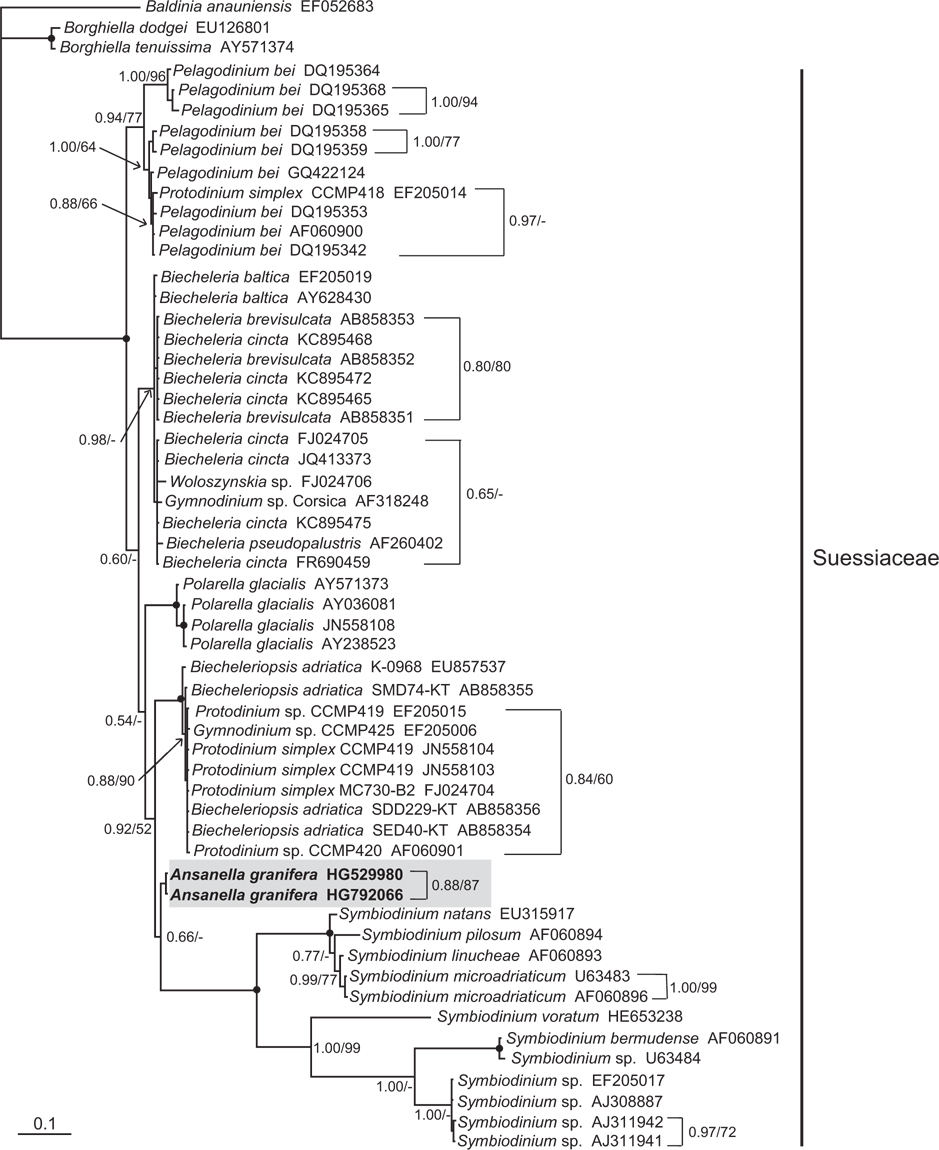
A small dinoflagellate, Ansanella granifera gen. et sp. nov., was isolated from estuarine and marine waters, and examined by light microscopy, scanning electron microscopy, and transmission electron microscopy. In addition, the identity of the sequences (3,663-bp product) of the small subunit (SSU), internal transcribed spacer (ITS) region (ITS1, 5.8S, ITS2), and D1-D3 large subunit (LSU) rDNA were determined. This newly isolated, thin-walled dinoflagellate has a type E eyespot and a single elongated apical vesicle, and it is closely related to species belonging to the family Suessiaceae. A. granifera has 10-14 horizontal rows of amphiesmal vesicles, comparable to Biecheleria spp. and Biecheleriopsis adriatica, but greater in number than in other species of the family Suessiaceae. Unlike Biecheleria spp. and B. adriatica, A. granifera has grana-like thylakoids. Further, A. granifera lacks a nuclear fibrous connective, which is present in B. adriatica. B. adriatica and A. granifera also show a morphological difference in the shape of the margin of the cingulum. In A. granifera, the cingular margin formed a zigzag line, and in B. adriatica a straight line, especially on the dorsal side of the cell. The episome is conical with a round apex, whereas the hyposome is trapezoidal. Cells growing photosynthetically are 10.0-15.0 µm long and 8.5-12.4 µm wide. The cingulum is descending, the two ends displaced about its own width. Cells of A. granifera contain 5-8 peripheral chloroplasts, stalked pyrenoids, and a pusule system, but lack nuclear envelope chambers, a nuclear fibrous connective, lamellar body, rhizocysts, and a peduncle. The main accessory pigment is peridinin. The SSU, ITS regions, and D1-D3 LSU rDNA sequences differ by 1.2-7.4%, >8.8%, and >2.5%, respectively, from those of the other known genera in the order Suessiales. Moreover, the SSU rDNA sequence differed by 1-2% from that of the three most closely related species, Polarella glacialis, Pelagodinium bei, and Protodinium simplex. In addition, the ITS1-5.8S-ITS2 rDNA sequence differed by 16-19% from that of the three most closely related species, Gymnodinium corii, Pr. simplex, and Pel. bei, and the LSU rDNA sequence differed by 3-4% from that of the three most closely related species, Protodinium sp. CCMP419, B. adriatica, and Gymnodinium sp. CCMP425. A. granifera had a 51-base pair fragment in domain D2 of the large subunit of ribosomal DNA, which is absent in the genus Biecheleria. In the phylogenetic tree based on the SSU and LSU sequences, A. granifera is located in the large clade of the family Suessiaceae, but it forms an independent clade.
