Following a global trend of using natural products from insects, the Korean government has been encouraging the development of insect resources for environmental and dietary purposes, traditional medicine, and as pets (Kim et al., 2013; Lee et al., 2013; Suh and Kang, 2012; Yeo et al., 2013; Yoo et al., 2007; Youn et al., 2014). The order Coleoptera (beetles) contains edible insects that are used in Asian countries as an important food supplement because of their nutritional values and therapeutic effects. Among them, Protaetia brevitarsis seulensis Kolbe (Coleoptera: Cetoniidae) and Allomyrina dichotoma Linn. (Coleoptera: Scarabaeidae) are extensively farmed in Korea (Aronson et al., 1986; Choi et al., 2003; Jander et al., 2000; Lee et al., 1997). Several reports indicate that these beetles can be used as functional food because of their antioxidant (Suh and Kang, 2012; Suh et al., 2010), anti-hepatofibrotic, anti-cancer, and anti-diabetic properties (Koyama et al., 2006; Miyanoshita et al., 1996; Taketa et al., 1986; Yamada et al., 2004; Yeo et al., 2013; Yoo et al., 2007; Yoon et al., 2003; Youn et al., 2012), and in Korea, P. b. seulensis and A. dichotoma are grown in farms. The industrialization of insect breeding requires constant monitoring, identification, and prevention of insect diseases caused by various parasitic agents (Papp et al., 2014).
In this study, the incidence of insect diseases and causative pathogens in insect farms was surveyed. In general, insect sample collection was performed by overnight delivery of the diseased insects from the farms to our research laboratory. Our choice of the investigated pathogens was based on the survey of insect farms in Korea; as a result, six major insect pathogens were selected: fungi Beauveria bassiana (Bals.-Criv.) Vuill. (Hypocreales: Cordycipitaceae) and Metarhizium anisopliae (Metschn.) Sorokin (Hypocreales: Clavicipitaceae), bacteria Bacillus thuringiensis Berliner (Bacillales: Bacillaceae), Pseudomonas aeruginosa Migula (Pseudomonadales: Pseudomonadaceae), and Serratia marcescens Bizio (Enterobacteriales: Enterobacteriaceae), and Oryctes rhinoceros nudivirus. Beauveria bassiana is an anamorphic entomophogenic fungus that grows naturally in soils and causes white muscardine disease in various arthropod species; it is used as an effective mycoinsecticide to protect against insect pests (Meyling and Eilenberg, 2007). Metarhizium anisopliae is also known as an entomopathogenic fungus and is applied to control insect pests; Metarhizium anisopliae-extracted toxin, destruxin, is lethal to insects (Sharif, 2010; Smagghe et al., 2013). Bacillus thuringiensis is a widely represented entomopathogenic bacterium that produce crystal (Cry) toxins 1, 2, 4, 10, and 11 and is used as the first choice effective biological control against many kinds of insects from the orders Coleoptera, Lepidoptera, and Diptera (Ben-Dov, 2014; Bergamasco et al., 2013). Pseudomonas aeruginosa is reported as a virulent bacterial pathogen not only for insects but also for mammals (Jander et al., 2000), where it causes infection of the respiratory tract, especially in cystic fibrosis patients (Spiker et al., 2004). Similar to Pseudomonas aeruginosa, Serratia marcescens is known as an opportunistic bacterium (El-Aasar et al., 2013); it is also used to control apple maggot fly population and as a source of insecticidal agents such as toxin luxA (Hejazi and Falkiner, 1997; Lauzon et al., 2003). Oryctes rhinoceros virus Rhabdionvirus oryctes Hüger was first discovered in oil palms in Malaysia 40 years ago and is currently used for the control of rhinoceros beetle population in Malaysia (Huger, 2005; Ramle et al., 2005). We have identified this virus in an A. dichotoma-breeding farm in 2014 in Korea (Lee, 2015).
