Baculoviruses are one of the largest and most diverse groups of viruses, and are phatogenic only for insects mostly of order Lepidoptera, Hymenoptera, and Coleoptera. The nuclear polyhedrosis virus isolated from Autographa californica (AcNPV), has a relatively broad host range(Ikonomou
The baculoviruses used in the BEVS(baculovirus expression vector system) are AcNPV and BmNPV(Motohashi
Currently, Sf9, Sf21, and Tn368 cell lines are used, since these make the reproduction of AcNPV considerably easier. The AcNPVSf9 host system is widely used as a representative cassette in BEVS (Kondo
On the other hand, the use of
AcNPV hosted by
In this study, the differences in AcNPV proliferation between different strains of
>
B. mori strains and virus injection
Seventy different strains from among the
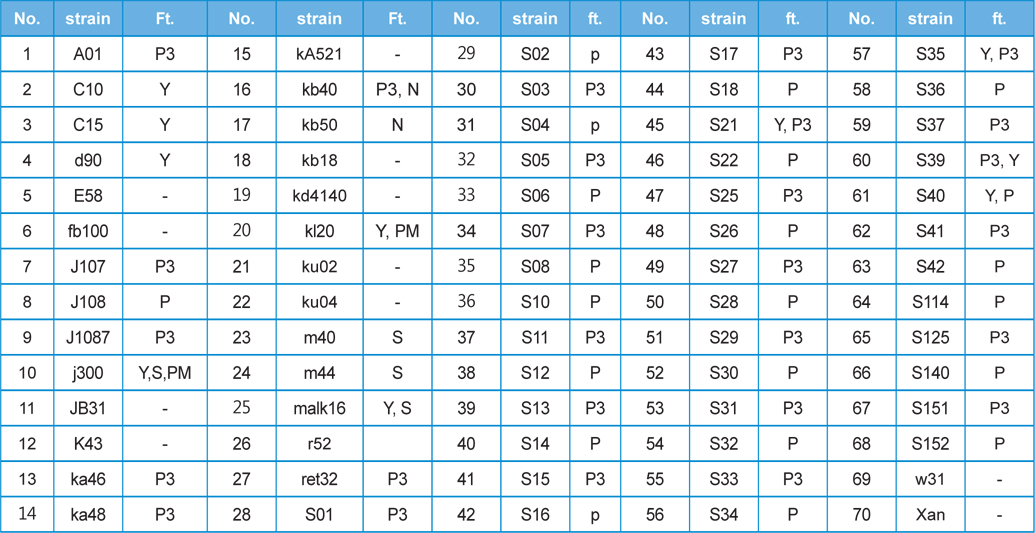
[Table 1] This study used 70 kinds of silkworm strain and features(ft.)
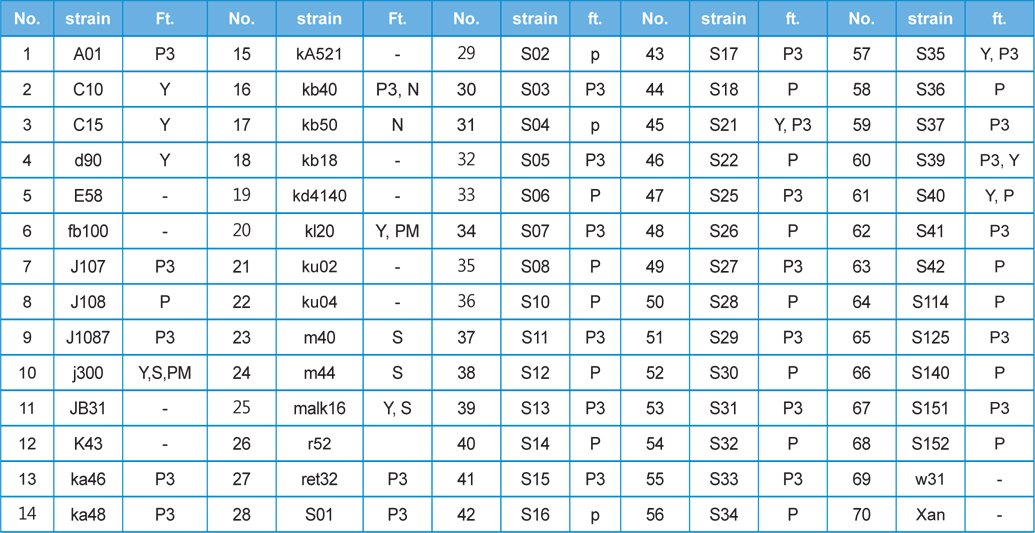
This study used 70 kinds of silkworm strain and features(ft.)
