The dinoflagellate genus Pelagodinium is genetically classified in distinct sub-clades and subgroups. However, it is difficult to determine whether this genetic diversity represents intra- or interspecific divergence within the genus since only the morphology of the type strain of the genus Pelagodinium, Pelagodinium bei, is available. An isolate associated with the genus Pelagodinium from Shiwha Bay, Korea, was recently cultured. This isolate was clustered with 3 to 4 strains from the Atlantic Ocean, Mediterranean Sea, and Indian Ocean. This cluster was distinct from the subgroup more closely associated with P. bei. The morphology of the isolate was analyzed using optical and scanning electron microscopy and was almost identical to that of P. bei except that this isolate had two series of amphiesmal vesicles (AVs) in the cingulum, unlike P. bei that has one series. When the pigment compositions of the isolate and P. bei were analyzed using high-performance liquid chromatography, these two strains had peridinin as a major accessory pigment and their pigment compositions were almost identical. In addition, the swimming behaviors of these two strains were very similar. The reexamination of the type culture of P. bei revealed two series in the cingulum as for the isolate. The new findings on the number of series of AVs in the cingulum, the pigment composition, and the swimming behaviors suggest that P. bei and the isolate are conspecific despite their genetic divergence. This study provides a basis to further understand the molecular classification within Pelagodinium combining genetic, morphological, pigment, and behavioral data.
Marine dinoflagellates are ubiquitous and play diverse roles in marine ecosystems (Jeong et al. 2010, 2012, 2015). The dinoflagellate
Extraction from pooled symbionts, combined host and symbionts, culture of symbionts established after microdissection, and bulk microplanktonic communities, followed by amplification or cloning, and sequencing have increased our knowledge on the distribution of symbionts associated with foraminifera around the world (Gast and Caron 1996, Shaked and De Vargas 2006, Siano et al. 2010, Decelle et al. 2012, Kok et al. 2014, De Vargas et al. 2015). The distribution of
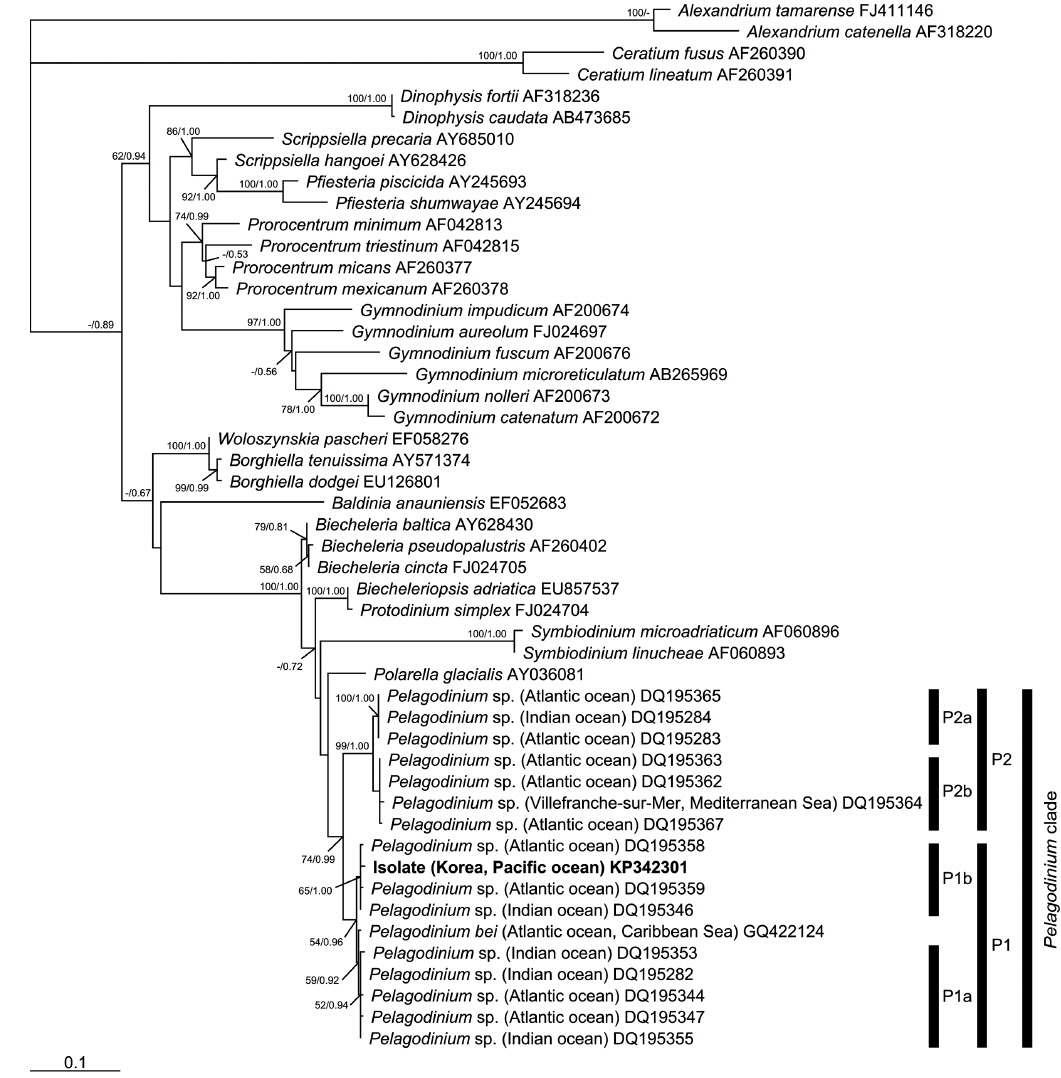
In the phylogenetic analysis, the type sequence of
The increase of studies assessing the diversity and distribution in the field using molecular tools makes the detailed characterization of strains associated to the various sequences and their comparison with the type species particularly relevant. However, details of the sulcal area, the internal transcribed spacers and 5.8S ribosomal RNA gene (referred to as ITS rDNA), and the pigment composition of the type species
