The family Bramidae in the order Perciformes comprises 7 genera and 22 species worldwide (Nelson, 2006), 6 genera and 10 species in Japan (Hatooka and Kai, 2013), and 4 genera and 6 species in Korea (Kim, 2011; Kim et al., 2012; Lee et al., 2014). The 6 species in Korea are
We collected 10 specimens that looked like
Ten specimens of
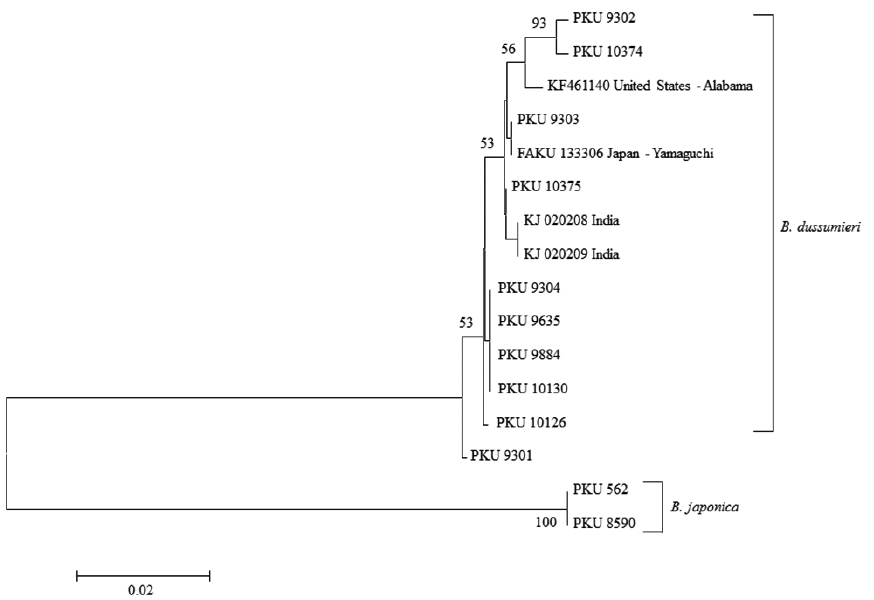
Genomic DNA was extracted from muscle tissue using Chelex 100 resin (Bio-Rad, USA), and polymerase chain reaction (PCR) was conducted using universal primer sets VF2 (5′-TCAACCAACCACAAAGACATTGGCAC-3′) and FishR2 (5′-ACTTCAGGGTGACCGAAGAATCAGAA-3′), which amplify mitochondrial DNA (mtDNA) cytochrome c oxidase subunit I (COI) (Ward et al., 2005; Ivanova et al., 2007). The PCR was performed on a total volume of 30 μL of sample, containing 5 μL DNA template, 4 μL dNTP, 5 μL 10X buffer, 0.5 μL Taq polymerase, 1 μL reverse primer, 1 μL forward primer, and distilled water. The PCR was conducted under the following conditions: initial denaturation for 1 min at 95°C, followed by 35 cycles of 1 min at 95°C for denaturation, 1 min at 50°C for annealing, and 1 min at 72°C for extension, with a final extension for 5 min at 72°C. The PCR products were purified using ExoSAP-IT (Biochemical Co., USA) and sequenced using an ABI PRISM Big Dye Terminator v3.1 Ready Reaction Cycle Sequence Kit (Applied Biosystems, USA) on an ABI3730xi DNA Analyzer (Applied Biosystems). The sequence was aligned using ClustalW (Thompson et al., 1994) in BioEdit (ver. 7) (Hall, 1999). A neighbor-joining (NJ) tree (Saitou and Nei, 1987) was constructed using the Kimura two-parameter model (Kimura, 1980) in MEGA 5 (Tamura et al., 2011).
