The genus of leafy green algae, Prasiola Meneghini includes marine, terrestrial (mostly supralittoral), and freshwater species characterized by monostromatic blades generally expanding above and narrowing to a short stipitate region at the base (Rindi et al. 2004, 2007, Moniz et al. 2012a, 2012b, Guiry and Guiry 2015). A total of 36 species have been reported in this genus, at least 14 of which are freshwater organisms (Moniz et al. 2012b, Guiry and Guiry 2015).
In east Asia, 11 species and one variety have been reported as freshwater or subaerial Prasiola species: P. crispa (Lightfoot) Kützing, P. elongata Hu, P. fluviatilis (Sommerfelt) Areschoug ex Lagerstedt, P. formosana Okada, P. formosana var. coreana Okada, P. hubeica Bi L.-J., P. japonica Yatabe, P. lanpingensis C. Qian et R. -N. Wang, P. sinica C. -C. Jao, P. subareolata Skuja, P. tibetica C. -C. Jao, and P. yunnanica C. -C. Jao. All of these species, except one variety, have been reported in China (including Taiwan), and only one freshwater species, P. japonica has been reported in Japan (Yatabe 1891). In Korea, two Prasiola species, P. formosana var. coreana and Prasiola sp., have been identified. Prasiola formosana var. coreana Okada was firstly reported from Muncheon in North Korea (Okada 1939). This variety is distinguished from P. formosana on the basis of the thickness of the thallus, the dimension of cells in cross-sections, and the outward form of the frond (Okada 1936, 1939). Okada (1939) pointed out the similarity of this variety to P. japonica, which is commonly distributed in Japan, in terms of the shape of cells in cross-sections and the external appearance of the frond. In South Korea, a different type of Prasiola species was reported by Park et al. (1970). This species was found growing along a small stream originating from Chodanggul, a limestone cave in Samcheok, Gangwon Province. This species was later identified as P. japonica by Chung (1993).
Molecular phylogenetic studies and DNA barcode marker analyses in Prasiola have primarily been conducted in marine species from Europe, North America, Tasmania, New Zealand, and the Antarctic (Sherwood et al. 2000, Rindi et al. 2004, 2007, Saunders and Kucera 2010, Heesch et al. 2012, Moniz et al. 2012a, 2012b, 2014).In contrast, the molecular phylogenies of Asian freshwater species have rarely been reported. Only Naw and Hara (2002) reported on the molecular phylogenetic position of a freshwater Prasiola species collected from Myanmar. These authors analyzed the 18S rRNA gene from Prasiola sp. as well as from P. japonica collected in Japan. Moniz et al. (2012b, 2014) generated plastid rbcL, psaB, and tufA genes of P. yunnanica collected from China to determine the phylogenetic position of this species. However, no Korean species have been examined in molecular phylogenetic analyses.
Therefore, the objective of the present study was to assess the molecular phylogenetic identity of the South Korean Prasiola species and to compare it with Japanese P. japonica and published representative species. To this end, we observed the morphological features and analyzed the plastid rbcL, psaB, and tufA genes sequences of the South Korean Prasiola species.
