Several transgenic marine medaka Oryzias dancena strains harboring a green fluorescent protein (GFP) reporter construct regulated by an endogenous β-actin promoter were established and their expression characteristics in relation to transgene copy numbers were examined in 21 transgene genotypes. Most of the transgenic strains displayed transgene insertion patterns typical of microinjection-mediated introduction of foreign DNA into fish embryos, characterized by the random integration of multiple transgene copies (ranging from 1 – 282 copies per cell), often accompanied by the formation of concatemer(s), as assessed by genomic Southern blot hybridization analysis and qPCR. Transgenic strains showed ubiquitous and continued temporal and spatial expression patterns of the transgenic GFP during most of their life cycle, from the embryonic stage to adulthood, enabling assessment of the expression pattern of the endogenous β-actin gene. However, a comparative evaluation of transgene copy numbers and expression levels showed that copy number-dependent expression, the stability of the ubiquitous distribution and expression efficiency per transgene copy varied among the transgenic strains. Fluorescence expression levels were positively correlated with absolute transgene copy numbers, whereas the expression efficiency per transgene copy was inversely related to the number of transgene integrant copies. Data from this study will guide the selection of potentially desirable transgenic strains with ubiquitous expression of a fluorescent transgene, not only in this marine medaka species but also in other related model fish species.
Experimental transgenesis in a laboratory fish model has received much attention as a potential means to study the function of vertebrate genetics, and a variety of transgenic fish strains have been investigated with particular focus on developmental genetics and functional genomics (Chen et al., 1996; Gong et al., 2001; Rembold et al., 2006). Transgenic strains that acquire the ability to express a visible reporter throughout their body in a constitutive and ubiquitous manner throughout their life cycle provide an invaluable resource for research into
Postmortem studies on the induction of ubiquitous transgenic expression in fish have reported a frequent loss of transgenicity (
The euryhaline medaka species
In this study, experimental transgenesis of the
>
Transgene construct and experimental transgenesis
The transgene was constructed by fusing the marine medaka
>
Southern blot hybridization analysis and qPCR-based estimation of transgene copy-numbers
Five micrograms of
To estimate the transgene copy number per cell for each transgenic line, a quantitative PCR assay was conducted using 5 ng of genomic DNA (measured spectrophotometrically) from each individual. To ensure that an equal amount of DNA was used across all samples, each template sample was pre-confirmed as having almost identical Ct values (± 0.4) based on the qPCR amplification of the endogenous actin gene (GenBank accession no. HM347346) segments (data not shown). Thermal cycling and detection of the fluorescent signal of the amplification products were carried out using 2 × iQ™ SYBR® Green Supermix and Optic Module from the iCycler® iQ™ Real-Time Detection System (Bio-Rad, Hercules, CA, USA) according to the manufacturer’s recommendations under the following cycling conditions: 45 cycles at 94℃ for 15 s, 60℃ for 15 s and 72℃ for 15 s with an initial denaturation step at 94℃ for 2 min. The PCR primers were qGFP-F (5ʹ-AACGAGAAGCGCGATCACAT-3ʹ) and qGFP-R (5ʹ-TACCGTCGACTGCAGAATTC-3ʹ). The resulting amplicon was 130 bp. Based on standard curves prepared using known copies of positive plasmids (podβ-actGFP), the PCR efficiency for each amplification reaction was confirmed to range from 97.1 – 101.6%. From preliminary tests to determine the appropriate ranges of the standard copy numbers, two different standard curves scales were applied (0 – 200 copies for low-copy-number transgenic lines and 0 – 2,500 copies for high-copy-number transgenic lines) to ensure optimum resolution. For each transgenic genotype, at least four individuals from the F1 and F2 generations were tested, and three independent experiments were carried out per individual to calculate the average copy number.