We recently established a clonal culture of a small dinoflagellate able to grow photosynthetically from Shiwha Bay, Korea, that was related to the genus
The culture of the isolate from Korea was established from surface sediment samples collected on September 30, 2010 (depth, 11.5 m; surface temperature, 19.6℃; surface salinity, 11.7) from Shiwha Bay, Korea (37°18′ N, 126°36′ E). The surface sediment was collected from an Eckman grab (WILDCO; Wildlife Supply Company, Buffalo, NY, USA) and stored in the dark at 4℃ until further analyses. To concentrate potentially viable cells, between 1 and 2 cm3 of sediment were sieved through 100-μm and 15-μm Nytex meshes with filtered seawater. The 15-100 μm fraction was then transferred to a 100-mL beaker with filtered seawater. A manual vortex was applied and the suspended fraction was recovered. The remnant fraction was incubated in F/2-Si culture medium (Guillard and Ryther 1962) with a salinity of 32 at 20℃ under a light-dark cycle 14 : 10 at a photon flux of 20 μmol m−2 s−1. A cell swimming in the medium was isolated by micromanipulation and a monoclonal culture was established after two serial single-cell isolations.
>
Type culture of Pelagodinium bei
The culture that was used to establish the holotype of the type species
Cells were observed using a transmitted light inverted microscope (Zeiss Axiovert 200M; Carl Zeiss Ltd., Göttingen, Germany) at magnifications of ×50-1,000 to determine the general morphology and behavior. The measurements were determined with a Zeiss AxioCam MRc5 digital camera (Carl Zeiss Ltd.).
>
Scanning electron microscopy
For SEM, cells were fixed with 0.5-1% osmium tetroxide and rinsed in a dilution of 1 : 1 filtered seawater: distilled water. The cells were then washed in distilled water only. Cells were then subjected to a dehydration series in ethanol (10, 30, 50, 70, 90, and 100%). The cells were dried using a critical point dryer (CPD 030; BAL-TEC, Balzers, Liechtenstein). Finally, the cells were mounted on stubs, sputter coated with gold-palladium (SCD 005; BAL-TEC), and observed with an FE-scanning electron microscope (AURIGA; Carl Zeiss Ltd.).
For genetic analyses, a culture growing in the conditions described above was filtered through a polycarbonate membrane (25 mm, 3-μm pore size Whatman Nucleopore Track-Etch; GE healthcare, Buckinghamshire, UK) and resuspended by vortexing in distilled water in a 1.5-mL microtube (Scientific Specialties Inc., Lodi, CA, USA). The sample was subsequently centrifuged (WiseSpin CF-10 Microcentrifuge; DAIHAN Scientific Co., Ltd., Namyangju, Korea) at 7,500 ×g for 5 min at room temperature. The cells were immediately subjected to total DNA extraction using the AccuPrep Genomic DNA extraction kit (Bioneer Corp., Daejeon, Korea) according to the manufacturer’s instructions.