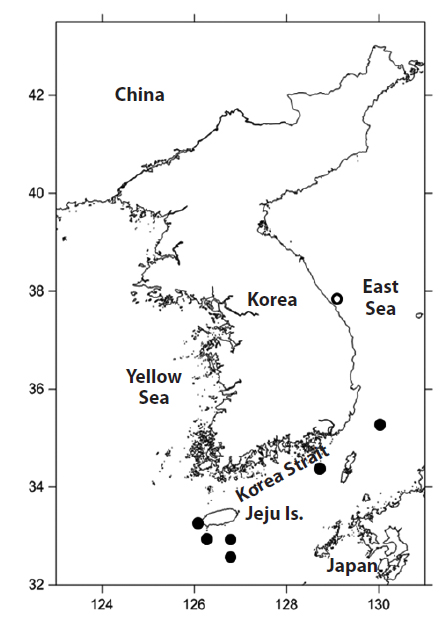
Material examined: PKU 9301 and 9302, 2 specimens, 102.1-153.6 mm standard length (SL), Seogwipo-si, Jeju Island, Korea, 4 July 2013; PKU 9303 and 9304, 2 specimens, 148.7-157.1 mm SL, Seogwipo-si, Jeju Island, Korea, 8 July 2013; PKU 9635, 1 specimen, 162.1 mm SL, Seogwipo-si, Jeju Island, Korea, 13 August 2013; PKU 9884, 1 specimen, 181.1 mm SL, Gangneung-si, Gangwon-do, Korea, 25 September 2013; PKU 10126, 1 specimen, 199.9 mm SL, Geoje-do, Gyeongsangnam-do, Korea, 26 December 2013; PKU 10130, 1 specimen, 200 mm SL, Busan, Korea, 2 January 2014; PKU 10374 and 10375, 2 specimens, 191.3–192.0 mm SL, Seogwipo-si, Jeju Island, Korea, 12 March 2014.
Comparative material examined:
>
Brama dussumieri
(New Korean name: Wae-sae-da-rae) (Fig. 2)
Dorsal fin rays, 32-34; pectoral fin rays, 19-20; anal fin rays, 26-29; lateral line scales, 58-64; gill rakers, 13-15; vertebrae, 39-42 (Table 1).
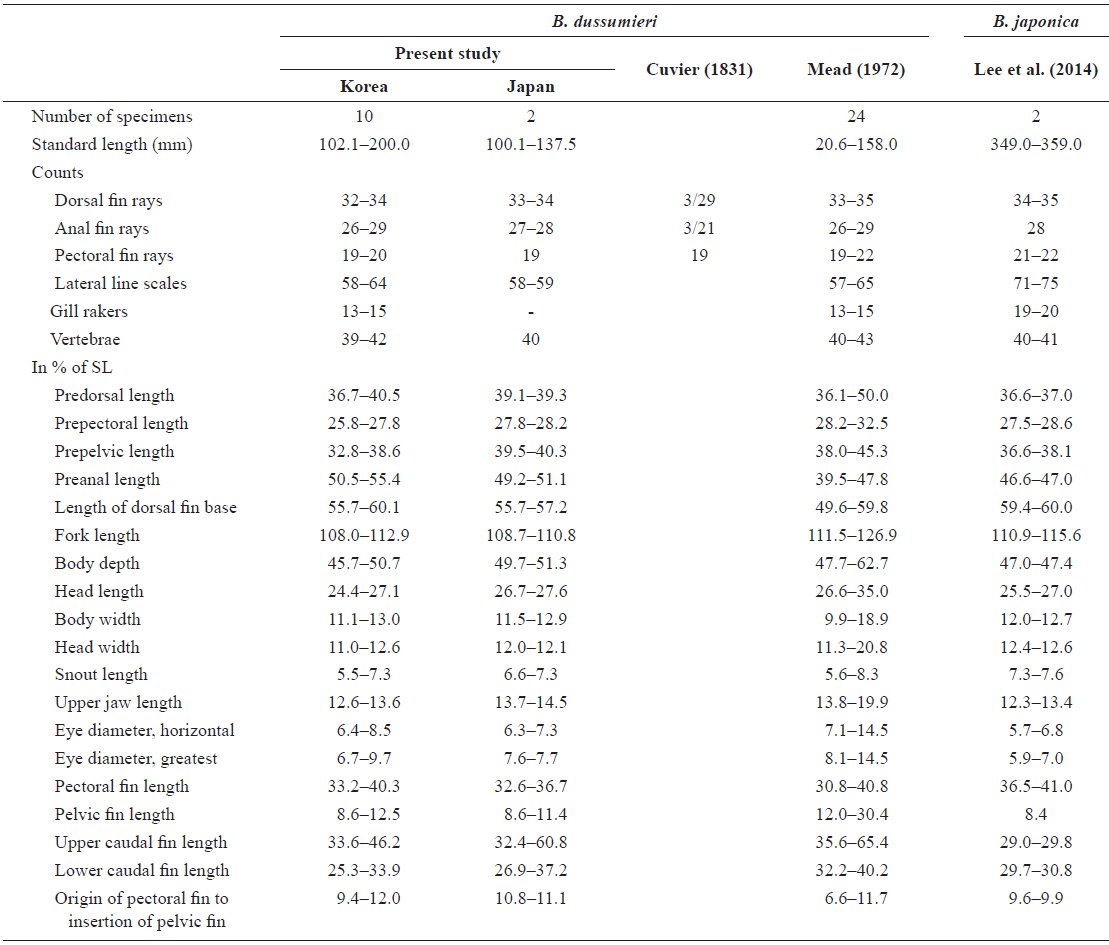
