Worldwide, the family Gonostomatidae (order Stomiiformes) consist of 23 species in 5 genera (
In February 2012, a juvenile of a Gonostomatidae species was collected at a depth of 100 m at Jeju Island, Korea (32.54°N, 126.05°E) in a RN80 net (mouth opening 0.8 m, mesh size 0.33 mm). The specimen was fixed in 99% EtOH and later deposited at the National Institute of Biological Resources (NIBR-P 0000018076), Korea.
>
DNA extraction, PCR, and sequencing
Total DNA was extracted from the eyeball of the specimen using 10% Chelex 100 resin. Polymerase chain reaction (PCR) was used to amplify the mitochondrial DNA cytochrome oxidase subunit I (mtDNA COI) gene using the previously reported PCR primers VF2 (5′-TCA ACC AAC CAC AAA GAC ATT GGC AC-3′) and FishR1 (5′-TAC ACT TCT GGG TGG CCA AAG AAT CA-3′) (Ivanova et al., 2007). The
PCR conditions were as follows: initial denaturation at 94℃ for 2 min, 35 cycles of denaturation at 94℃ for 30 s, annealing at 50℃ for 40 s, extension at 72℃ for 1 min, and a final extension at 72℃ for 10 min.
The DNA was sequenced using an ABI 3730XL sequencer and an ABI PRISM BigDye Terminator v3.0 Ready Reaction Cycle Sequencing Kit (Applied Biosystems Inc., Foster City, CA, USA). For comparison we obtained mtDNA COI sequences for gonostomatid species from the National Center for Biological Information (NCBI) database, including
Counts and measurements followed the methods of Nakabo (2002), and each body part was measured to the nearest 0.1 mm using the Active measure program (Korea). Morphological identification followed Okiyama (1988) and Ozawa and Katayama (2003).
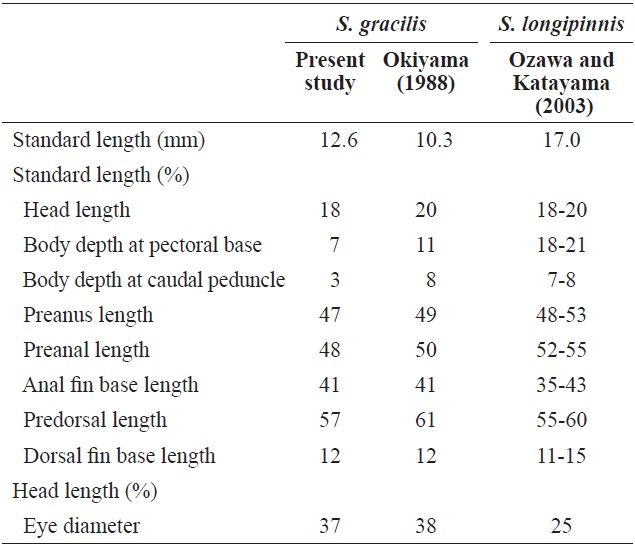
[Table 1.] Comparison of meristic characters of Sigmops gracilis and three Sigmops species

Comparison of meristic characters of Sigmops gracilis and three Sigmops species
Counts and proportional measurements for the
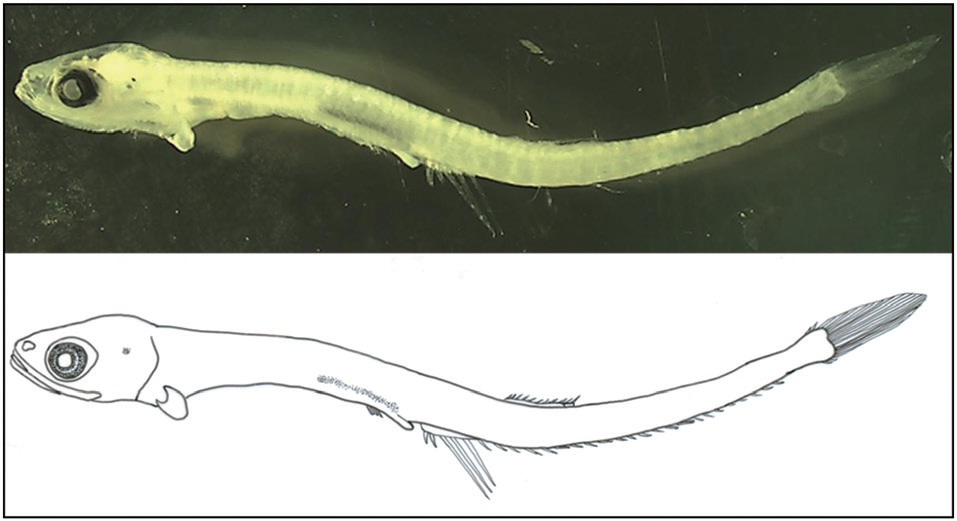
Body moderately compressed and elongate. Head moderate and eyes round. Eye diameter longer than the snout length. Body depth uniform except at the caudal peduncle. Mouth very large with hooked teeth, with the posterior tip of the upper jaw reaching to the posterior border of the eye. Notochord flexion complete and all fins develop at their fixed locations. Pelvic and pectoral fins small. Anus not attached to anal fin base, but very close to it. Origin of dorsal fin below the 7th-8th anal fin ray. Anal fin base length much longer than the dorsal fin. Melanophores developed on lateral occipital and dorsal gut. Photophores not observed.