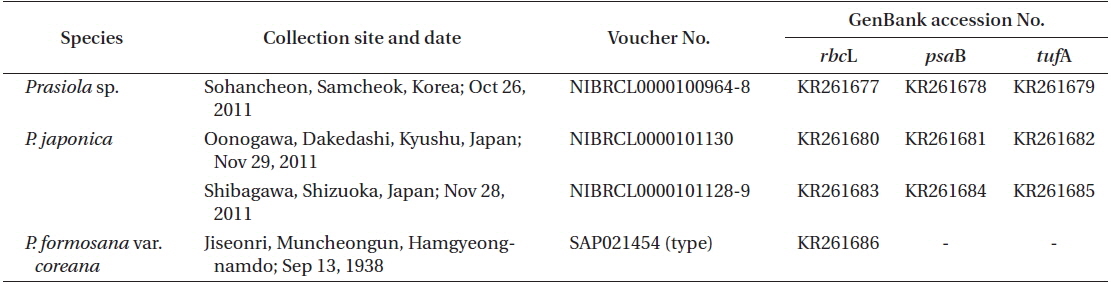
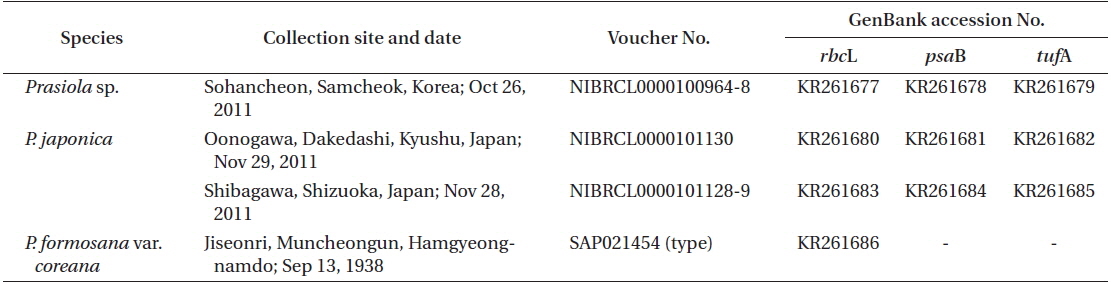
Thalli of Prasiola species were collected from the stream originating from Chodanggul Cave in Samcheok for both morphological observation and molecular analyses. We also collected P. japonica samples from Kyushu and Shizuoka in Japan for molecular analysis. Fresh material was used for surface observations using a microscope. Photographs were taken with a digital camera (C-4040 zoom; Olympus, Tokyo, Japan) attached to a light microscope (BX50; Olympus). All voucher specimens analyzed in this study have been deposited in the Plant Collections Storage of the National Institute of Biological Resources (NIBR) in Incheon, Korea (Table 1).
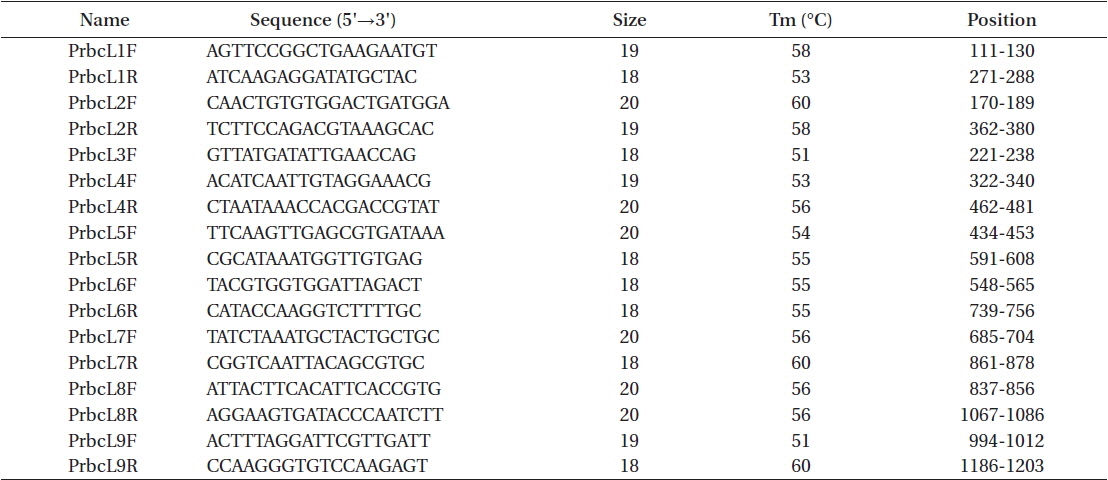
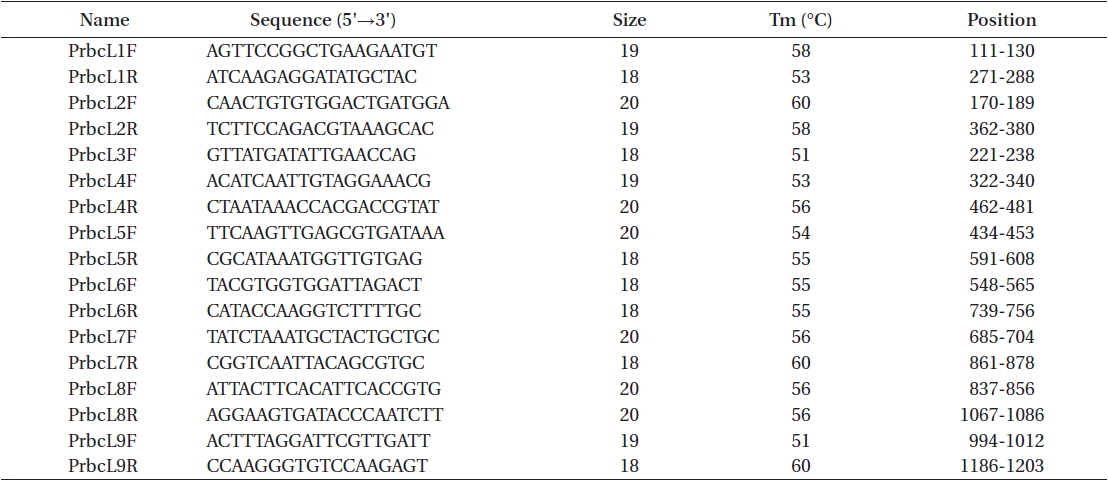
Total genomic DNA was extracted from silica gel-preserved dry materials. Approximately 0.05 g dried thallus was ground using TissueLyser (Qiagen, Austin, TX, USA) with two tungsten carbide beads (3 mm; Qiagen) at 25/s frequencies for 2 min. Ground powder was used for the DNA extraction procedure using a NucleoSpin Plant II kit (Macherey-Nagel, Düren, Germany), following the manufacturer’s protocol. Plastid rbcL regions were amplified using the PF2 / PR2 primer pair (Rindi et al. 2004) for fresh materials and nine newly designed primer sets for the type specimen of P. formosana var. coreana (Table 2, Fig. 1). Polymerase chain reaction (PCR) amplifications were conducted using AccuPower PCR premix (Bioneer, Daejeon, Korea) and a HotStarTaq Plus Master Mix Kit (for herbarium samples; Qiagen) on a GeneAmp PCR System 9700 (Applied Biosystems, Foster City, CA, USA). The amplification cycle consisted