Amplicons of the ITS and LSU rDNA were obtained. The polymerase chain reaction (PCR) final mix concentrations were as follows: 1× PCR f-Taq buffer (fTaq DNA polymerase; SolGent Co., Ltd., Daejeon, Korea), 0.2 mM of dNTP (fTaq DNA Polymerase; SolGent Co., Ltd.), 0.4 μM of each primer, 0.025 U μL−1 of f-Taq DNA polymerase (fTaq DNA polymerase; SolGent Co., Ltd.), and 1.5 mM of MgCl2. A volume of 1.0 μL of the DNA extraction was used as template with a final reaction volume of 50 μL. The amplicons were obtained with the primer pairs ITSF2 and ITSR2 and 5.8 SF and LSUB (Litaker et al. 2003) with 54 and 50℃ as annealing temperatures (AT), respectively. PCRs were conducted using a thermal cycler (Mastercycler ep, model 5341; Eppendorf AG, Hamburg, Germany) as follows: one activation step at 95℃ for 2 min, followed by 35 cycles at 95℃ for 20 s, AT for 40 s, 72℃ for 1 min, and a final elongation step at 72℃ for 5 min.
Positive and negative controls were used for all amplification reactions. The size of the amplicons was verified on a 1.0% agarose gel. The products were visualized under a UV lamp. The PCR products were purified using the AccuPrep PCR purification kit (Bioneer Corp.) according to the manufacturer’s instructions. The purified PCR products were sent to the Genome Research Facility (School of Biological Science, Seoul National University, Korea) where they were sequenced on an ABI PRISM 3700 DNA Analyzer (Applied Biosystems, Foster City, CA, USA) with the primers used in the PCR.
The sequences of taxa used to construct the phylogenies were obtained from NCBI GenBank. The sequence of the isolate from Korea (strain HJ-2010), and the ITS rDNA sequence of the type species
The matrixes were also analyzed with MrBayes v3.2.3 (Ronquist and Huelsenbeck 2003) for Bayesian analyses. The models previously selected by Modeltest 7.3 were used. Four independent Markov chain Monte Carlo simulations were run simultaneously for 2,000,000 generations. Trees were sampled every 1,000 generations and the first 800 trees were deleted to ensure that the likelihood had reached convergence. A majority-rule consensus tree was created from the remaining 1,201 trees to examine the posterior probabilities of each clade.
The pigments were analyzed using HPLC (LC-10A system; Shimadzu Co., Kyoto, Japan) as in Zapata et al. (2000). A volume of culture containing 2,000,000 cells growing in the conditions mentioned previously was used in the analysis. The culture was filtered through a 1.2 μm pore-sized GF/C filter. Three milliliters of 95% methanol were used for extraction and a Waters C8 column (150 × 4.6 mm, 3.5-μm particle size, 0.01-μm pore size; Waters Corporation, Milford, MA, USA) for separation. Pigments were identified by retention times and absorption spectra identical to those of authentic standards, and quantified against standards purchased from DHI Water & Environment (Hørsholm, Denmark).
>
Morphological characterization
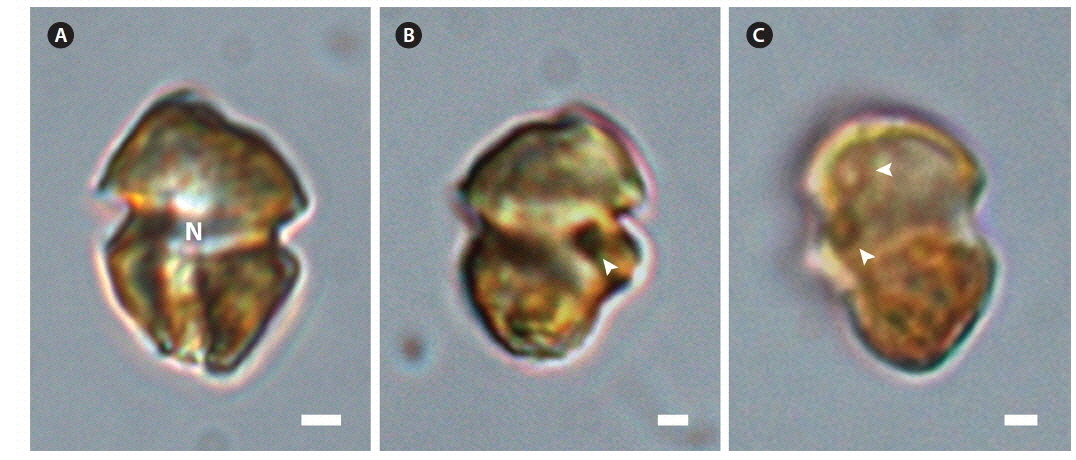
The ranges (and mean ± standard deviation) of cell length and cell width of living cells from the Korean isolate were 7.9-14.1 μm (11.1 ± 1.3, n = 100) and 5.1-10.8 μm (8.3 ± 1.1, n = 100), respectively. The chloroplasts were golden-yellow (Fig. 1). The epicone and hypocone were similar in size (Fig. 1A & B). The epicone was typically round to elliptical, while the hypocone was either round or slightly asymmetrical in ventral view (Fig. 1). An eyespot was present (Fig. 1B). Pyrenoids were occasionally visible with light microscopy (Fig. 1C).