>
Recombinant AcNPV construction
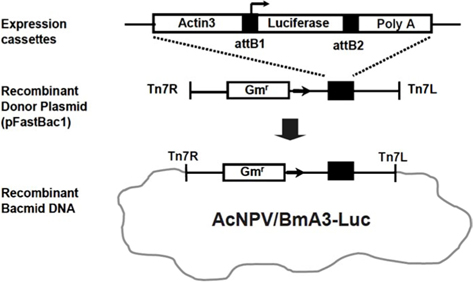
Recombinant AcNPV was produced using the luciferase gene as a reporter, using the Bac-to-Bac Baculovirus Expression System (Invitrogen, USA). In order to express the luciferase gene, an expression cassette containing the
>
Luciferase Activity measurement
Hemocytes were collected from 200 ¥ìL of hemolymph from virus infected individuals by centrifugation at 3000 rpm for five minutes. Seventy μL of cell solution (25 mM Tris-phosphate pH 7.8, 2 mM dithiothreitol, 2 mM 1,2-diaminocyclohexane- N,N,N’,N’-tetraacetic acid, 10% Glycerol and 1% Triton X-100) was added to the collected hemocytes and gently mixed for 30 minutes. The cell suspension was centrifuged at 10,000 rpm for 1 minute at 4°C and 20 μL of the supernatant and 50 μL of luciferase (Promega, USA) were mixed, after which the luciferase activity was measured (Lumistar Galaxy, BMG).
Recombinant AcNPV/BmA3-Luc was propagated using insect cells (Invitrogen, Carlsbad, CA, USA). Sf9 insect cells were cultured in Grace’s insect medium (Gibco-BRL, USA) with 10% fetal bovine serum (FBS, Invitrogen) at 27°C.
>
Reverse Transcriptase PCR (RT-PCR) and Real Time PCR
cDNA was synthesized from 1 μg of total RNA. Using a high capacity cDNA archive kit (Applied Bio systems, USA), PCR was performed on the obtained cDNA. A gene specific primer was made in order to partially amplify the luciferase gene inserted into the recombinant AcNPV (Fig. 1). One μL of cDNA was used of 30 amplification cycles, seconds at 94°C, 30 seconds at 48°C, and 1 minute at 72°C. Real-time PCR reaction mixture including the designed primer and SYBR green (TaKaRa) were used to apply the PCR. (Applied Biosystem 7300 Real-time PCR system). The amplification consisted of 10 seconds at 94°C, followed by 40 cycles of 95°C for 5 seconds, 62°C for 32 seconds and then a final cycle of 95°C for 5 seconds, 62°C for 32 seconds and 72°C for 30 seconds. 18S rRNA was used as a control group.
Dot blot analysis was performed by dotting 500 ng of extracted DNA from the hemocytes and fat cells onto Hybond N+ membranes (Amersham Pharmacia Biotech), and fixing with a UV-cross linker (Spectrolinker XL-100, USA). The luciferase gene obtained by PCR was labeled with radioisotopes (32P-dCTP) (Perkin Elmer) and used as a probe. The membranes were pre-hydrated in a hybridization solution without the probe for two hours at 68°C, and then hydrated with the probe in an hybridization solution overnight at 68°C. The film was then washed in 2X SSC, 0.1X SSC and exposed onto a film at −70°C.
>
Reproduction of AcNPV in B. mori
>
Screening of B. mori luciferase activity
The differences in the susceptibility of the 70 strains of
[Table 2] The results of silkworm host range against the propagation of recombinant AcNPV.