of an initial denaturation step at 95℃ for 4 min, followed by 30 cycles of denaturation at 95℃ for 30 s, annealing at 47 or 50℃ for 30 s, and extension at 72℃ for 1 min, with a final extension at 72℃ for 7 min. The psaB and tufA genes were amplified using the Pp1F / Pp4R (Novis et al. 2010) and tufGF4 / tufAR (Saunders and Kucera 2010) primer pairs respectively, as recommended by Moniz et al. (2012b, 2014) for Prasiola specifically. PCR amplification parameters for the psaB and tufA regions were the same as those for rbcL, except that the annealing temperature was 58 or 60℃.
The sequencing reactions were performed using ABIPRISM BigDye Terminator Cycle Sequencing Kits, and the florescent signals were detected using an ABI PRISM 3730XL Analyzer (Applied Biosystems). The sequences were verified using Chromas 1.45 (McCarthy 1996) and aligned visually using previously published data (Sherwood et al. 2000, Rindi et al. 2004, 2007, Saunders and Kucera 2010, Heesch et al. 2012, Moniz et al. 2012a, 2012b, 2014).
Maximum likelihood (ML) analysis was performed using RAxML 7.2.8 (Stamatakis 2006) under the GTR + Γ model. We used 200 independent tree inferences using the default option of automatically optimized subtree pruning and regrafting (SPR) rearrangements and 25 “distinct rate categories” options to identify the best tree. Bootstrap analysis was conducted for 1,000 replications.
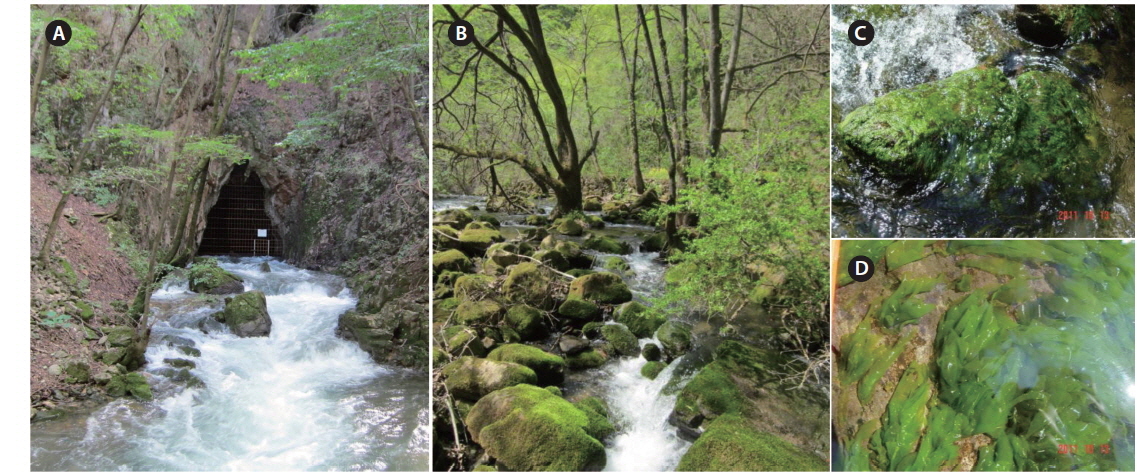
The South Korean Prasiola species grows on rocks along the Sohancheon stream, which is located 6.37 km from the East Sea. This species primarily grows along the central part of the stream, which shallows, exposed to light, and has a fast stream velocity (Fig. 2).