Extensive large-scale insect breeding has resulted in frequent occurrence of insect infection due to multiple pathogens. Here, we developed a multiplex PCR-based approach that allowed simultaneous detection of six insect pathogens using specific primer sets. Multiplex PCR-based methods have been previously applied for the detection of mixed infections in humans (Aguilera-Arreola et al., 2014; Ishaq and Wright, 2014). Thus, it has been recently shown that in humans, multiplex PCR can be effectively used for the detection of bacteria and fungi in blood (Gosiewski et al., 2014), influenza and parainfluenza viruses in the respiratory tract, and neurological viruses such as herpes simplex virus, cytomegalovirus, and Epstein-Barr virus (Kalvatchev et al., 2004). Multiplex PCR has been applied to detect food-associated bacterial pathogens that can infect the gastrointestinal tract, including Clostridium perfringens, Escherichia coli, Salmonella enterica, and Staphylococcus aureus (Gosiewski et al., 2014; Khare et al., 2014; Settanni and Corsetti, 2007), and thus, can represent an effective diagnostic method to be used for the prevention of foodborne infection spreading. The method has also been successfully applied to rapid simultaneous identification of pig viruses commonly occurring in pig farms (Haines et al., 2013; Liu et al., 2013; Monavari et al., 2014; Zeng et al., 2014). Furthermore, multiplex PCR has been used to detect infection in invertebrate species such as bee viral diseases (McNeil, 2012; Sguazza et al., 2013a). To rapidly perform simultaneous identification and quantification of infectious agents, the concentration of pathogen genomic DNA could be calculated based on fluorescence intensity in Multiplex Ligation-dependent Probe Amplification (MPLA), which is a variation of multiplex PCR that permits amplification of multiple targets with a single primer pair (Chung et al., 2014; Thomas et al., 2010a; Uno and Yanagihara, 2014). This system provides several advantages compared to real-time PCR. It takes a shorter time for product amplification and data analysis; in addition, the performance of real-time PCR is limited within a certain number of cycles providing linear increase in DNA amplification, and when the cycle number is further increased, false results can be expected (Freeman et al., 1999; Siebert, 1999), which can be avoided by using multiplex PCR. The QIAxcel Advanced system performs fully automated sensitive, high-resolution capillary electrophoresis of up to 96 samples per run (De Smet et al., 2012; Thomas et al., 2010a; Xiao et al., 2012); therefore, this system, which does not needed fluorescently labeled primers and uses one primer set, is more sensitive and effective for the detection of multiple pathogens compared to real-time PCR, and can be an alternative to conventional gel electrophoresis (Xiao et al., 2012). Based on previous research demonstrating the advantages of multiplex PCR and capillary electrophoresis for the detection of multiple infections in various systems, in this study, we applied these methods to successfully identify various insect pathogens, including fungi, bacteria, and a virus, which are known as common causes of infection in farm-raised A. dichotoma and P. b. seulensis in Korea.
Healthy larvae of P. b. seulensis and A. dichotoma used as experimental controls were raised in the laboratory of the National Agricultural Academy of Science in Wanju, Korea. P. b. seulensis and A. dichotoma larvae of three stages were reared in 35 × 25-cm cages (20 per cage) containing sawdust and water, at 25℃ and 60% humidity; their uninfected state was constantly monitored.
Fungal strains B. bassiana KACC40039 and M. anisopliae KACC40969, bacterial strains B. thuringiensis KACC10169, P. aeruginosa KACC10232, and S. marcescens KACC11892 were obtained from the Korean Agricultural Culture Collection, Rural Development Administration, Suwon, Korea. B. bassiana KACC40039 was cultured on potato dextrose agar (PDA; BD, Franklin Lakes, NJ, USA) and M. anisopliae KACC40969 was cultured on Sabouraud dextrose agar (SDA; BD) for 2 wk in an incubator at 24℃ . Three bacterial strains were grown on Nutrient Agar (NA; BD) at 24℃ . O. rhinoceros virus was isolated from virus-infected A. dichotoma raised in the insect farm in Youngdong, Korea; we assumed that it originated from coconut palm rhinoceros beetle O. rhinoceros (Lee et al., 2015).
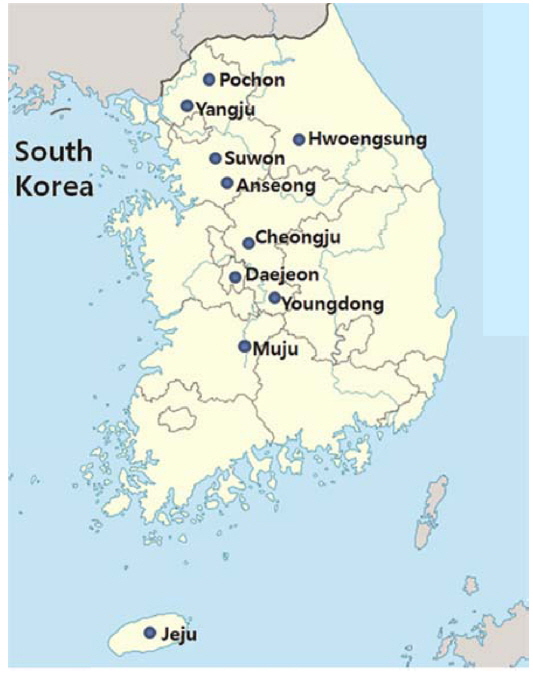
Farm-raised pathogen-infected P. b. seulensis was delivered from Hwoengsung, Yangju, Suwon, Cheongju, Jeonju, and Jeju, and A. dichotoma was obtained from the insect farms in Pochon, Anseong, and Youngdong (Fig. 1) Infected farm-raised A. dichotoma and P. b. seulensis were analyzed for the presence of insect pathogens frequently causing severe diseases in insects; these included Cordyceps spp., Enterobacter spp., Bacillus spp., Acinetobacter spp., Lecanicillium spp., and protozoa (data not shown), and B. bassiana, M. anisopliae, B. thuringiensis, P. aeruginosa, S. marcescens, and O. rhinoceros virus (Table 1).