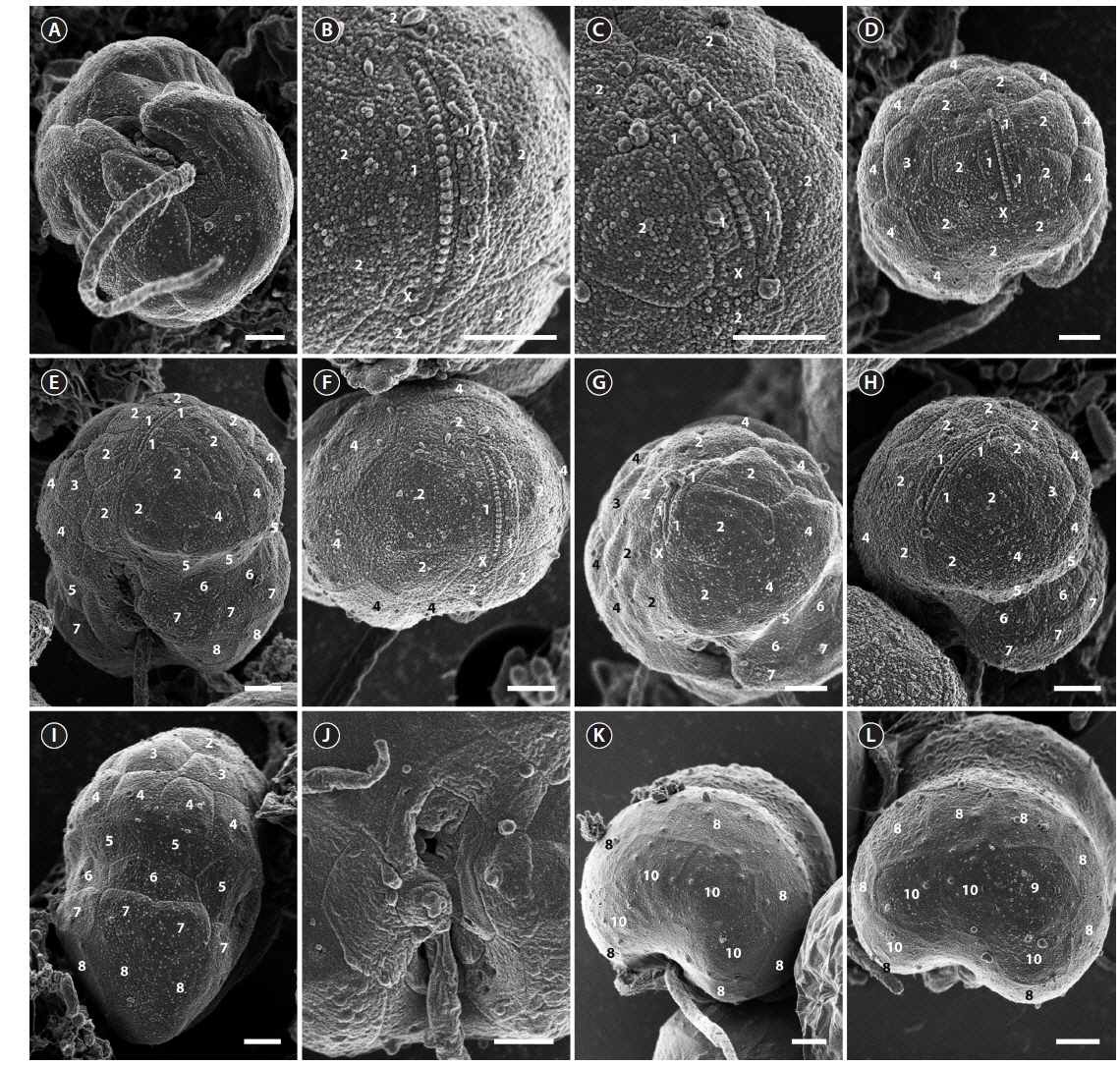
The cell surface of the isolate from Korea was ornamented with globular knobs and pores that were dispersed randomly (Fig. 2). A hypoconal flange was present, but not clearly expressed in all observed individuals (Fig. 2A). A single EAV ornamented with a row of globular knobs (Fig. 2B & C) was surrounded by a series of 3-4 quadrangular vesicles and a small squared vesicle (X vesicle) forming the apicals (series 1) (Fig. 2B & C). However, the apicals were constituted of 4 quadrangular vesicles only once. Another series of vesicles was posterior to the apicals and followed by an anterior