More than 20 and 10 clades / ecotypes of Synechococcus and Prochlorococcus, respectively, have been identified in various oceanic regions. However, their diversity has yet to be thoroughly studied in the northwest Pacific Ocean. Further, spatial distribution of Synechococcus clades in the oligotrophic oceans has been scarcely characterized. To elucidate picocyanobacterial lineage distribution in the northwest Pacific Ocean, 16S-23S internal transcribed spacer sequences of picocyanobacteria were sequenced by barcoded amplicon pyrosequencing method. Additional pyrosequencing library using a primer specific for the Synechococcus subcluster-5.1 was constructed to thoroughly understand Synechococcus diversity in the oligotrophic oceans. In warm pool area, Prochlorococcus was predominant and showed a distinct depth-partitioning between HLII and LL ecotypes. Despite low abundances, diverse Synechococcus clades appeared in the oligo-trophic open ocean, showing both vertical and horizontal niche partitioning. Clade II was the predominant Synechococcus clade, especially in upper euphotic depths. In shallow and middle euphotic depths, clades UC-A, III, and CRD1 were distributed broadly. However, a distinct shift in the horizontal distribution was found at ca. 20° N. Conversely, clades XVII and CRD2 dominated at deep euphotic depths and constituted a higher proportion than clade II. These niche-partitioning of Synechococcus clades seemed to be related with temperature, nutrient concentration as well as iron concentration.
The lineage diversity of picocyanobacteria has been studied using various genetic markers, and this has revealed the existence of very diverse
Picocyanobacterial diversity can be unveiled using a clone library approach or probe-based molecular techniques, such as dot-blot hybridization and quantitative polymerase chain reaction for specific clades or ecotypes (Ahlgren et al. 2006, Fuller et al. 2006, Zinser et al. 2006, Martiny et al. 2009, Tai and Palenik 2009, Lavin et al. 2010, Rusch et al. 2010, Huang et al. 2012, Sohm et al. 2015). Recently, barcoded amplicon pyrosequencing methods were developed to reveal picocyanobacterial and
The northwestern Pacific Ocean is composed of diverse ecological environments, oligotrophic tropical / subtropical warm pools, oligotrophic Kuroshio Current (KC) and mesotrophic / eutrophic marginal seas. The East China Sea (ECS) is the largest marginal sea affected by oligotrophic pelagic water at the east and by coastal water at the west. Furthermore, the ECS is experiencing most rapid warming (Belkin 2009). The warm pool area has the warmest surface water of the ocean with oligotrophic features (McClain et al. 1999) and is expected to undergo significant changes as consequences of ENSO (El Niño Southern Oscillation) variability (Collins et al. 2010). Therefore, NW Pacific Ocean is an important region to study tele-connections between open oceans and marginal seas and to understand the ecological impacts of changing ocean, such as warming and acidification. Despite the importance, NW Pacific steadily remains one of least explored areas on earth.
