The intimate physical interaction between food algae and sacoglossan sea slug is a pertinent system to test the theory that “you are what you eat.” Some sacoglossan mollusks ingest and maintain chloroplasts that they acquire from the algae for photosynthesis. The basis of photosynthesis maintenance in these sea slugs was often explained by extensive horizontal gene transfer (HGT) from the food algae to the animal nucleus. Two large-scale expressed sequence tags databases of the green alga Bryopsis plumosa and sea slug Placida dendritica were established using 454 pyrosequencing. Comparison of the transcriptomes showed no possible case of putative HGT, except an actin gene from P. dendritica, designated as PdActin04, which showed 98.9% identity in DNA sequence with the complementary gene from B. plumosa, BpActin03. Highly conserved homologues of this actin gene were found from related green algae, but not in other photosynthetic sea slugs. Phylogenetic analysis showed incongruence between the gene and known organismal phylogenies of the two species. Our data suggest that HGT is not the primary reason underlying the maintenance of short-term kleptoplastidy in Placida dendritica.
Photosynthesis in some sacoglossan sea slugs offers a unique model to study the possibility of horizontal gene transfer (HGT) between multicellular predator and prey. Sacoglossan mollusks ingest and actively maintain chloroplasts that they acquire from large coenocytic green algae and keep them for up to several months (Wägele et al. 2011). These kleptoplastidic associations vary greatly in terms of specificity of the animal towards its algal prey and in retention time and functionality of the captured plastids (e.g., Rumpho et al. 2011, Klochkova et al. 2013). The basis for long-term maintenance of photosynthesis in these sea slugs has often been explained by extensive HGT from the nucleus of the alga to the animal nucleus, followed by expression of algal genes in the gut to provide essential plastid-destined proteins (Bhattacharya et al. 2013).
Sacoglossan mollusk
In this study, we present initial results from the comparative analysis of transcriptomes of
Adult animals of
For transcriptome analysis, sea slugs were kept without any food algae (i.e., starved) for 28 days and the Petri dish with culture medium was changed every day. Egg ribbons were harvested from one week after starvation, when no chloroplasts remained in the digestive tract of the sea slugs, because defecation stopped by that time and their body color was turning yellowish by each passing day. The harvested eggs were rinsed with 3% H2O2, frozen in liquid nitrogen and kept in deep freezer (−70℃) until use. Other sacoglossan sea slugs,
All algae (
>
Isolation of total RNA and mRNA purification
Total RNA from
>
Genomic DNA isolation, polymerase chain reaction (PCR) and sequence determination
DNA was purified from algal samples from different localities (
Transcriptome of