intercalary and precingular series that were constituted of 5-8 (series 2), 0-3 (series 3), and 6-8 vesicles (series 4), respectively (Fig. 2D-I). The cingulum was wide, shallow and distinct, located in the median portion of the cell, descending, and displaced by approximately its own width (Fig. 2E). The cingulum contained two series of AVs (series 5 and 6) (Fig. 2E & I). The number of series in the cingulum was reduced to one when approaching the sulcus, particularly on the right side (Fig. 2E & I). While the sulcus could be deep and narrow, it was in most cases shallow and wide enabling most of the AVs that constituted it to be seen (Fig. 2J). The sulcus contained 13 AVs (Fig. 2J). The hypocone was composed of a series of postcingular small vesicles (series 7) (Fig. 2E & G-I), anterior to another series of 6-8 hypoconal vesicles (series 8) (Fig. 2K & L), and 3-4 antapical vesicles (series 10) (Fig. 2K & L). An intercalary vesicle was sometimes observed preceding the antapicals (series 9) (Fig. 2L).
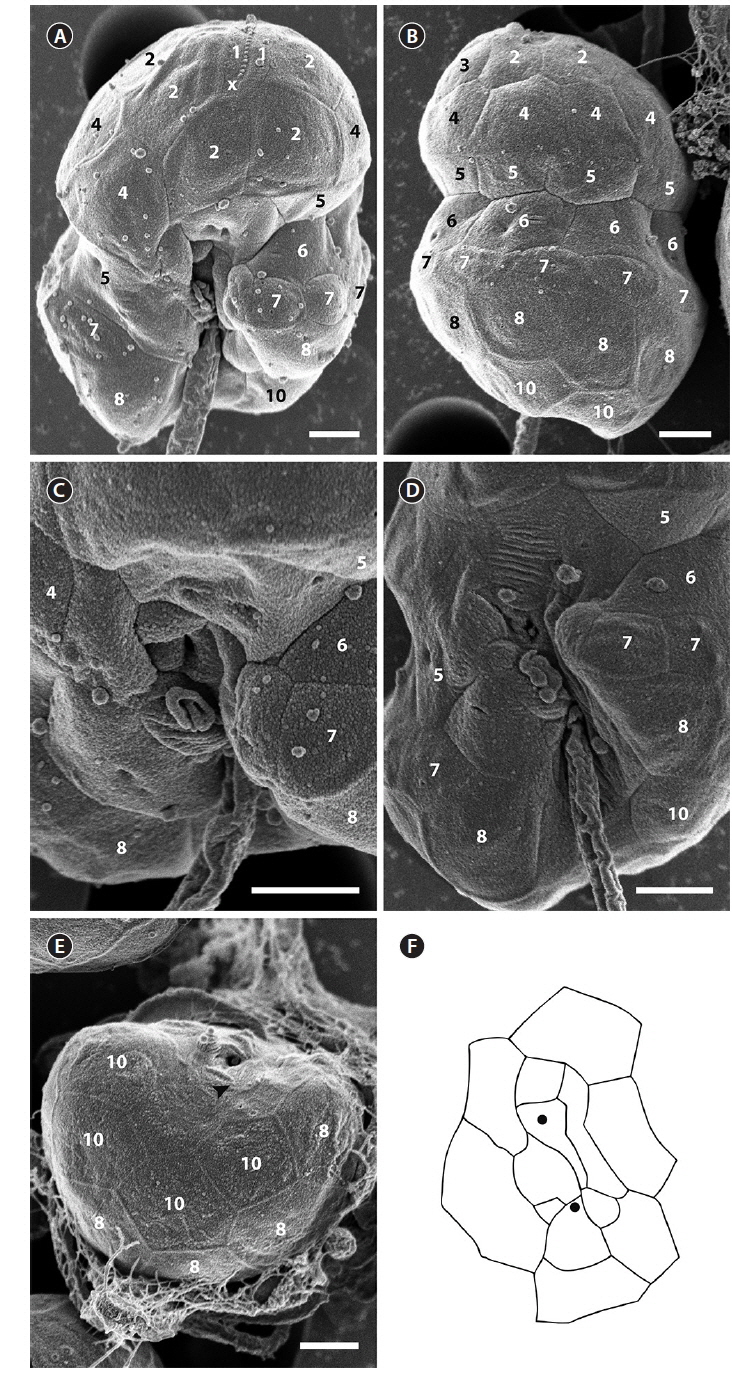
The SEM fixation of the type culture of
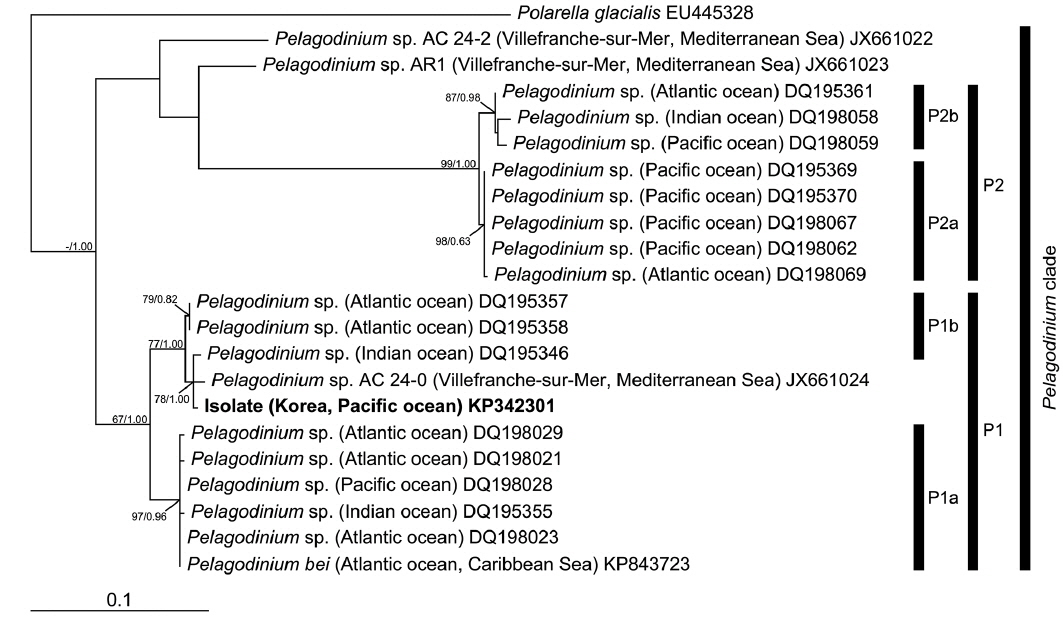
In the phylogenies based on ITS rDNA (Fig. 4) and LSU rDNA (Fig. 5), the isolate from Korea clustered with other sequences associated to the genus