Nucleotide sequences of all tested pathogens except B. thuringiensis were retrieved from GenBank and used to design specific primers with similar annealing temperatures using the MFEprimer 2.0 program (Gardner and Slezak, 2014); B. thuringiensis-specific primers were the same as previously reported (Yamada et al., 1999). The primers were synthesized by Macrogen (Seoul, Korea). Primer sequences and GenBank accession numbers of the target genes are shown in Table 2. The fungi, bacteria, and virus were isolated and identified by Nucleotide sequence of NCBI (National Center for Biotechnology Information). The accession numbers of GenBank were based on the nucleotide sequence alignment of classified fungi, bacteria and virus from diseased insect in Korea from 2013 to 2014. For primer design, the nucleotides sequence was compared using Basic Local Alignment Search Tool (BLAST) and aligned by ClustalW (Multiple Sequence Alignment, UK).
Fungal mycelia were ground in liquid nitrogen with a mortar and pestle, and genomic (g)DNA was extracted and purified using the DNeasy Plant Mini kit (Qiagen, Germantown, MD, USA). Bacterial and viral gDNA was extracted from the samples diluted in 500 μL distilled water using the Wizard Genomic DNA Purification kit (Promega, Madison, WI, USA). Healthy and pathogen-infected larvae were washed with distilled water, and 100 μL and 200 μL hemolymph per larva was obtained from P. b. seulensis and A. dichotoma, respectively. The hemolymph was then diluted 1/10 with distilled water and the extracted gDNA was used as a template for PCR amplification.
PCR was performed at the following cycling conditions: initial denaturation at 94℃ for 5 min, 35 cycles of denaturation at 94℃ for 30 s, annealing at 55℃ for 30 s, extension at 72℃ for 30 s, followed by final extension at 72℃ for 6 min. The reaction was carried out using the AccuPower multiplex PCR Premix kit (Bioneer, Korea) in a total volume of 25 μL containing gDNA (1 ng/μL) and primer pairs specific for B. bassiana (30 pM/μL), M. anisopliae (5 pM/μL), B. thuringiensis (200 pM/μL), P. aeruginosa (20 pM/μL), S. marcescens (20 pM/μL), and O. rhinoceros virus (20 pM/μL). First, single PCR detection of each of the six pathogens was performed. Then, multiplex PCR was carried out using different primer combinations: B. bassiana and M. anisopliae; B. thuringiensis, P. aeruginosa, and S. marcescens; B. bassiana, M. anisopliae, and O. rhinoceros virus. B. thuringiensis, P. aeruginosa, S. marcescens, and O. rhinoceros virus. PCR products were separated on 1% agarose gels in 1 × TEA buffer for 85 min and visualized using an UV trans-illuminator. Then, 10 μL of each PCR product was added to capillary tubes of the microfluidic QIAxcel capillary electrophoresis system (Qiagen, Valencia, CA, USA) and separated for 30 min at 6 kV. The resulting electropherograms and DNA patterns were analyzed using the QIAxcel software.
Using uniplex PCR with specific primer pairs, we were able to detect the following DNA bands: 577 bp (O. rhinoceros virus), 368 bp (B. bassiana), 269 bp (P. aeruginosa), 202 bp (S. marcescens), and 87 bp (M. anisopliae) (Fig. 2A). We also tested the following primer combinations: B. bassiana and M. anisopliae; B. thuringiensis, P. aeruginosa, and S. marcescens; B. bassiana, M. anisopliae, and O. rhinoceros virus; B. thuringiensis, P. aeruginosa, S. marcescens, and O. rhinoceros virus in multiplex PCR for simultaneous detection of multiple infectious agents. The results demonstrated the same band pattern obtained with uniplex PCR (Fig. 2B). These data indicate a sufficient sensitivity of multiplex PCR for the detection of several parasites in only 0.5 ng gDNA as a PCR template (Fig. 2).