The actin sequences examined in this study were aligned with the actin sequences from GenBank using MacClade 4.08. The nucleotide alignment contained 65 sequences and was trimmed to 778 nucleotide comparable positions without third codon position using Paup*v. b10. Maximum likelihood analysis was performed using RAxML 7.0.4 with rapid bootstrapping option and 1,000 replicates under GTR + I + Γ model. Tree was visualized and graphic versions were exported using FigTree v1.4.0. New actin sequences generated in this study have been deposited in NCBI under the accession numbers listed in
Functional annotation on
Comparison of two large-scale ESTs databases of
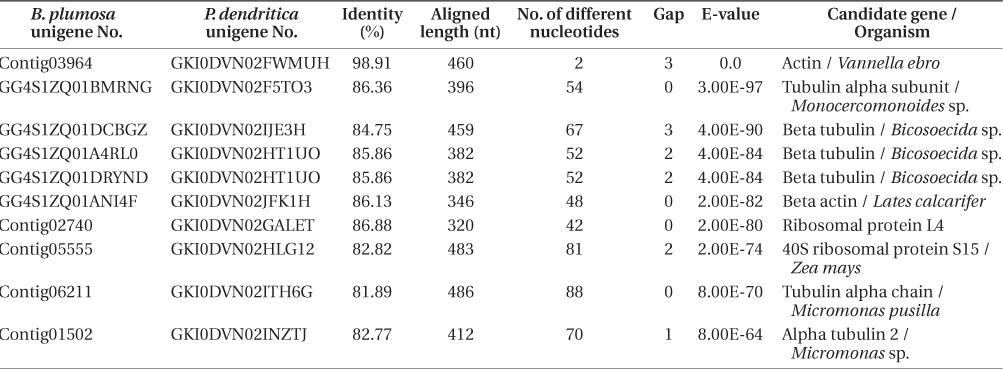
Results of BLAST analysis between the transcriptomes of Bryopsis plumosa and Placida dendritica
[Table 2.] Actin homologues from Placida dendritica and Bryopsis plumosa

Actin homologues from Placida dendritica and Bryopsis plumosa
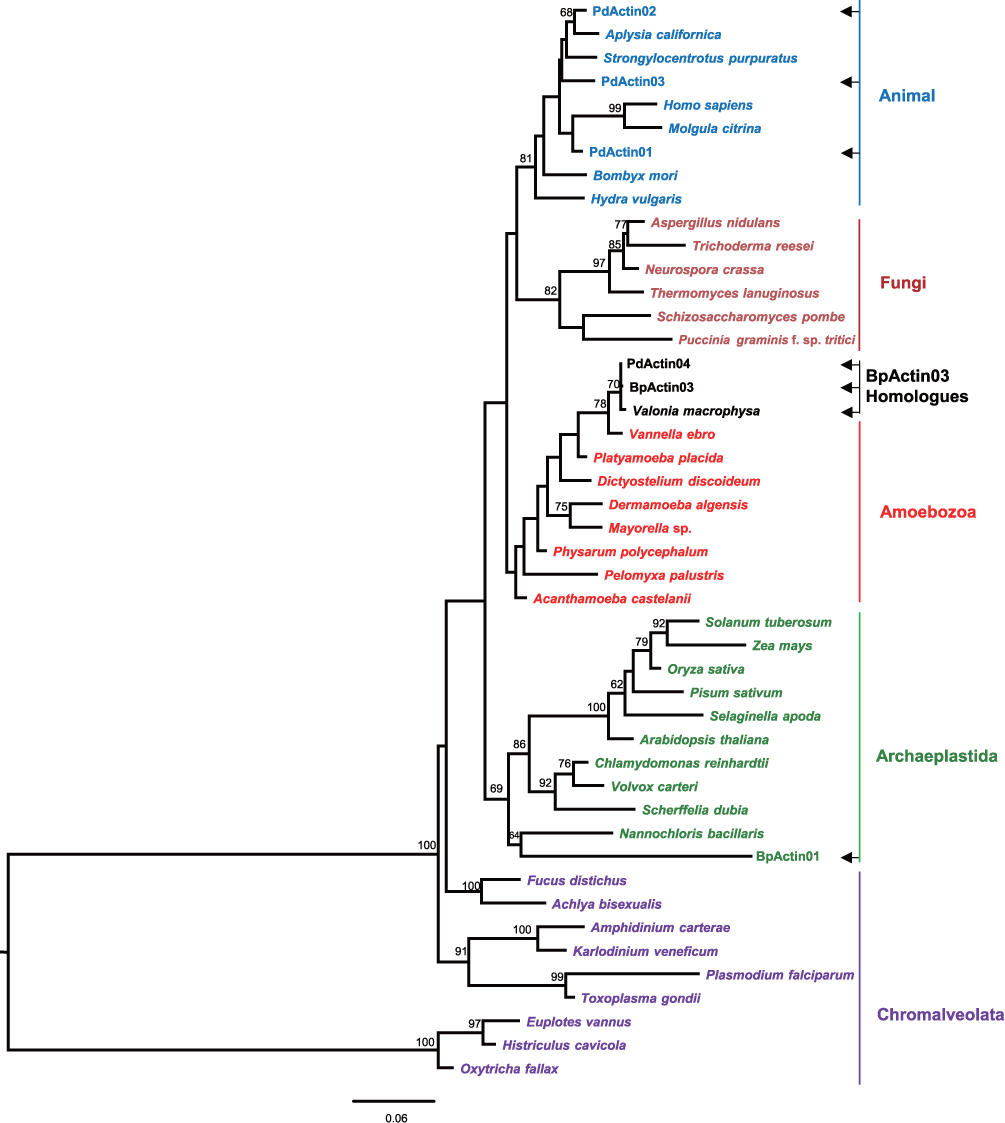
Phylogenetic analysis showed incongruence between the actin gene and known organismal phylogenies of the animals and algae (Fig. 3). Surprisingly, all BpActin03 homologues were grouped within a branch of Amoebozoa, while other actin genes from the two species were grouped appropriately in the Archaeplastida and Opisthokonta (Fig. 3). To check if PdActin04 homologues are from simultaneous contamination of some Amoebozoa species, the transcriptomes of
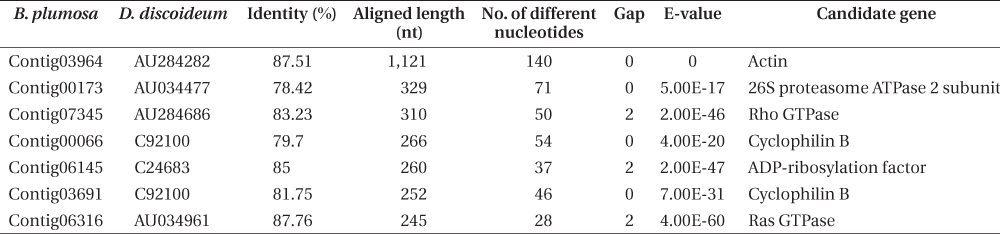
Results of BLAST analysis between the transcriptomes of Bryopsis plumosa and Dictyostelium discoideum
Our results did not show any photosynthesis-related genes in
The gold standard for identifying HGT with confidence is phylogenetic incongruence and this occurs if there is strong conflict between the phylogenies of the gene and of the organism (Keeling and Palmer 2008). The incongruence between the gene and known organismal phylogenies of the herbivore and algae did not support that PdActin04 has been horizontally transferred from its food algae. PdActin04 and all of its homologues found in green algae were nested in a branch of the Amoebozoa phylogeny (Fig. 3). It is possible that BpActin03 homologues are from a common amoeba contaminating all algal strains as well as the sea slugs simultaneously. If the contamination was from laboratory culture, all BpActin03 homologues should have the same sequence.
However, DNA sequences of nine BpActin03 homologues were clearly different among species. It means there must be nine different amoebas specifically contaminating algal strains, as well as the sea slug. It is hard to believe that each algal strain, collected from different localities of the world in different times, carry a specific amoeba and never mixed during years of laboratory culture. Most of all, the sequence difference among the homologues reflected the phylogenetic distance among ulvophyceaen algae; species closely related showed less sequence difference (
Direct comparison of large EST databases enabled us to avoid a long-lasting concern with HGT studies, that of contamination of targeted genes. If the materials used for building EST database of
The intimate physical interaction between herbivore and food algae may lead to horizontal transfer of certain genes. The short-term kleptoplastidy occurring in