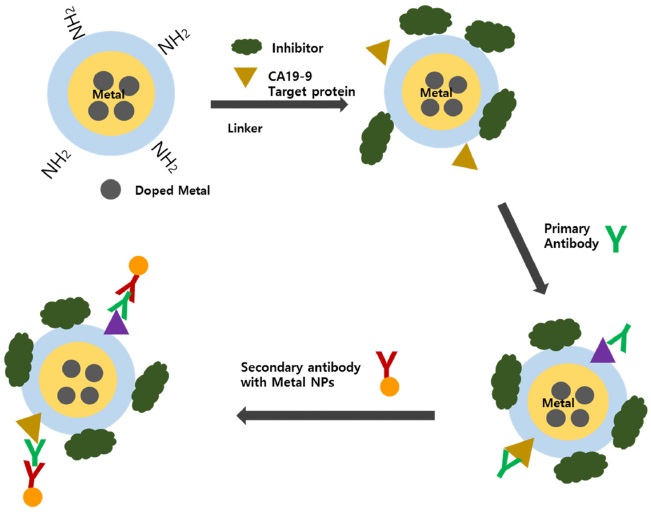
However, those hyphenated techniques showed limitations for separation and extraction of targets due to complexity and analysis time. In order to overcome the issue, Jang and Lim suggested a magnetic separation technique for sample pretreatment using magnetic nanoparticles.38 They modified magnetic nanoparticles and demonstrated them to determine various protein targets through magnetic separation. So far, many research has explored sandwich-type indirect analysis using the magnetic separation. For example, Peng et al used magnetic nanoparticles and mercury ions for target extraction and a probe, respectively.39 The proposed procedure was compared with chemiluminescent immunoassay (CLIA), commonly used for clinical purposes. The established method showed a lower LOD of 0.041 ng mL-1 than the other diagnostic methods, such as chemiluminescent multiplex immunoassay, multi-analyte electrochemical immunoassay, and surface plasmon resonance immunosensors.40,41,42 Noticeably, their work showed no signal amplification because mercury atom was used as a probe. Meanwhile, Ko and Lim synthesized metal-doped silica nanoparticles as a tagging material for multiplex diagnosis with high sensitivity and reliability.43 Silica nanoparticles have many suitable properties for bio application, such as easy surface modification and size control, good dispersion in hydrophilic solutions, and a large surface area. However, silica nanoparticles produce polyatomic interferences as a result of rearrangement or recombination in the plasma, which have the same mass as 28Si and 29Si (i.e., 14N2+, 12C16O+, 14N15N+, 14N21H+, 13C16O+, and 12C16O1H+). Although those interfering molecular ions can be removed through collisional induced dissociation or dynamic chemical reactions, the method can still suffer from poor sensitivity. If the metal-doped silica nanoparticles were used, high sensitivity as well as multiplex detection can be achieved because of signal amplification and variety of the doped metal ions. As an example, the authors designed three different metal-doped silica nanoparticles, Cs, Cd, and Pb. The prepared Csdoped core-shell SNPs were also used to monitor DNA cleavage induced by mercury or ROS.43 Furthermore, various kinds of metal-doped nanoparticles have been synthesized for sample treatment, including multicore magnetic nanoparticles (i.e., silica NP cores containing dye and metal ions and iron oxide magnetic cores).44 Since magnetic nanoparticles can provide benefits of easy target separation and concentration through magnetic extraction, a sandwich-type platform was preferred for sample preparation because of ratiometric measurement. The method was demonstrated for the determination of CA19- 9 in serums, as shown in Figure 1.
[Figure 1.] Schematic diagram of sandwich-type ICP-MS immunoassay for the determination of CA19-9.44

In this demonstration, the doped metal of the magnetic nanoparticles was used as an internal standard for ratiometric measurement in order to compensate sample loss during the sample treatment and measurement process. The obtained linear regression coefficient was significantly improved by revising the internal standard, and the LOD improved by 300 times, compared to the ICP-MS with element tagging and about 500-fold, compared to ELISA. Similarly, Zhao et al. used the sandwich-type platform for the detection of human r-thrombin that catalyzes many coagulation-related reactions for blood clotting.45 Noticeably, they used the affinity between aptamers instead of antibodies for target extraction. The LOD for human rthrombin was 0.5 fmol, and the calibration curve without ratiometric measurement showed a long dynamic range by three orders of magnitude.
Although direct analysis using ICP-MS has been used for a long time, indirect analysis using tagging has been extended its application from small biomolecules to cell analysis. In particular, laser ablation ICP-MS provides unique advantages of not only elemental distribution but also fast analysis with a small sample volume of less than a nanoliter. In addition, the development of various metaldoped inorganic nanoparticles will help to enhance the application potential of indirect analysis for bio and clinical analysis.