To understand distribution and ecological niches of clades / ecotypes of
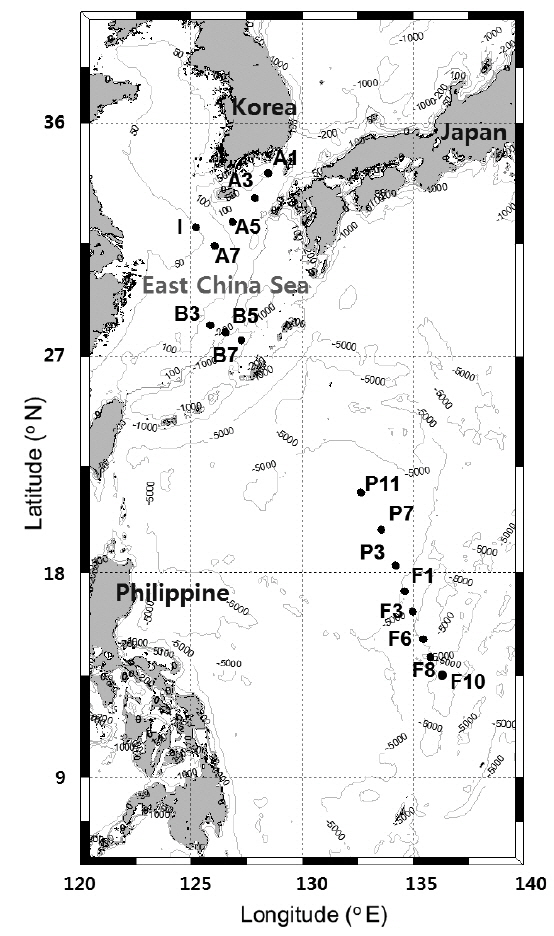
Water samples were collected at 16 stations during the POSEIDON (Northwestern Pacific Ocean Study on Environment and Interactions between Deep Ocean and marginal seas) cruise in the northwestern Pacific from May 26 to June 12, 2010 aboard the R/V Onnuri (Fig. 1). Stations (Stns) on lines F and P are located in a tropical area affected by the oligotrophic North Equatorial Current (NEC). Stations on line B are located in a subtropical area mainly affected by the KC. Stations on line A and Stn I are located in the ECS, where coastal currents and a branch current (Korea / Tsushima Warm Current) from the KC meet, forming oceanic fronts in the central continental shelf region (Ichikawa and Beardsley 2002). At each station, seawater was sampled from four to six depths between the surface and deep chlorophyll maximum (DCM) depth using Niskin bottles attached to a rosette sampler.
>
DNA extraction, polymerase chain reaction (PCR) amplification, and pyrosequencing
The genetic diversity of picocyanobacteria was analyzed in samples collected between the surface and DCM depth. Two liter water samples were passed through a 0.2-μm Supor filter (Gelman Sciences, Ann Arbor, MI, USA), and the filters were frozen at -80°C after adding 1 mL STE buffer (100 mM NaCl, 10 mM Tris-HCl, 1 mM EDTA, pH 8.0). DNA was extracted using the method of Choi et al. (2015).
To elucidate the picocyanobacterial ITS sequence diversity, barcoded amplicon pyrosequencing with GS-FLX Titanium (454 Life Sciences, Branford, CT, USA) was carried out. To amplify partial picocyanobacterial ITS sequences, picocyanobacteria-specific ITS-af forward and ITS-ar reverse primers were used (Choi et al. 2013). The amplification and pyrosequencing was conducted following the procedures shown in Choi et al. (2013). As
>
Analyses of pyrosequencing data
The obtained sequences were analyzed using the mothur program (Schloss et al. 2009) according to the overall procedures presented in Choi et al. (2014). Briefly, raw reads were filtered to remove reads associated with errors by allowing only a perfect match to the barcode and forward primer (in case of picocyanobacterial library, two mismatches to forward primer were allowed) sequences. The allowed number of maximum homopolymers was six. Reads with initial noisy flow (0.5-0.7) before 150 were removed and flows beyond 350 were ignored. The filtered reads were de-noised using the ‘shhh.flows’ command and the chimeric sequences were removed by ‘chimera.perseus’ command. The alignment and classification of each read were conducted using the database constructed in the previous study (Choi et al. 2014).