Dinoflagellates are ubiquitous protists in marine environments (Jeong et al. 2010, Gomez 2012
The order Suessiales was established by Fensome et al. (1993), with the two new families Suessiaceae and Symbiodiniaceae. Species in the family Suessiaceae comprised fossil dinosporin cysts with a 7-9 latitudinal paraplate series (i.e., arrangement of amphiesmal vesicles [AVs] in latitudinal series), whereas species in the family Symbiodiniaceae comprised extant symbionts with a 7-latitudinal plate series. However, the number of latitudinal series of AVs has since been extended to include species with more than 10 (Kremp et al. 2005). Moestrup et al. (2009
Recently, we isolated thin-walled dinoflagellates from Shiwha Bay, Korea, and established two clonal cultures. The morphology of the cells from the cultures was almost identical, and there was only one base pair difference in the sequences of their SSU, internal transcribed spacer (ITS) region (ITS1, 5.8S, ITS2), and D1-D3 large subunit (LSU) rDNA. On the basis of morphological characters (in particular, the structure of the eyespot, thylakoids, and apical apparatus) and molecular analyses, these dinoflagellates are classified as a species of the family Suessiaceae, of which Symbiodiniaceae is considered to be a synonym. However, there is no other genus or species matching the characteristics of these dinoflagellates within the family. Therefore, in this study, we propose classifying these thin-walled dinoflagellates in the new genus
This study describes the morphological features of
Samples of surface sediment were collected from Shiwha Bay, Korea (37˚18′ N, 126˚36′ E) in 2010 and 2012, using an Ekman grab (Wildco; Wildlife Supply Company, Buffalo, NY, USA). Samples were stored in the dark at 4℃ until processed further. The sediments were sieved consecutively through 100-µm and 15-µm Nitex meshes. The material on the 15-µm mesh was then transferred to a 50-mL beaker containing filtered seawater. A manual vortex was applied and the suspended sediment fraction was recovered. The sediment fraction was incubated in f/2 medium without Si (Guillard and Ryther 1962), in a growth chamber at 20℃ under 20 µmol m-2s-1 illumination, in a 14-h light, 10-h dark cycle. The incubated sediments were observed regularly for the presence of motile cells, using a stereomicroscope (SZX-12; Olympus, Tokyo, Japan).