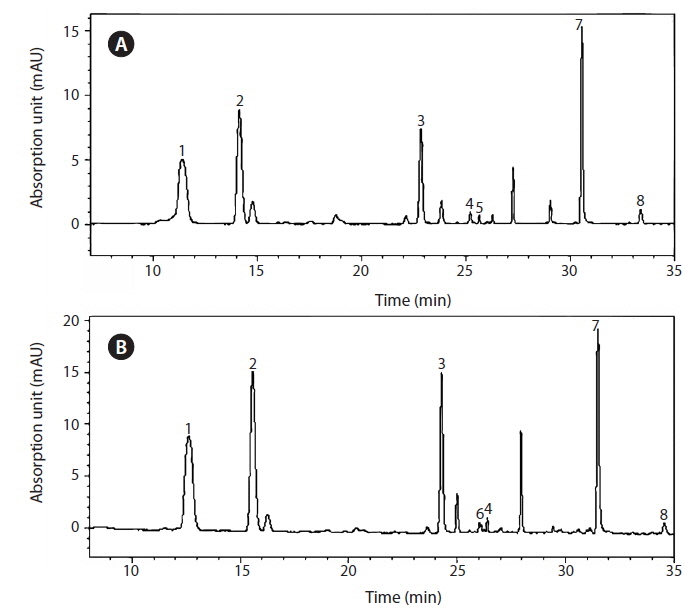
The isolate contained chlorophyll
The isolate usually swam fast in a straight line. It stopped quickly, changed direction at different angles, and backtracked repetitively. These behaviors were also observed previously for
The identification of dinoflagellates in the order Suessiales is difficult because many species in this order are small and fragile and SEM fixation does not always result in a clear distinction of the AVs. Thus, classification based on genetic characterizations has been used within several genera such as