>
GFP image analysis and qRT-PCR assay of gfp transcripts
The GFP signals were measured using the MetaVueTM Research Imaging System (Molecular Devices Corp., Downingtown, PA, USA) and NIS-Elements BR image analysis software (ver. 3.1) equipped in an AZ100 fluorescence microscope system (Nikon Corporation Instruments Company, Japan) with a Nikon GFP-B-2A filter (450–490 nm excitation; dichromatic mirror cut-on at 500 nm and a barrier filter at 515 nm). The arbitrary values (statistical mean of intensity values of the pixels) relative to the serial dilutions of a standard purified GFP protein (Clontech Laboratories Inc.) were measured using the image analysis software to assess the relative strength of GFP expression across the transgenic strains. Living GFP-transgenic strains (six specimens per transgene genotype; 1-week-old fry) were subjected to fluorescence microscopy analysis. To examine the distribution pattern of transgenic GFP expression in adult tissues, somatic (brain, eye, fin, gill, heart, intestine, kidney, liver, skeletal muscle, spleen) and gonadic (ovary or testis) tissues were obtained from F1 and/or F2 individuals (
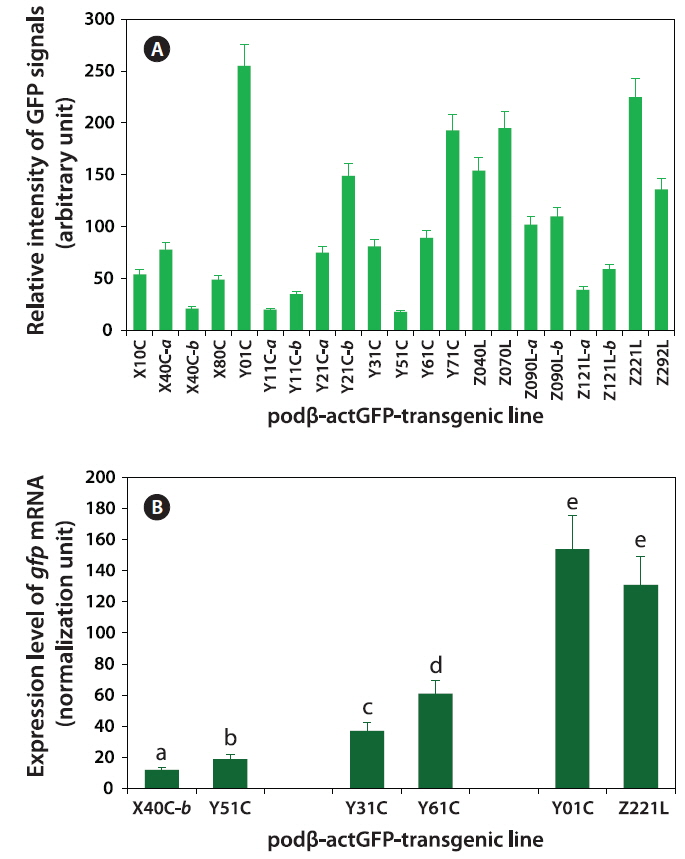
To ascertain whether the GFP expression levels measured using fluorescence microscopy were positively correlated with the transcriptional levels of the
>
Generation of transgenic germlines
Following multiple microinjection trials, 21 and 14 GFP-positive founder fishes were selected at adulthood from embryo batches injected with the circular and
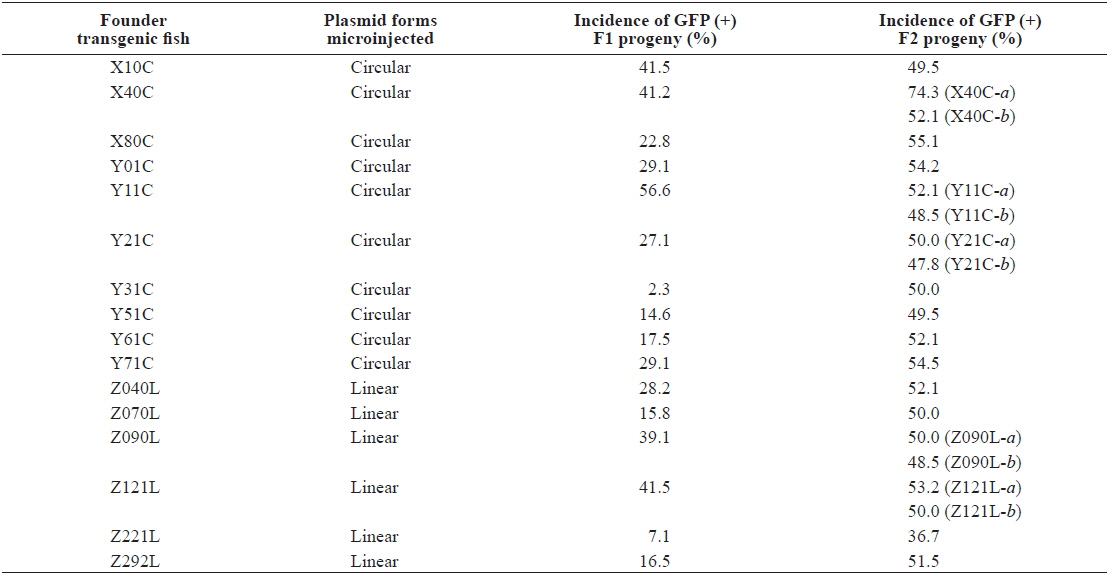
Percent germline transmission frequency of podβ-actGFP transgene from founder to F1 and F1 to F2 offspring
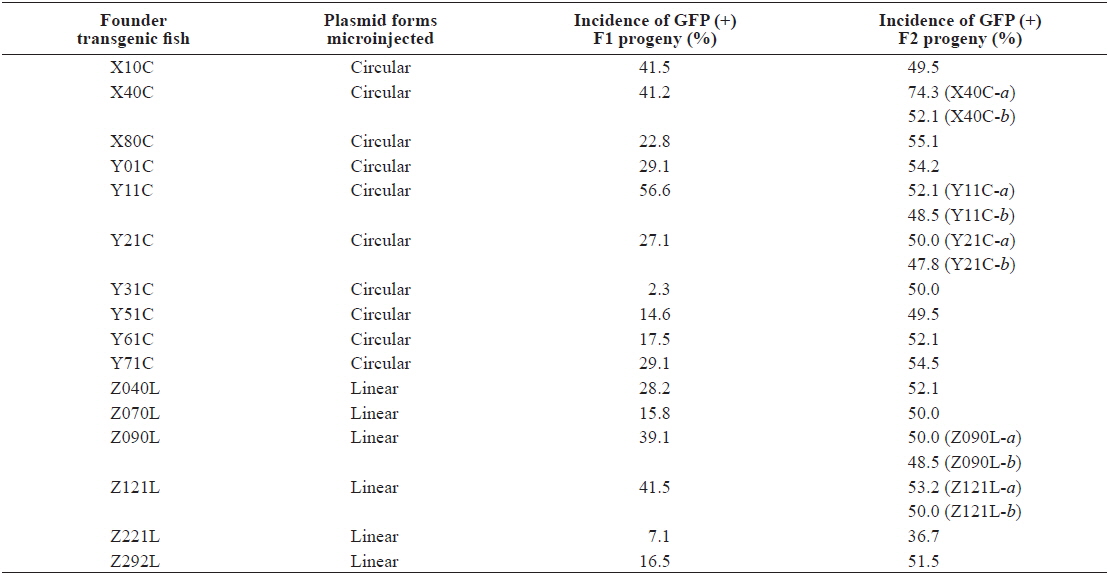