To further evaluate the specificity of the designed primers, we used them for the analysis of a mixture of six parasite-derived gDNA samples. The results demonstrated the same band patterns in uniplex and multiplex PCR: each target product migrated according to the expected size (Fig. 3), confirming the sensitivity and specificity of the designed primers.
Next, the primers were applied for the detection of infectious agents in diseased P. b. seulensis and A. dichotoma larvae obtained from three areas in Korea. As shown in Fig. 4A, M. anisopliae (87 bp) and S. marcescens (144 bp) were detected in all P. b. seulensis samples and P. aeruginosa (202 bp) was identified in 50% of them as cross infection. We also found that three of four A. dichotoma samples from Youngdong were infected with O. rhinoceros virus (577 bp), and one of them was co-infected with M. anisopliae (87 bp); one of the four analyzed A. dichotoma samples was infected only with M. anisopliae (Fig. 4B). In healthy larvae, no amplification was observed, indicating the specificity of the primer set for the tested pathogens.
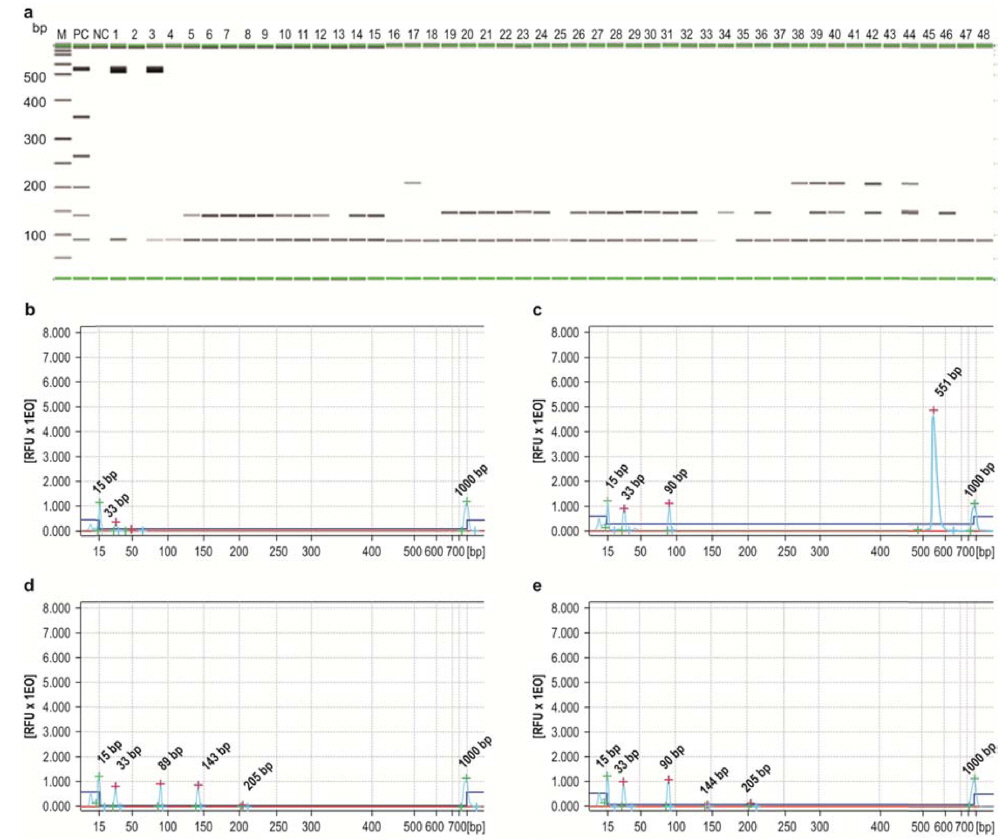
Quantification of multiplex PCR results was performed by capillary electrophoresis using the QIAxcel Advanced system, and the quantitative data (ng/μL) of multiplex PCR were integrated by the QIAxcel software (Fig. 5). The multiplex PCR products obtained from infected P. b. seulensis and A. dichotoma larvae were transferred to gel cartridges in the QIAxcel system and analyzed by capillary electrophoresis. The resulting data show the frequency of insect pathogens in diseased insects and infection incidence in local insect farms in Korea (Fig. 5). Most of P. b. seulensis samples from Yangju, Suwon, and Jeonju were co-infected with M. anisopliae and S. marcescens; two of the samples (lanes 13 and 14) were co-infected with M. anisopliae, S. marcescens, and P. aeruginosa (Fig. 5A). Although A. dichotoma larvae from Youngdong were co-infected with O. rhinoceros virus and M. anisopliae, the quantity of viral DNA suggests that A. dichotoma from Youngdong was likely to die from viral infection (Fig 5C). This notion should be confirmed by further bioassay experiments aimed to evaluate comparative toxicity of insect pathogens in order to determine the contribution of each pathogen to the mortality of cross-infected larvae. We are currently conducting the experiments to analyze A. dichotoma mortality as a result of O. rhinoceros virus infection.