[Table 1.] Comparison of counts and measurements for Brama dussumieri and Brama japonica
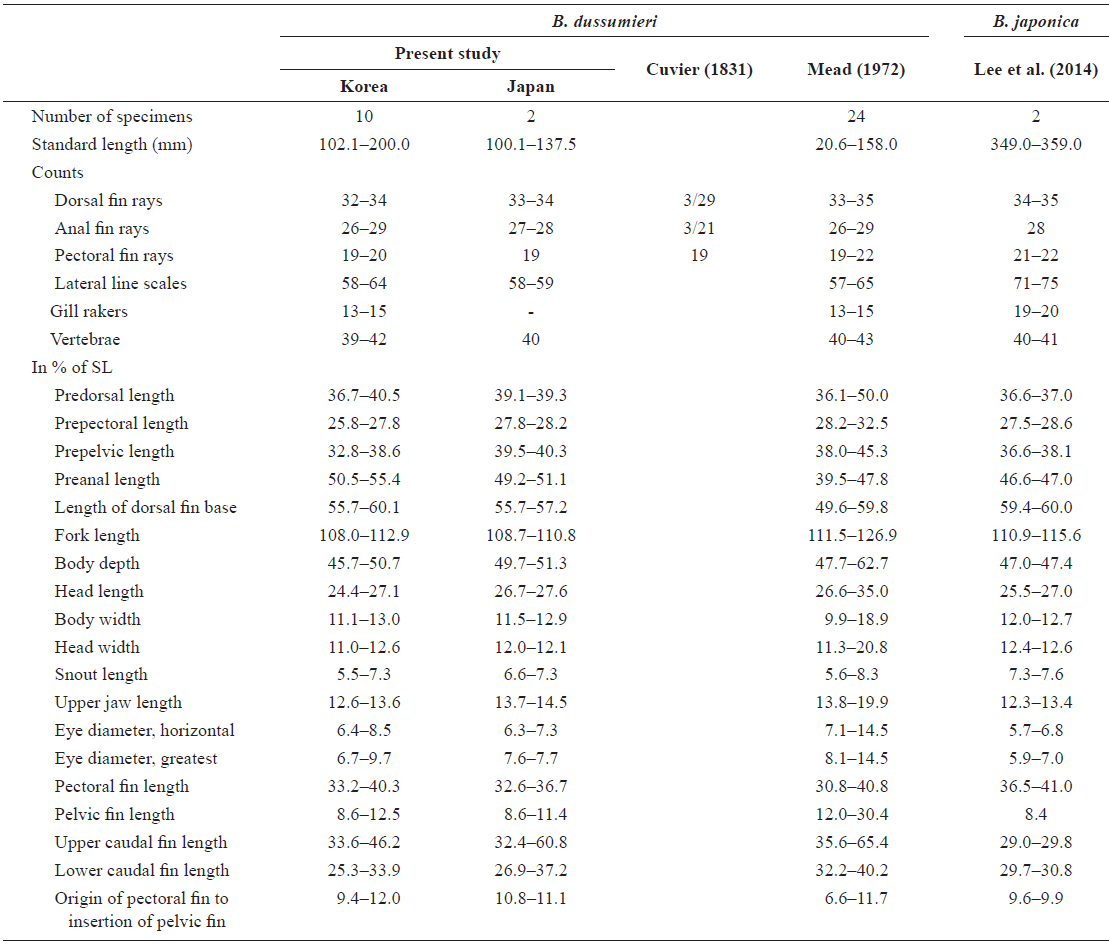
Comparison of counts and measurements for Brama dussumieri and Brama japonica
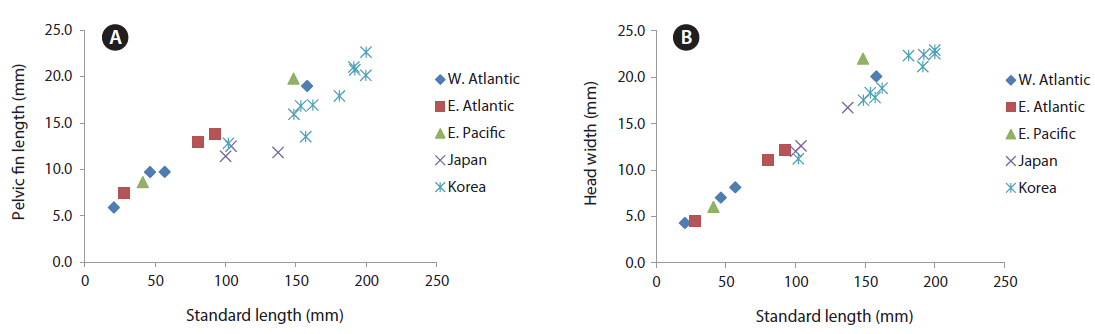
Proportion of measurements as a percentage of SL: head length 24.4-27.1; predorsal length 36.7-40.5; length of dorsal fin base 55.7-60.1; snout to pectoral fin 25.8-27.8; snout to pelvic fin 32.8-38.6; snout to anal fin 50.5-55.4; fork length 108.0-112.9; body depth 45.7-50.7; body width 11.1-13.0; upper caudal fin length 33.6-46.2; lower caudal fin length, 25.3-33.9
Body ovate, strongly compressed; head large (24.4-27.1% SL), forehead feebly arched; snout short (5.5-7.3% SL), slightly blunt, mouth strongly oblique; interorbital region convex, anterior nostril round and located at anterior tip of snout, posterior nostril is a slit midway between anterior nostril and orbit; eye elliptical in shape, with