Although high variability of the germline transmission frequency was common in the passage from F0 to F1, most, but not all, transgenic F1 fishes transmitted the fluorescent transgene to the F2 generation following Mendelian inheritance patterns. Moreover, the GFP (-) but PCR (+) transgenic strains, present in a few strains in the F1 generation, were no longer detected in the F2 generation (data not shown), suggesting that hemizygous transgenic genotypes were successfully established in these transgenic lines (Nam et al., 2000). However, unlike these stable transgenic strains, two exceptional transgenic lines, X40C-
>
Genomic integration and transgene copy numbers
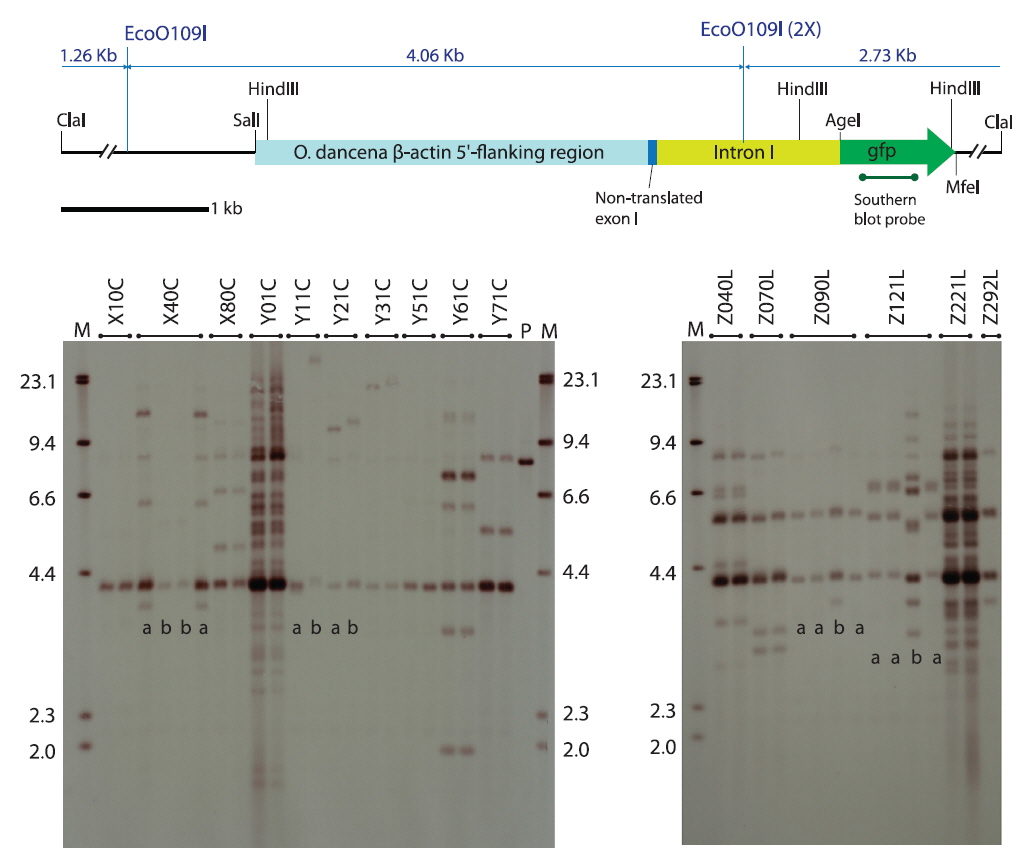
Southern blot hybridization analysis did not reveal any notable hybridization signals from the non-transgenic control fishes under the present hybridization conditions (data not shown). Transgenic strains carrying podβ-actGFP revealed various hybridization patterns in the transgenic lines, indicating independent and random integration of multiple copies of the microinjected DNA in most of the transgenic groups (Fig. 1), consistent with previous microinjection experiments in fish embryos (Hackett and Alvarez, 2000; Kim et al., 2004; Cho et al., 2013a). In the transgenic strains developed by microinjection with
As evidenced by the Southern blot hybridization analysis, five transgenic founders passed on each of two different transgene genotypes to their F1 offspring. These were X40C, Y11C and Y21C from the circular transgene-injected groups and Z090L and Z121L from the linear transgene-injected groups (see blots in Fig. 1). For both the X40C and Z121L lineages, the two transgenic genotypes detected among the F1 fishes within a given family differed greatly from one another in terms of hybridization pattern and transgene copy number (labeled