In 2013, one clonal culture of
The morphology of
For SEM, a 20-mL aliquot of a dense culture of
For TEM, cells from a dense culture were transferred to a 10-mL tube, and fixed in 2.5% glutaraldehyde (final concentration). After 1.5-2 h, the contents of the tube were placed in a 10-mL centrifuge tube, and concentrated at 1,610 ×g for 10 min in a Vision Centrifuge VS-5500 (Vision Scientific Company, Bucheon, Korea). The pellet from the tube was subsequently transferred to a 1.5-mL tube and rinsed in 0.2 M sodium cacodylate buffer at pH 7.4. After several rinses, the cells were post-fixed for 90 min in 1% osmium tetroxide (final concentration) in deionized water. The pellet was then embedded in agar. Dehydration was performed using a graded ethanol series (50, 60, 70, 80, 90, and 100% ethanol), followed by two changes of 100% ethanol. The material was embedded in Spurr’s low-viscosity resin (Electron Microscopy Sciences, Fort Washington, PA, USA) (Spurr 1969). Sections were prepared using an RMC MT-XL ultramicrotome (Boeckeler Instruments Inc., Tucson, AZ, USA) and stained with 3% aqueous uranyl acetate followed by lead citrate. The sections were viewed with a JEOL-1010 transmission electron microscope (JEOL Ltd., Tokyo, Japan).
>
DNA extraction, polymerase chain reaction (PCR) amplification, sequencing, and data analysis
Three to five cells of
>
Sequence availability and phylogenetic analysis
Phylogenetic analyses of the SSU and LSU rDNA regions of
Approximately 100,000 cells of
Phylum Dinoflagellata Butschli 1885 sensu Fensome et al. 1993
Class Dinophyceae Pascher 1914
Order Suessiales Fensome et al. 1993
Family Suessiaceae Fensome et al. 1993
>
Ansanella H. J. Jeong, S. H. Jang, Moestrup and N. S. Kang gen. nov.
Description. Free-living dinoflagellates with very thin transparent polygonal AVs arranged in longitudinal rows. The AVs are further arranged in 10-14 horizontal rows: 4-6 rows on the episome, 3-4 rows in the cingulum, and 3-4 rows on the hyposome. A single straight elongated apical vesicle, containing a single row of globular knobs, is present on the cell apex. Grana-like thylakoids in the chloroplasts and an eyespot of Type E are present. The peridinin-containing chloroplasts are lined by three membranes. A 51-base pair fragment of domain D2 of the large subunit rDNA, which is not found in genus
Type species.
Etymology. The generic name “ansanella” refers to the name of the city, Ansan, enclosing Shiwha Bay where this species was collected.
>
Ansanella granifera H. J. Jeong, S. H. Jang, Moestrup and N. S. Kang sp. nov.
Description. Episome conical with a round apex, larger than the trapezoidal hyposome. The cell has a wide and distinctive cingulum, which is displaced about its own width. The length and width of living cells are 10-15 µm and 9-12 µm, respectively. The ratio of cell length to cell width of living cells is 1.1-1.3. The nucleus is oval and located in the anterior to central part of the cell. Chloroplasts are yellowish-brown with the lobes arranged mainly near the surface of the cell; three or four starchenveloped pyrenoids are penetrated by thylakoids; pusule present. Peduncle and trichocysts absent. Estuarine and marine dinoflagellate.
Holotype. A holotype slide labeled USNM slide 1231539, of a culture fixed in 1% (final concentration) osmium tetroxide, was deposited in the Protist Type Specimen Slide Collection, US Natural History Museum, Smithsonian Institution, Washington, DC, USA.
Molecular characterization of
Type locality. Shiwha Bay, Korea (37˚18ʹ N, 126˚ 36ʹ E).
Etymology. The specific name “granifera”, indicates the characteristic chloroplast grana-like structures of this organism.