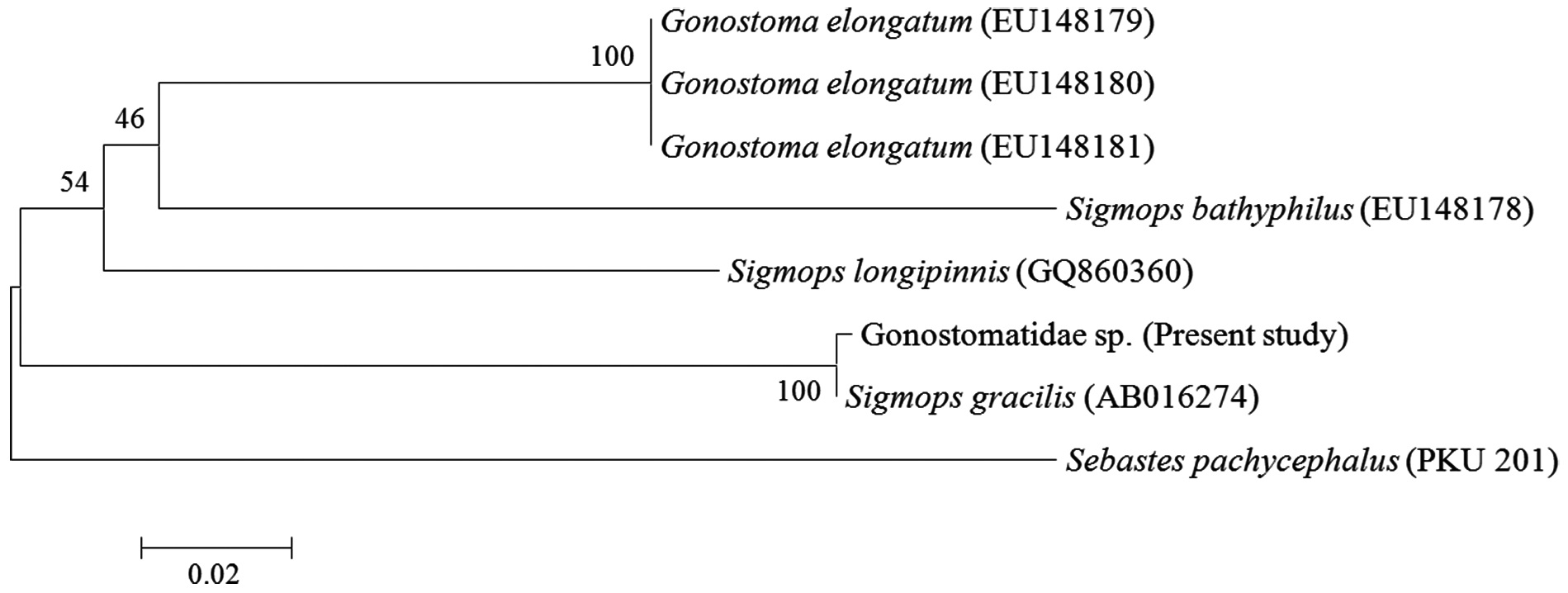
A mtDNA COI sequence of 499 base pairs (bp) derived from the eyeball of the Gonostomatidae juvenile was compared with sequences from other gonostomatid species and an outgroup. The smallest genetic distance was between the juvenile and
was identified as
Alaska (Mecklenburg et al., 2002), China (Yang et al., 1996), Japan (Masuda et al., 1984), Kuril Islands (Mecklenburg et al., 2002), Taiwan (Shao, unpublished data), and Korea (present study).
Juveniles of
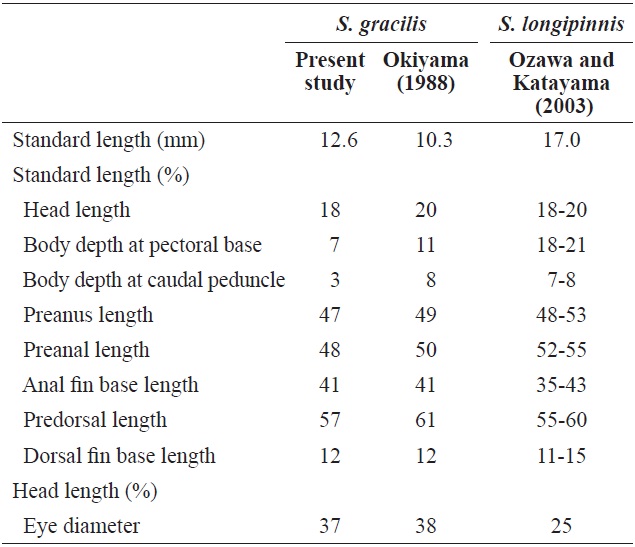
Comparison with the proportional measurements of Sigmops gracilis and Sigmops longipinnis juveniles
opercle at 8.0 mm SL, while