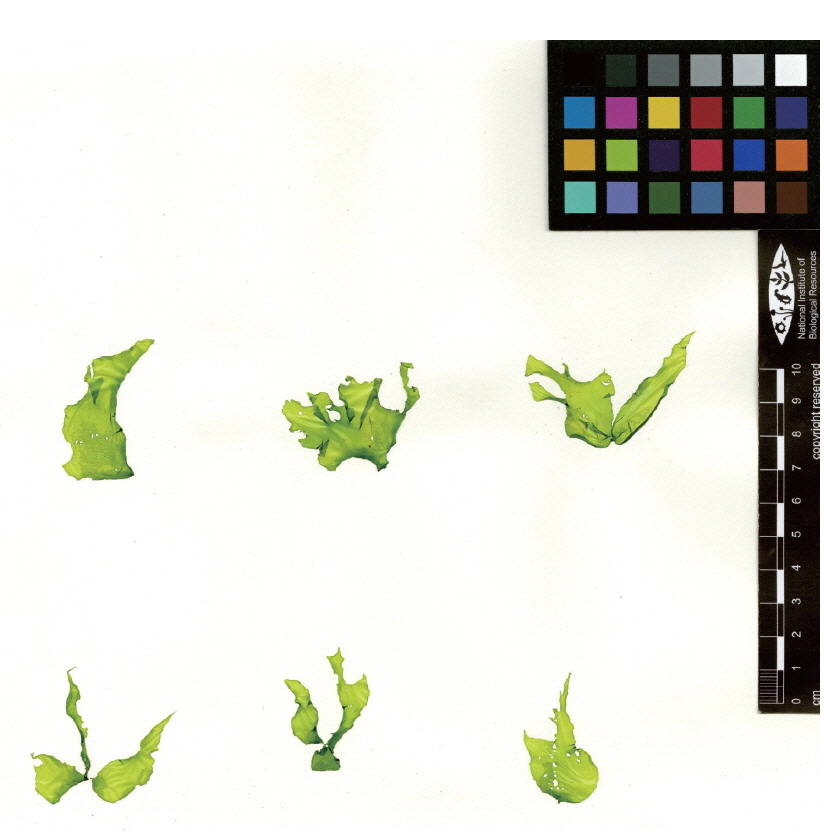
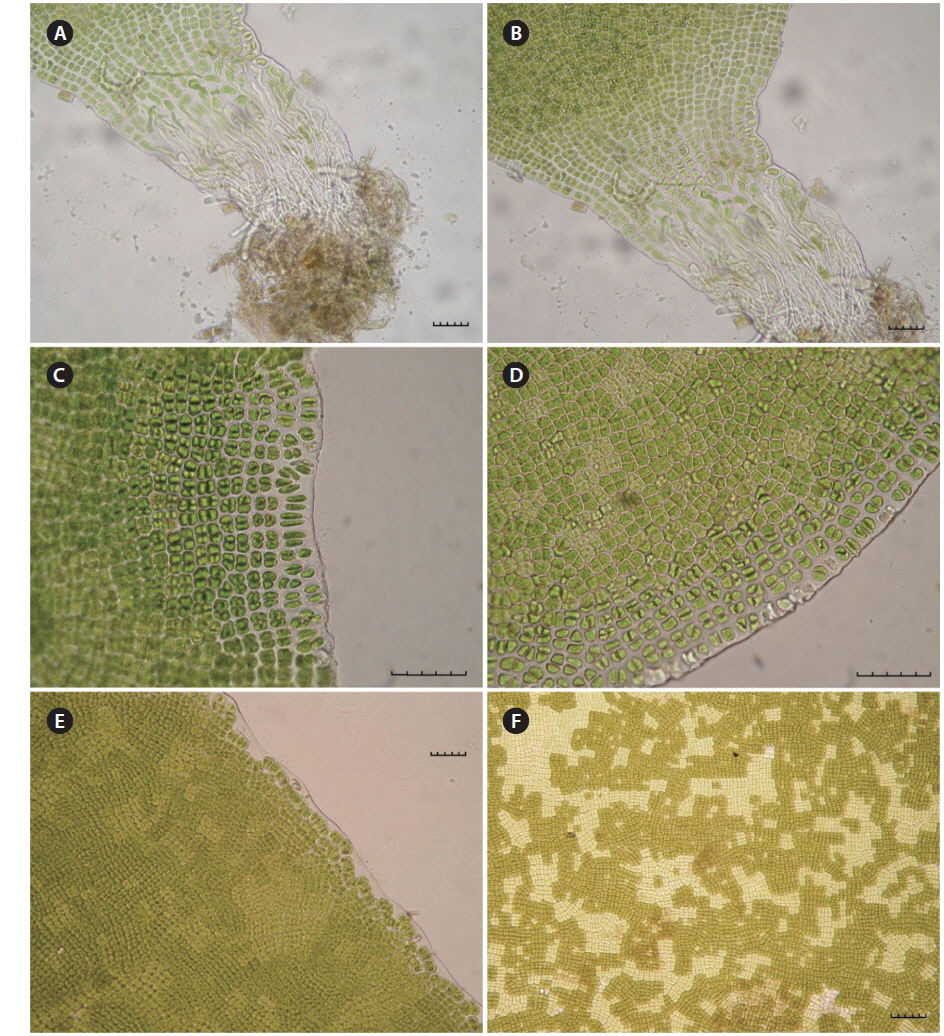
The thallus is light green, leafy, linear, lanceolate or ovate, 1-7 cm long, 0.5-3.5 cm wide (Fig. 3), and attached to the surface of rocks with a small mass of fibrous cells comprising the rhizoid (Fig. 4A & B). Vegetative cells are single-layered, round-square, and arranged in two or four cells together. At the margin of the thallus, cells are elliptical or lanceolate, and the space between cells gets wider than the inner part (Fig. 4C & D). Gametangial cells arise from vegetative cells and are arranged as an irregular mosaic from early winter to spring (Fig. 4E & F).
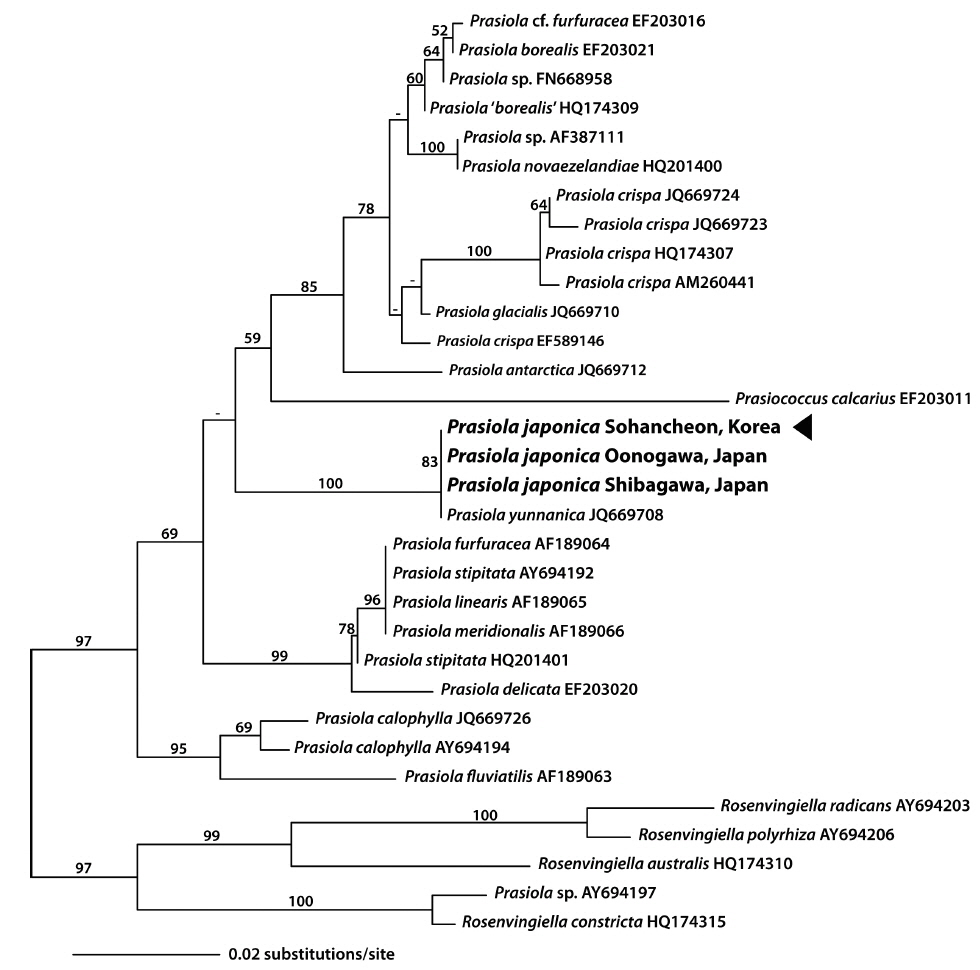
In the present study, we obtained 1,094 bp of three new rbcL sequences from a South Korean Prasiola species and two Japanese P. japonica. The plastid rbcL sequences of the Korean Prasiola species were identical to those of Japanese P. japonica and Chinese P. yunnanica (JQ669708), with the exception of one undesignated base (N) in the sequences by Moniz et al. (2012b). Pairwise distances of rbcL sequences ranged from 0 to 6.5% (average 3.5%) in