>
Characteristics of the pyrosequencing runs
In the analyses of the amplicon library of picocyanobacteria, 49,116 sequences passed our trimming and screening procedures. The number of sequences per sample varied between 97 and 3,290, and averaged 572. In the
>
Distribution of Synechococcus and Prochlorococcus in different oceanic regions
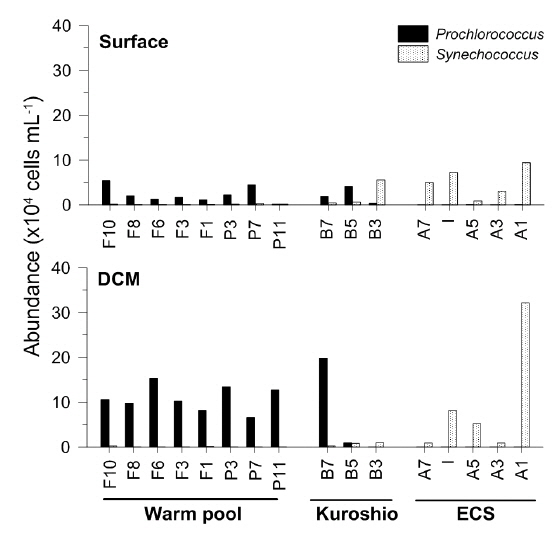
The abundance of
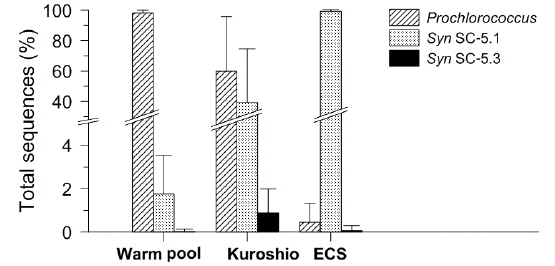
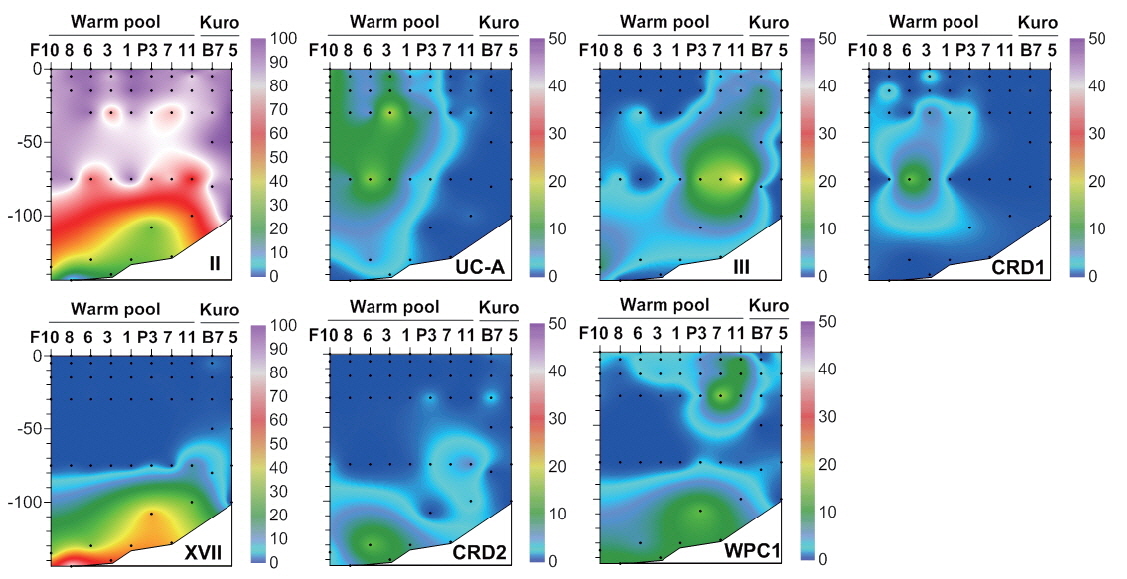
Similarly, the picocyanobacterial amplicon library showed distinct picocyanobacterial diversity among regions (Fig. 3). In tropical warm pool,
>
Phylogenetic diversity of picocyanobacteria in the NW Pacific Ocean
In the warm pool area of the NW Pacific, HLII ecotype
Similarly, the dominance of
>