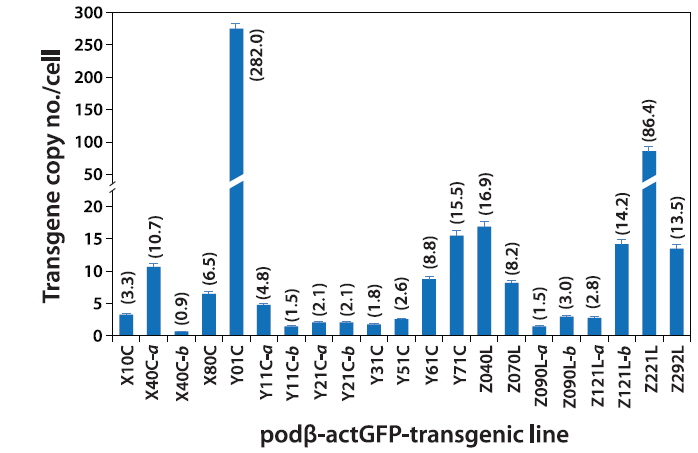
Based on the qPCR assay, the average transgene copy number per cell varied greatly among the transgenic lines, ranging from 0.9 (almost a single copy; transgenic line X40C-
>
Expression characteristics of the GFP transgene among the transgenic lines
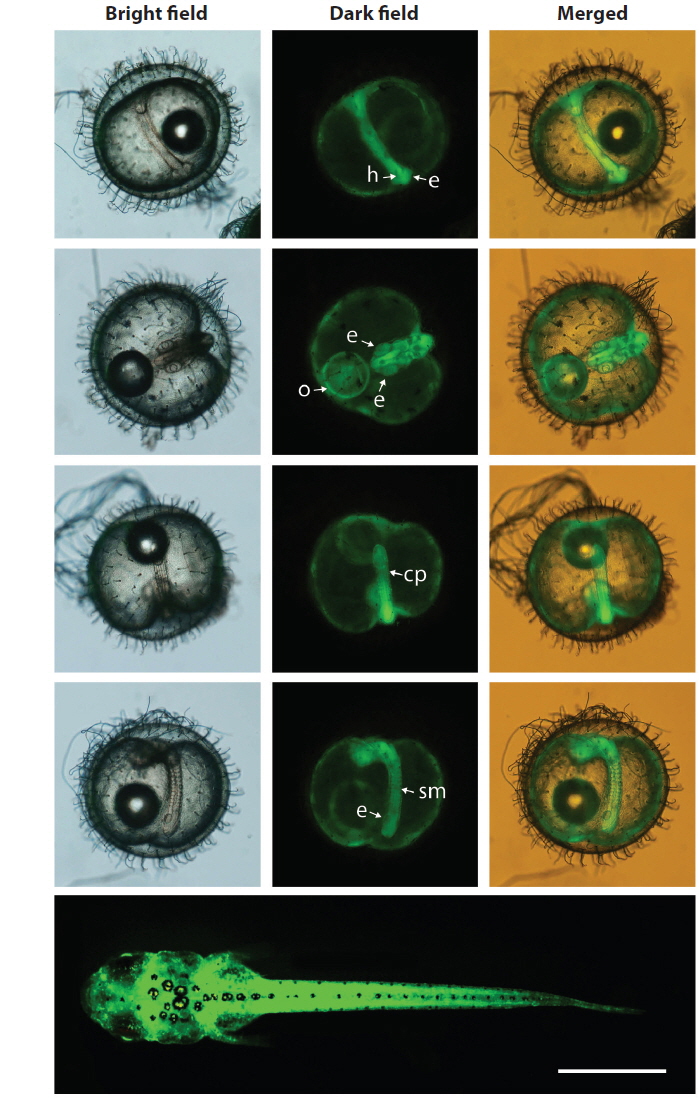
During embryonic development, spatial and temporal patterns of transgenic GFP expression were identified in all of the transgenic lines tested. The expression pattern of the GFP signals could be characterized at the onset of expression at the neurula stage, which intensified at the somite formation stage and was ubiquitous throughout the embryonic body, including the head, body trunk, eye and caudal peduncle, although the intensity of the GFP signal varied among the transgenic lines. This ubiquitous pattern was persistent in hatchlings that showed whole body expression of the GFP signals. Representative images showing the ubiquitous distribution of the GFP signals, exemplified by the Y71C transgenic line are shown in Fig. 3. The developmental expression pattern of the GFP reporter in this study was in agreement with previous observations of the expression of the endogenous
Fluorescence microscopic image analysis revealed that levels of GFP signals in the transgenic fry varied across the transgenic lines, with the difference between the weakest and strongest GFP expression being ~14-fold (Fig. 4A). Usually, the transgenic strains belonging to groups with robust GFP expression were easily distinguishable by eye from their non-transgenic siblings. However, the microscopic assay in this study revealed