the genus Prasiola. Based on the ML tree of rbcL, the South Korean Prasiola species was included in a monophyletic clade with P. japonica and P. yunnanica with a 100% bootstrap value (Fig. 5). The sister relationship of this clade was not clear in rbcL gene tree.
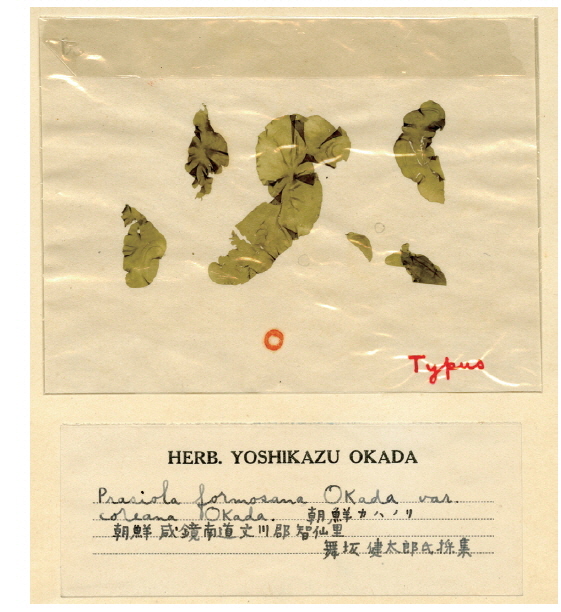
We also attempted to amplify the rbcL region from the type specimen (SAP021454) of Prasiola formosana var. coreana and succeeded in obtaining 196 bp of rbcL-3P using the PrbcL9F / 9R primer set. This short fragment was also identical to those of the South Korean Prasiola species and Japanese P. japonica. Although the resolutions for several species were low when the phylogenetic tree was generated using the 196 bp of rbcL-3P, the Asian Prasiola specimens were clustered as a strong monophyletic group (figure is not shown). Only 75 bp from these short sequences overlapped in comparison with sequences from P. yunnanica, but these sequences were identical among all species. Pairwise distances of these short rbcL-3P sequences ranged from 0 to 4.6% (average 2.2%) in the genus Prasiola.
A total of 1,292 bp of psaB and 813 bp of tufA genes sequences were generated from one specimen of South Korean Prasiola species and two specimens of Japanese P. japonica. These sequences were identical to published data for P. yunnanica (JQ669686 for psaB and KF993445 for tufA), except for one undesignated base (N) in the sequences of psaB by Moniz et al. (2012b, 2014).
Our morphological observations indicated that the South Korean Prasiola species had an external morphology that was mostly lanceolate or ovate, as described by Park et al. (1970), who also reported a similar arrangement of cells and gametangial mosaic pattern to what we found. According to the previous report of the Korean Prasiola species, morphological features and cell shape can vary among individuals (Park et al. 1970). Cell shape and arrangement were once regarded as diagnostic characteristics distinguishing Asian freshwater Prasiola species (Hu and Wei 2006). Indeed, the presence or absence of a stalk and disc-like holdfast can be used to distinguish several species. However, some characteristics, such as cell shape and arrangement, are quite variable not only at the species level, but also among individuals.