The results of silkworm host range against the propagation of recombinant AcNPV.
>
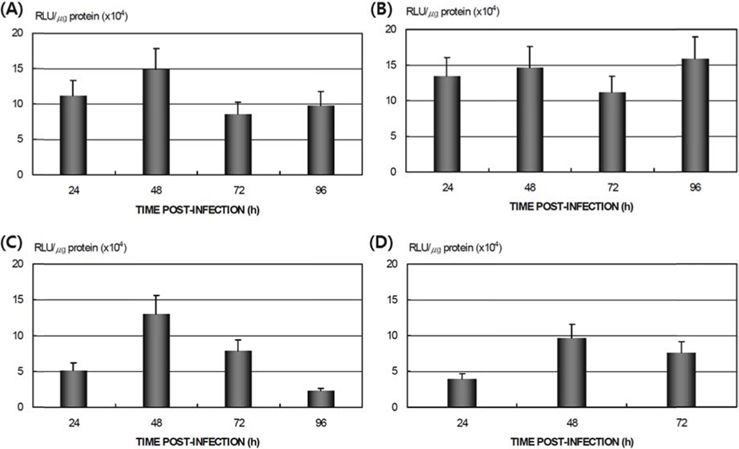
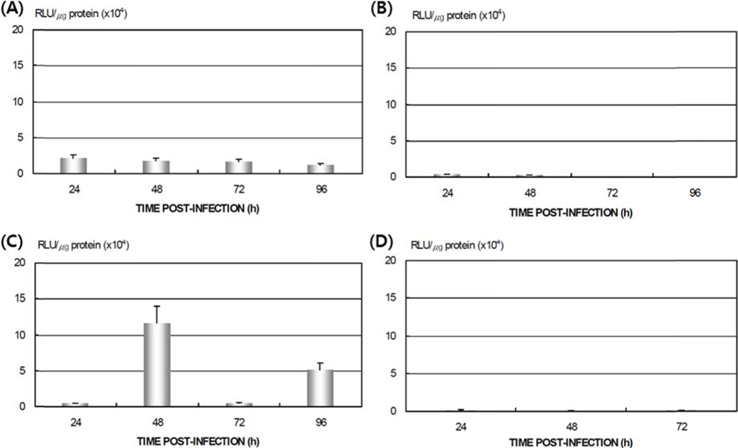
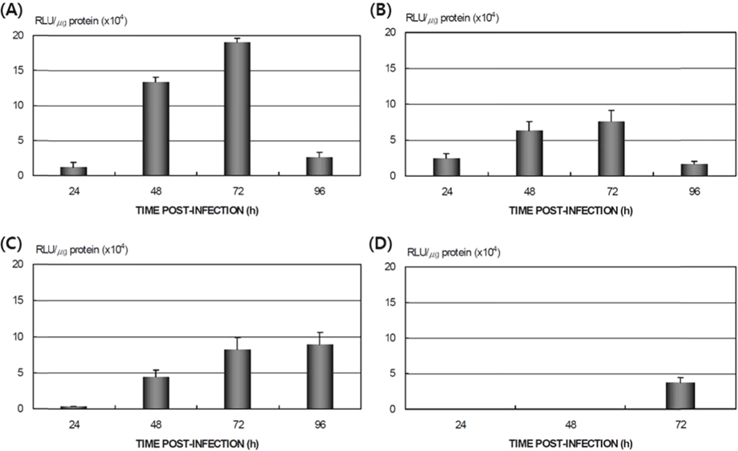
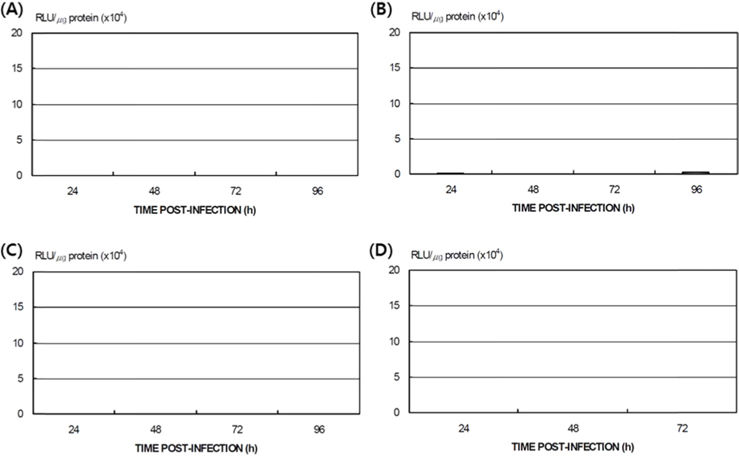
Differences in AcNPV reproduction in the HPS and LPS
To study the reproduction of AcNPV in
>
Expression of AcNPV in B. mori tissue
>
Proliferation of AcNPV/BmA3-Luc DNA in B. mori tissue
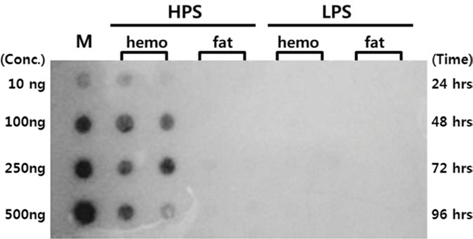
Dot blotting was performed to verify virus proliferation in the
>
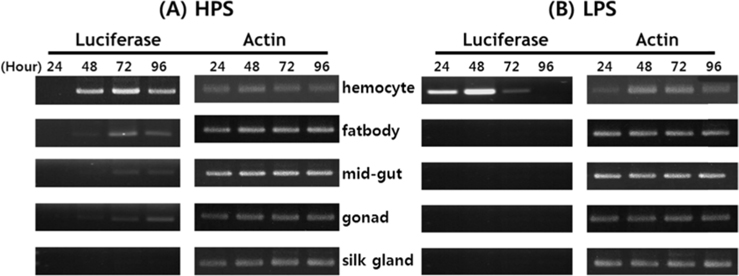
Viral RNA expression profile in B. mori tissue
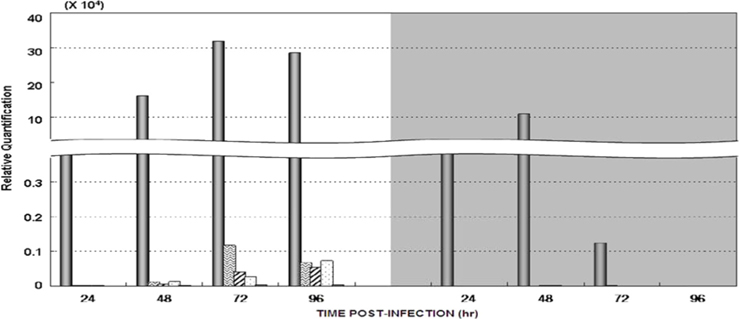
Dot blotting of recombinant viral DNA showed that virus proliferation was high in the HPS. In order to confirm this result, viral RNA expression within different tissues was studied using RT-PCR and real-time PCR. HPS S10 and LPS S39 were used The luciferase gene was observed in hemocytes, fat cells, the mid-gut, and the gonads. It was observed beginning 48 h after infection in blood cells and reached its maximum level within 72 h The expression level of luciferase in fat cells, the mid-gut, and the gonads was less than that seen in hemocytes, and viral RNA began to be observed after 72 h, 24 h later than in hemocytes (Fig. 8A). The luciferase gene was only detected in hemocytes inthe LPS(Fig. 8B). The real time PCR results corresponded with those from the RT-PCR. The luciferase gene was observed in all tissues from the HPS. only observed in hemocytes, and it reached its maximum levels within 48 h. Proliferation of the luciferase gene was not detected in tissues other than hemocytes (Fig. 9).
Functional development of function, the synthesis of active substances, and genes, of insects are receiving more attention in life science the utilization of insects is increasing. The useful production of biological material using insect tissues is increasing, by taking advantage of insect behavior and their specific characteristics (Tamura
AcNPV infection of
It has been reported previously that the proliferation of
Virus proliferation in