the external GFP expression in only an indirect way, and instead, this assay should include the mismatch of GFP quantification between GFP-expressing living organism and purified GFP protein in solution. For this reason, further quantification to assess the actual amount of GFP protein (
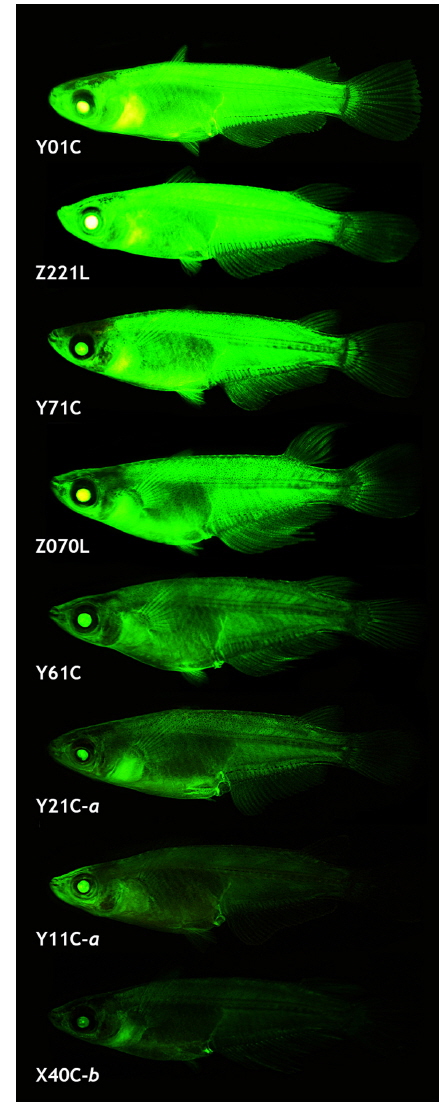
The transgenic expression pattern observed in the early stages generally persisted during growth to sexual maturity, although several transgenic lines lost GFP expression in certain tissues, similar to previous reports on transgenic strains with ubiquitous expression (Gibbs and Schmale, 2000; Burket et al., 2008; Er-meng et al., 2010). Of the 21 transgenic strains, 6 lost the GFP signals in one or more tissues, while the remaining 15 successfully retained the GFP phenotype in a ubiquitous manner. The number and type of GFP-negative organs observed in the six transgenic strains were not uniform: the transgenic lines failed to express GFP in 1 – 4 organs (the brain, eye, fin, heart, kidney, liver, spleen and/or testis). Although GFP expression was not quantified among the tissues, microscopic observations revealed that the tissue-dependent expression pattern of the transgenic GFP was possibly not homogeneous for all the transgenic lines (data not shown). Although tissue/organ-dependent expression levels were difficult to assign to a specific or uniform pattern, the external appearance of the GFP phenotype in the adult transgenic strains was in agreement with the GFP signals measured in the microscopic assays (Fig. 5). Under identical exposure conditions, the transgenic adults belonging to either the Y01C or Z221L strains always displayed strong GFP expression, which was barely seen in the other transgenic lines. On the other hand, transgenic lines—such as Y11C-