As shown in Fig. 5, the majority of P. b. seulensis samples (90%) were infected with M. anisopliae, and most of them were co-infected with S. marcescens. In the diseased P. b. seulensis, total co-infection rate was about 45%.
In this study, we designed six pairs of PCR primers targeting different types of insect pathogens most frequently identified in the insect farms breeding P. b. seulensis and A. dichotoma in Korea. Fungi B. bassiana and M. anisopliae infect insects through the mouth and then grow on insect epidermis; we could see the color specific for fungal spores. For example, B. bassiana is shown as white and M. anisopliae as dark green with light black covering on insect epidermis, which could be qualified as the first sign of infection with these fungi. However, it is not so easy to detect symptoms caused by bacterial infection. In this case, PCR performed on freshly dead insects can accurately and rapidly diagnose the parasite, as was the case with the bacterial pathogens S. marcescens and P. aeruginosa known to cause severe infection if injected in the hemocoel of Orthoptera, Coleoptera, Hymenoptera, Lepidoptera, and Diptera (Tanada et al., 1996). Thus, PCR-based detection can provide early diagnosis of insect diseases and prevent their spreading, because the delay in diagnosis and advanced symptoms could promote systemic infection, which would not be possible to properly control. Oryctes rhinoceros nudivirus was confirmed to be the cause of almost all viral diseases in Korean beetle farms in 2013–2014 (Lee et al., 2015). Therefore, in the present study, we designed primer sets specific to the most frequent causative agents of severe insect diseases, which enabled us to identify the stages of insect disease, the degree of sawdust contamination, and the pathogens responsible for the disease of beetles bred in insect farms in Korea.
In our study, using multiplex PCR, we demonstrated that P. b. seulensis was frequently infected with S. marcescens and co-infected with M. anisopliae in more than 80% of cases, indicating that such an analysis can be useful for pathogen identification, especially if different pathogens produce similar symptoms. For example, pathogenic bacteria Paenibacillus popilliae that cause milky disease in A. dichotoma larvae and the entomopathogenic fungus Nomuraea rileyi that appear in light yellow-green color may be confused with B. bassiana and M. anisopliae, respectively (Tanada et al., 1996). Viruses such as nuclear polyhedrosis virus and cytoplasmic polyhedrosis virus that often infect Bombyx mori silkworm can also attack Coleoptera spp. (Tanada et al., 1996); therefore, the primers specific for these viruses should be added to the panel of multiplex PCR primers for the identification of infectious agents in farmed beetles. Primer sets for simultaneous diagnostics of insect pathogens by multiplex PCR should be developed to efficiently control insect infection and prevent its further spreading. Our study shows that to minimize the damage to insect farming caused by pathogenic microorganisms, multiplex PCR-based detection and capillary electrophoresis-based quantification may be an effective and rapid approach to identify and prevent spreading of insect disease and massive insect death. Also, rapid and sensitive diagnosis of insect disease is necessary to promote further industrialization of insect farming and to obtain insect-derived biologically active substances. Compared to single PCR, multiplex PCR demonstrates advantages as a fast and cost-effective method for simultaneous detection of several insect pathogens; however, multiplex PCR has limitations when quantitative evaluation of insect co-infection is required or the contribution of each pathogen to disease progress should be assessed (De Smet et al., 2012; Khare et al., 2014; Romero-Pastrana, 2012; Sguazza et al., 2013b). In such cases, capillary electrophoresis can be used as a sensitive method to quantitatively analyze and compare the presence of each target pathogen gene (Barakat et al., 2014; Chung et al., 2012; Shin et al., 2014; Shin et al., 2010). Capillary electrophoresis can be successfully applied in cases of insect cross infection with several pathogens to quantitatively evaluate relative importance of infectious agents based on the ratio of target gene expression (Thomas et al., 2010b);