horizontal axis shorter than vertical axis; branchiostegal ray number, 7; membrane completely covered by opercular flap; posterior margin of upper jaw extending slightly beyond line vertical to the middle of the pupil; upper jaw protruding more than lower jaw; one row of small conical teeth in upper jaw, one row or two rows of teeth in lower jaw, some mid-section teeth markedly large and recurved; dorsal fin originating slightly posterior to base of pectoral fin, reaching anterior caudal peduncle; pectoral fin starts under edge of opercle and reaches to mid body; pelvic fin short, originating on a vertical line slightly posterior to the origin of the pectoral fin; caudal fin strongly forked, length of upper lobe (32.3-46.2% SL) longer than that of lower lobe (25.3-33.9% SL); body and head mostly covered with oblong ctenoid scales; dorsal and anal fins covered with large oblong ctenoid scales.
Tropical regions of all oceans (Mead, 1972), Ryukyu Is. and Ogasawara Is., Japan (Hatooka and Kai, 2013), and waters off Jeju Island, Busan, and Gangneung, Korea (present study).
The specimens in the present study were identified as belonging to the genus
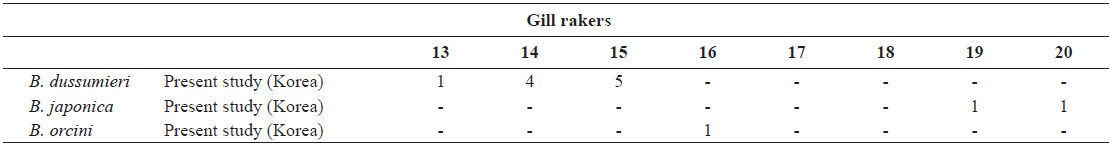
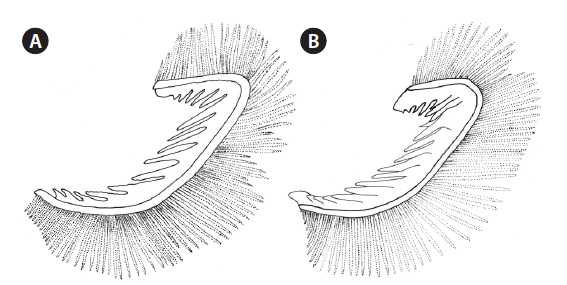
[Table 2.] Number of gill rakers in Brama dussumieri, Brama japonica, and Brama orcini
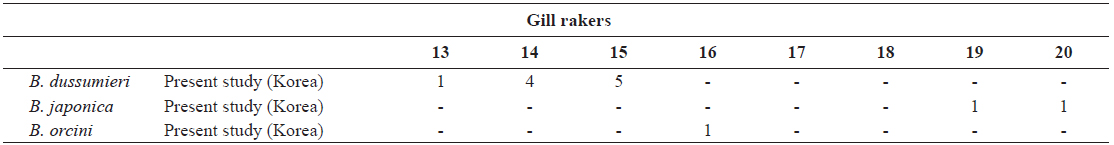

Number of gill rakers in Brama dussumieri, Brama japonica, and Brama orcini
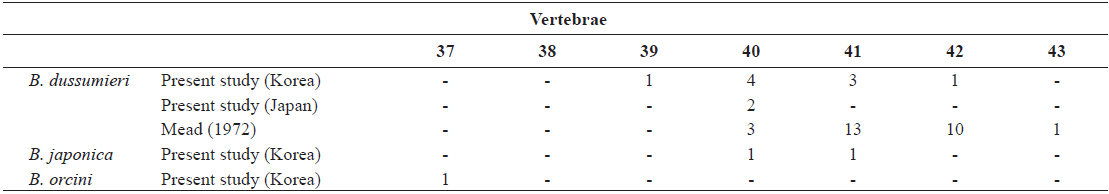
[Table 3.] Number of vertebrae in Brama dussumieri, Brama japonica, and Brama orcini
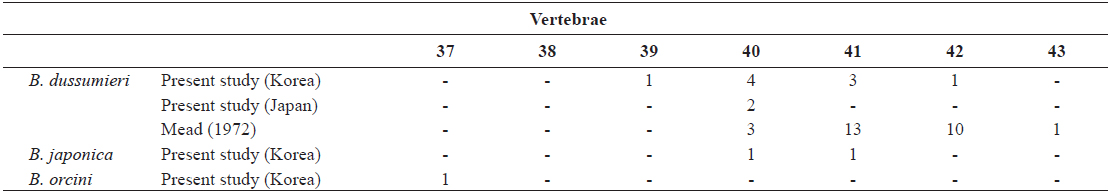

Number of vertebrae in Brama dussumieri, Brama japonica, and Brama orcini
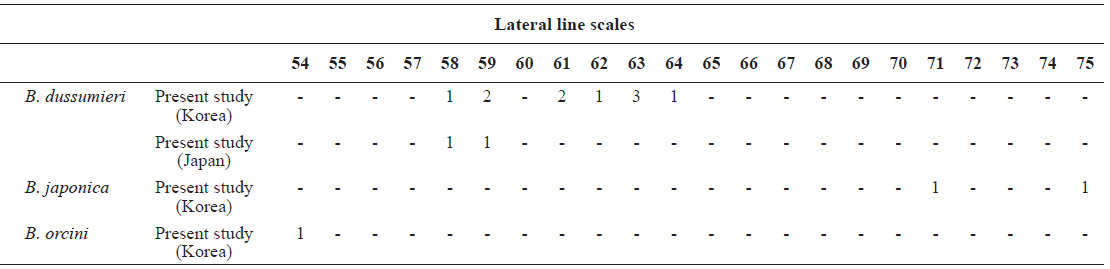
[Table 4.] Number of lateral line scales in Brama dussumieri, Brama japonica, and Brama orcini
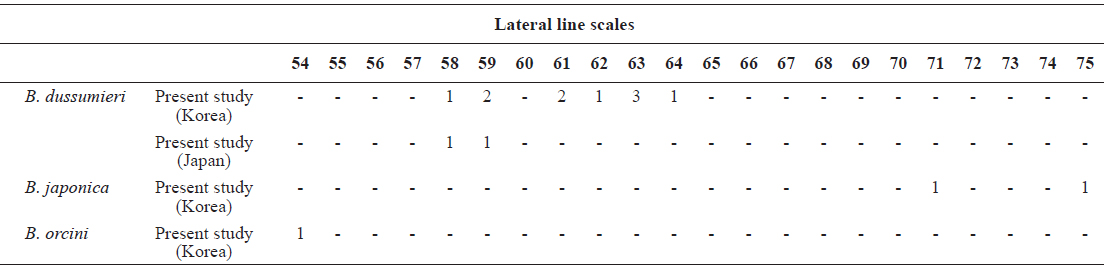
Number of lateral line scales in Brama dussumieri, Brama japonica, and Brama orcini