Synechococcus diversity in the oligotrophic oceans
Physiological and genetic diversity and ecological adaptation of
Fifteen clades belonging to
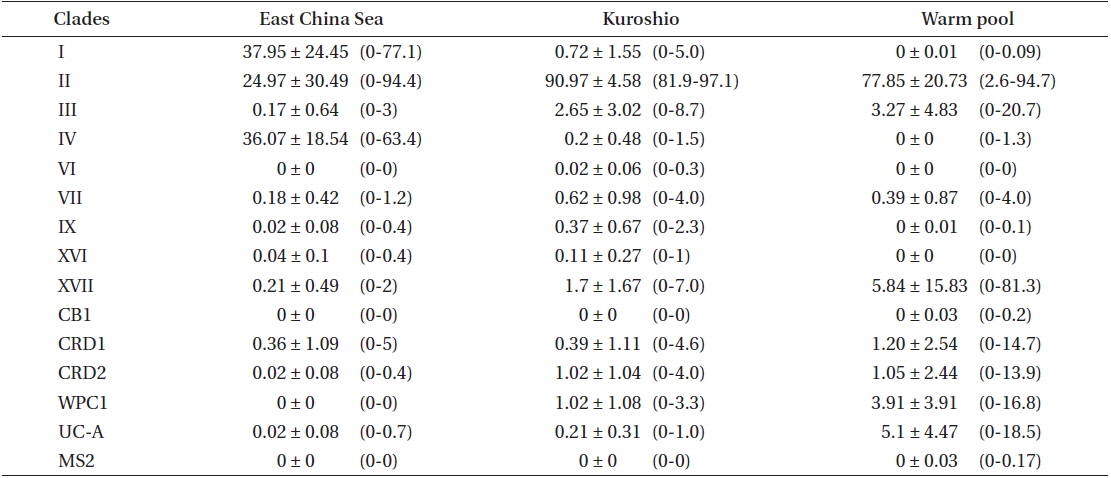
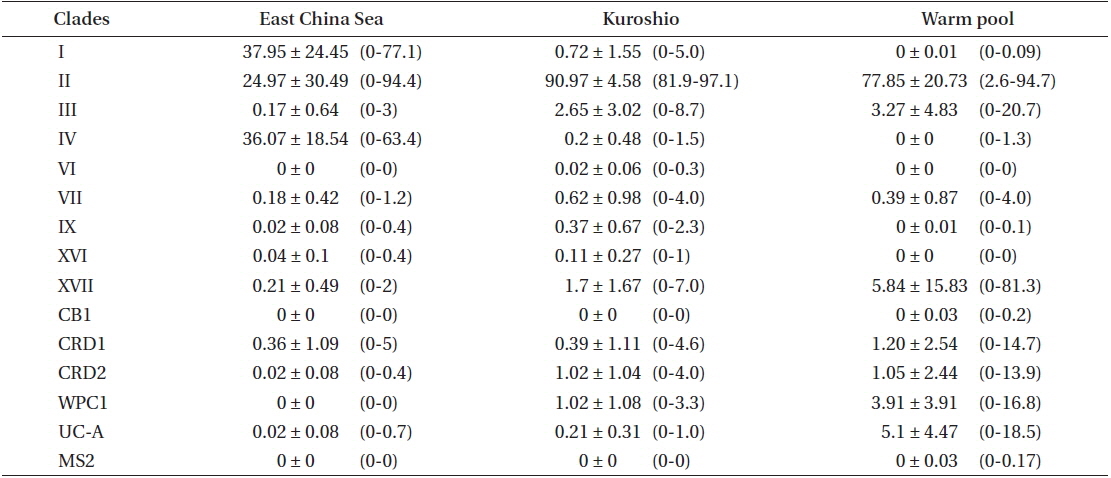
Percentages (mean ± 1 standard deviation, ranges) of Synechococcus lineages found in this study as proportions of all sequences belonging to Synechococcus subcluster-5.1 sequences
The distributions of clades V and VI have not been studied intensively due to methodological limitations. Most studies of the distribution of
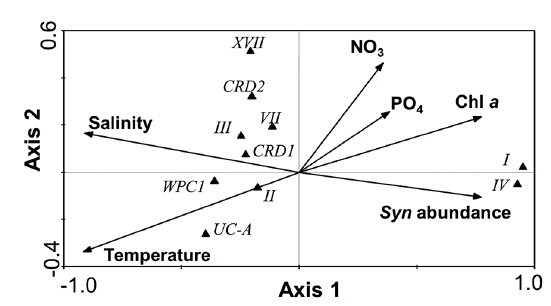
For
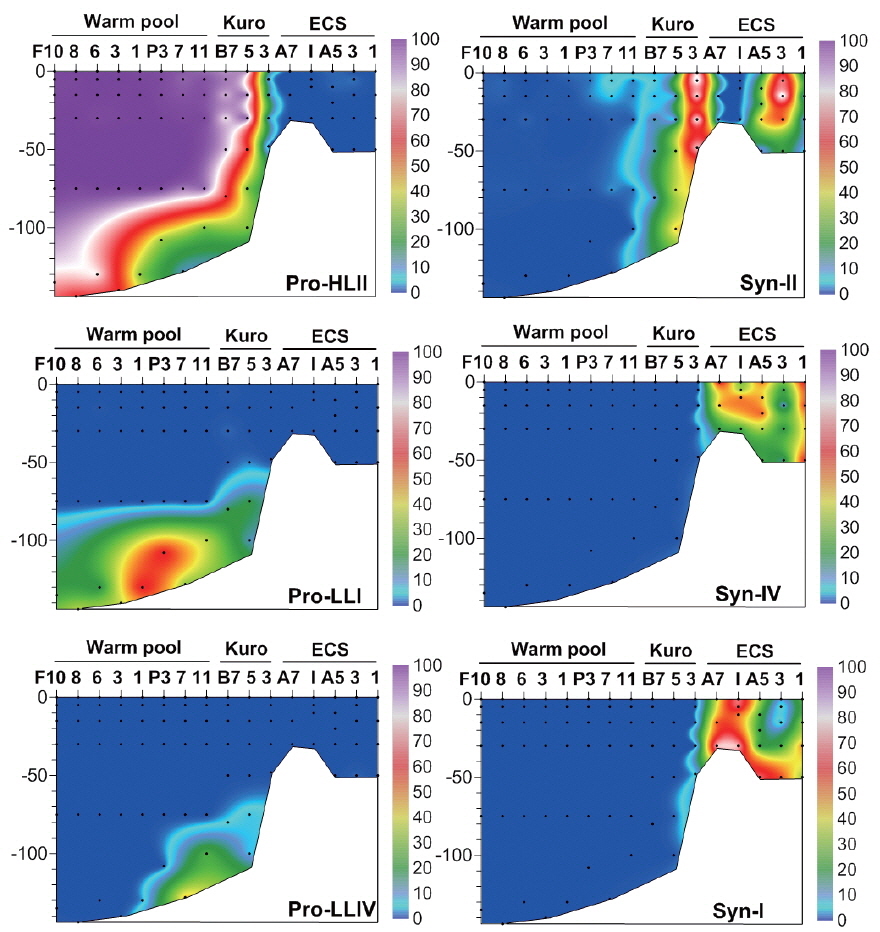
Clade CRD1, first identified as a dominant
Conversely, clades XVII and CRD2 had relatively high proportions at deeper euphotic zone (Fig. 5), suggesting their adaptation to relatively low temperature and low light (<3% of surface irradiance) or relatively high nutrient conditions. However, their niche was restricted to tropical waters. In the South China Sea, they were found only at mid-euphotic depths (Huang et al. 2012). However, their appearance in the surface waters of the equatorial Pacific Ocean (Huang et al. 2012) suggested that the distribution of clade XVII is not governed by light intensity only.
By definition, the deep euphotic zone is characterized by low light intensity, and also usually has relatively high nutrients and low temperatures compared to surface water. Light intensity plays an important role in discriminating the ecotypes of
Distribution of picocyanobacterial ecotypes / clades could be clearly differentiated among regions in the NW Pacific Ocean.
Using a