>
The relationship between transgene copy number and GFP expression levels
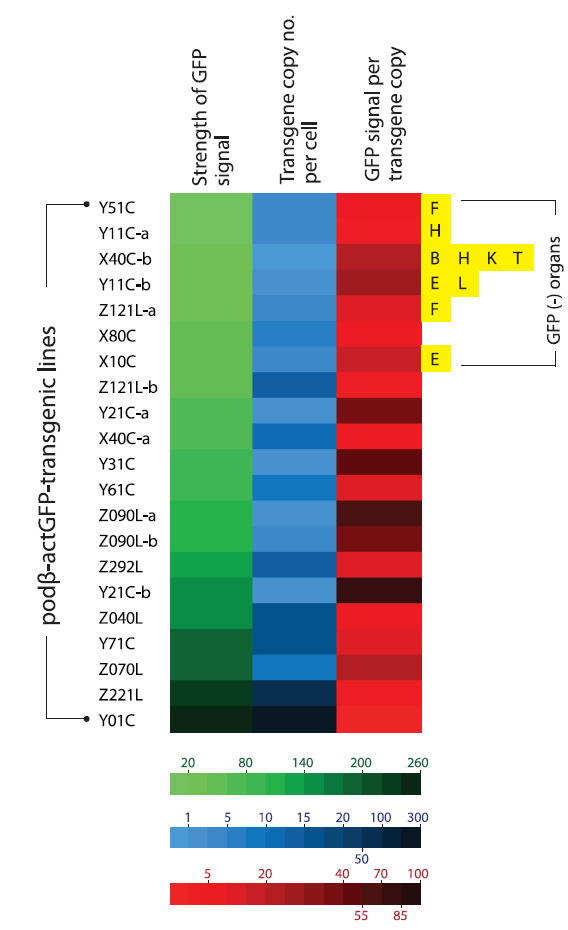
The data obtained at the transgene genotype and phenotype levels reveal a potential interrelationship between transgene copy number and GFP expression (Fig. 6). There exists neither a direct nor proportional correlation between the expression levels of transgenic GFP and copy numbers of transgene integrants. There is no simplified pattern of cross-talk between the two parameters. However, some common tendencies in the relationship between transgene copy number and expression characteristics should be considered. First, the highest GFP expression levels were achieved in the transgenic strains with the highest, absolute transgene copy numbers, as seen in the Z221L and Y01C lines. This is largely contradictory to previous claims that transgenic animals with very high numbers of transgene copies in long concatemers often fail to display high expression levels (Dorer and Henikoff 1997; Geurts et al. 2006). On the other hand, this finding is in agreement with previous observations of high-copy-number transgenic marine medaka strains carrying different transgenes, suggesting a relatively high capability of the marine medaka genome to maintain high copies of exogenously introduced genes (Cho et al., 2011; 2013a). Transgenic lines possessing single- or only a few transgene copies are more likely to display relatively weak expression levels than the high-copy-number transgenic strains. The bottom five ranking transgenic lines (
In summary, diverse transgenic marine medaka strains harboring a GFP reporter construct under the regulation of an endogenous