This study is the first to report the detailed morphological, behavioral, and pigment characterization of a strain of
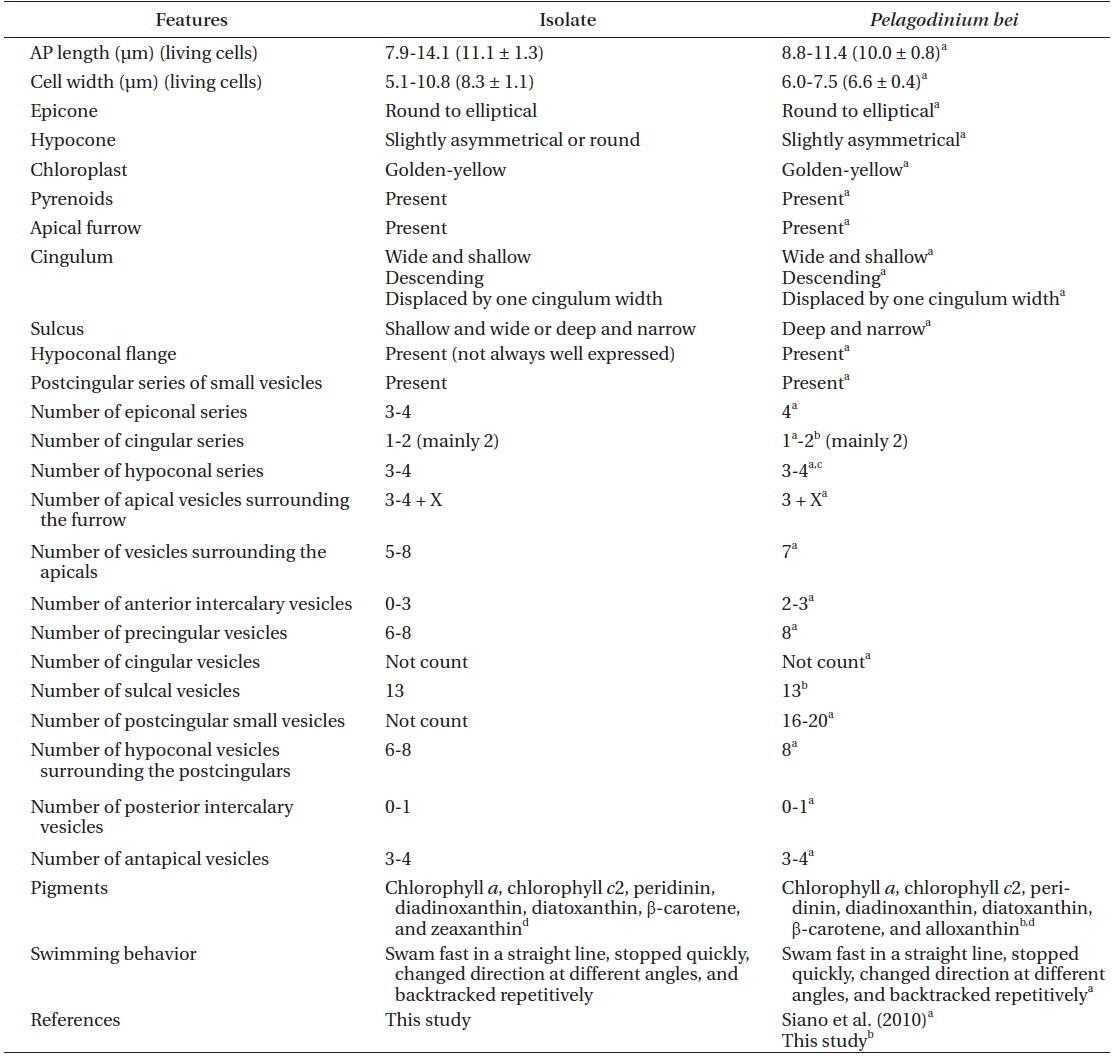
[Table 1.] Comparison between the isolate from Korea and Pelagodinium bei from the Caribbean Sea
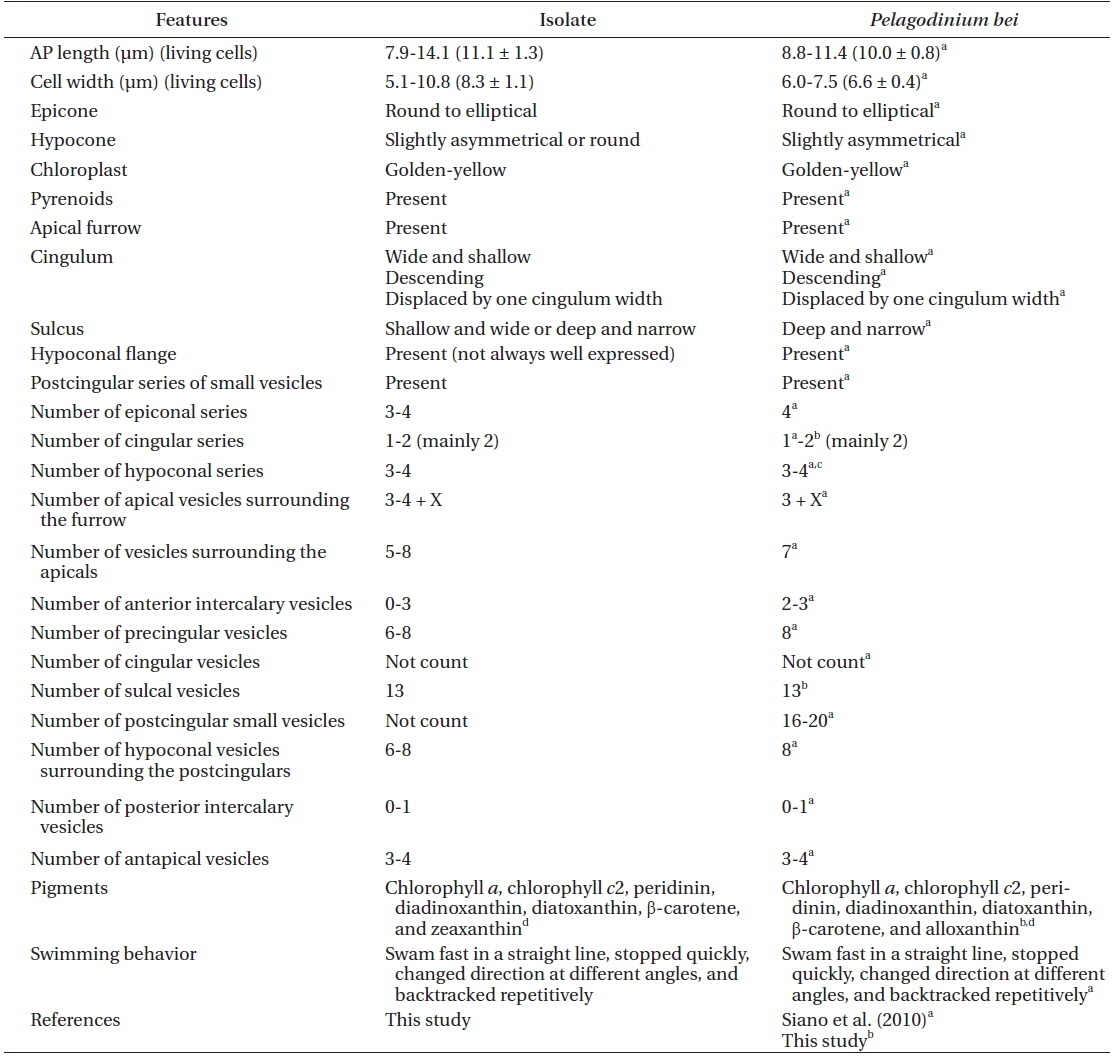
Comparison between the isolate from Korea and Pelagodinium bei from the Caribbean Sea
The observations of
The sulcal features were useful in determining the taxonomic identity within the same species since they are consistent between strains of
The swimming pattern is consistent between strains of
In conclusion, this study established detailed features of an isolate associated with the genus