Park et al. (1970) noted that Prasiola species from Sohancheon is lanceolate and ovate, which is similar to all morphological types of P. japonica, P. formosana, and P. formosana var. coreana. When Okada (1939) first described P. formosana var. coreana, he also noted that this new species resembled P. japonica in the shape of cells in cross section and the external appearances of the thallus. Jao (1947) documented P. yunnanica as a new species based on morphological differences from P. japonica, noting that the thickness and shape of the thallus and the length and form of cells in sectional views of this Chinese species were “quite dissimilar” from the Japanese species. Therefore, it can be quite difficult to distinguish Asian freshwater species using only external morphology and anatomical features.
Three plastid genes, rbcL, psaB, and tufA genes of the South Korean Prasiola species were identical to those of Japanese P. japonica and Chinese P. yunnanica, the latter of which was published by Moniz et al. (2012b, 2014). The phylogenetic trees for the South Korean species were also similar to those published previously (Moniz et al. 2012b). Moniz et al. (2012b) noted that the unrelated lineage of P. yunnanica was particularly interesting; however, because they only analyzed one Asian species, their results are not necessarily applicable to other Asian freshwater species. The present study is the first report of rbcL, psaB, and tufA genes of Prasiola species from South Korea and P. japonica from Japan, as well as of partial rbcL sequences of P. formosana var. coreana.
Even though the type specimen is most important when delineating species, amplifying an adequate length of the gene from old specimens can be quite challenging due to DNA degradation. Our partial rbcL sequences of the type specimen of P. formosana var. coreana (SAP021454) included the 3′ portion (rbcL-3P), which is one of the best barcode markers for green macroalgae (Saunders and Kucera 2010). In addition, small-sized “mini-barcodes” are considered adequate to represent a sufficient number of signals for species identification. In insects, short DNA barcode sequences of mitochondrial cytochrome oxidase subunit I (less than 150 bp) can provide effective taxonomic sequence tags (Meusnier et al. 2008). Here, the pairwise distance value was moderate in the genus Prasiola when only 196 bp of rbcL-3P were compared.
The distributions of Prasiola species in Korea and Japan are similar. In South Korea, Prasiola only grows attached to small limestone rocks along the Sohancheon stream in Samcheok, which is a geologically limestone region (Park et al. 1970). In Japan, P. japonica is mostly distributed along pristine streams connected to the Pacific Ocean from central Honshu (Tochigi Prefecture) to Kyushu. Iwamoto (1984) reviewed the literature to clarify the relationship between the distribution of P. japonica and the geological features of the region. He reported that P. japonica is distributed within geologically distinct regions, such as the areas of Fossa Magna and along the Median Tectonic Line. Although he did not address the relationship between the distribution of P. japonica and mineralogically distinct areas, the regions described above are mostly limestone areas. Many recent studies have reported that P. japonica grows well within concrete waterways (Ishikawa et al. 2005, 2007, Ishikawa 2009). The type locality of P. formosana var. coreana is Muncheon, North Korea, which is also a limestone area (Okada 1939). Moreover, P. yunnanica, which was also molecularly identical to the Korean Prasiola species, has only been reported in Yunnan, another limestone area (although quite distant from the ocean). Additional research is necessary to determine whether the distribution of Asian freshwater Prasiola species is related to the geological features of their habitat.
Although Park et al. (1970) concluded that the South Korean Prasiola species appeared to be a new species, they also noted that three eastern Asian species, Prasiola japonica, P. formosana, and P. formosana var. coreana, could be regarded as a single species. Here, we conclude that the South Korean Prasiola species be considered as P. japonica based on its habitat, morphology, and plastid genes phylogenies, and propose to synonymize P. formosana var. coreana and P. yunnanica with P. japonica.
Type: Unknown. Syntype localities: Nikko (Province of Shimotsuke), Kiriu (Kozuke), Shibakawa (Suruga), and Oimura (Mino), March 1891 (Yatabe 1891:188). Taxonomic (heterotypic) synonyms: Prasiola formosana var. coreana Okada 1939 (J. Jpn. Bot. 15:450), P. yunnanica C. -C. Jao 1947 (Bot. Bull., Acad. Sin. 1:110).