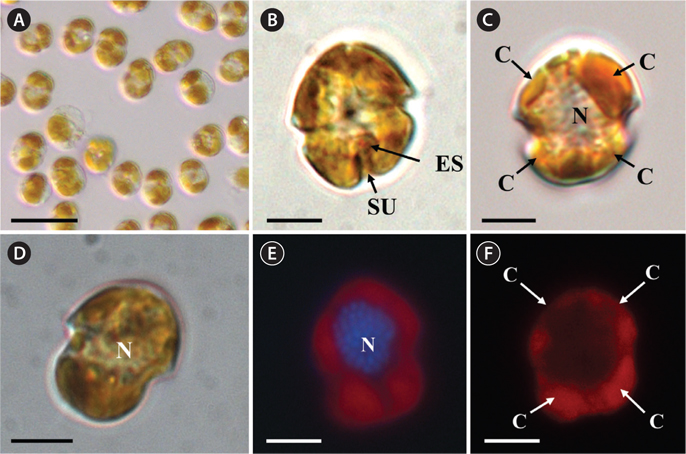
The morphologies of all the observed cells of
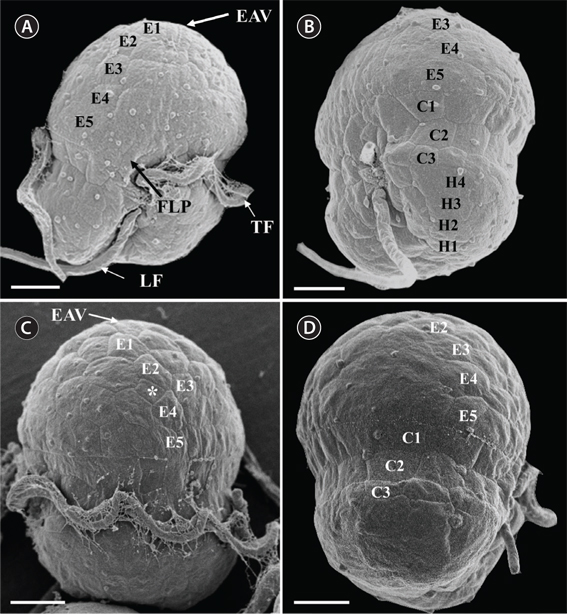
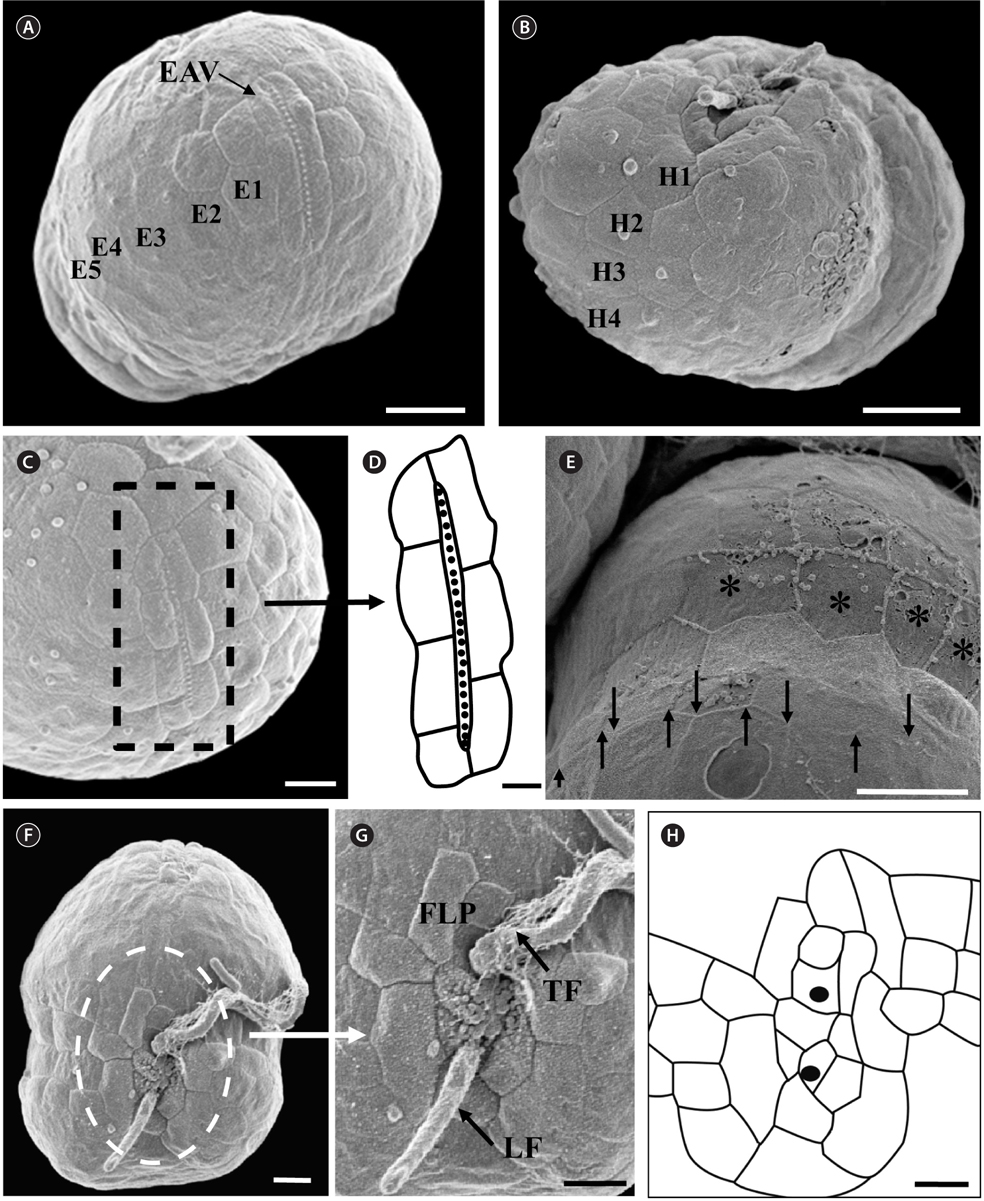
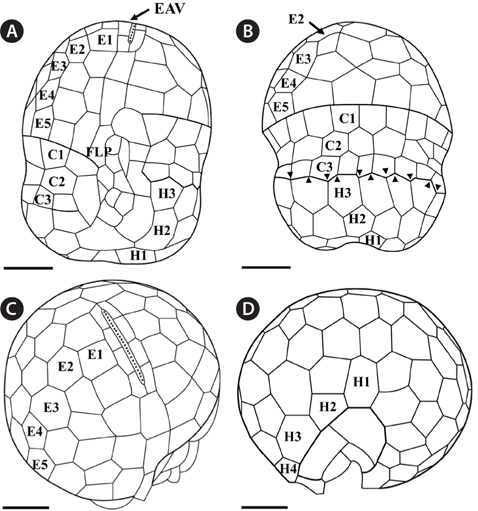
Under the SEM, the elliptical to round episome appeared larger than the trapezoidal hyposome (Fig. 2A-C). Cells were covered with hexagonal or pentagonal AVs. The length of each side of a hexagonal or pentagonal AV was 1.1-2.5 µm. The AVs were arranged in 10-14 rows (mostly 11, n = 100), as follows: 4-6 rows on the episome (mostly 5, n = 100), 3-4 rows in the cingulum (mostly 3, n = 100), and 3-4 rows on the hyposome (mostly 3, n = 100) (Figs 2A-D, 3A, B & 4A-D, Appendix 1D & E). The total number of AVs was ~160. The apical furrow apparatus consisted of a narrow EAV, approximately 4.1 µm, which extended from the mid-ventral side of the episome, over the apex to the dorsal side (Figs 3A, C, D & 4C). The EAV was ornamented with a central row of approximately 20-25 small knobs (Figs 3A, C, D & 4C), and was bordered by X+4-6 platelets (Figs 3A, C, D & 4C, Appendix 1F & G). In the dorsal part of
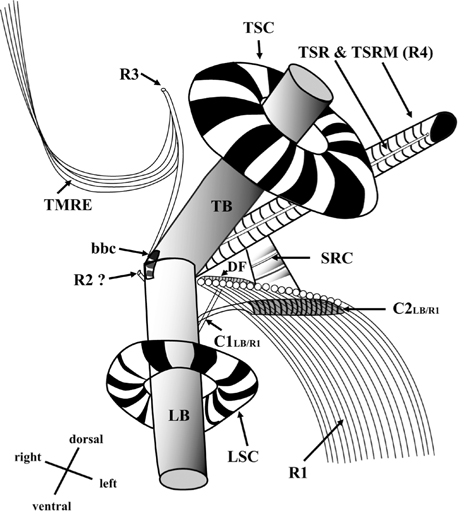
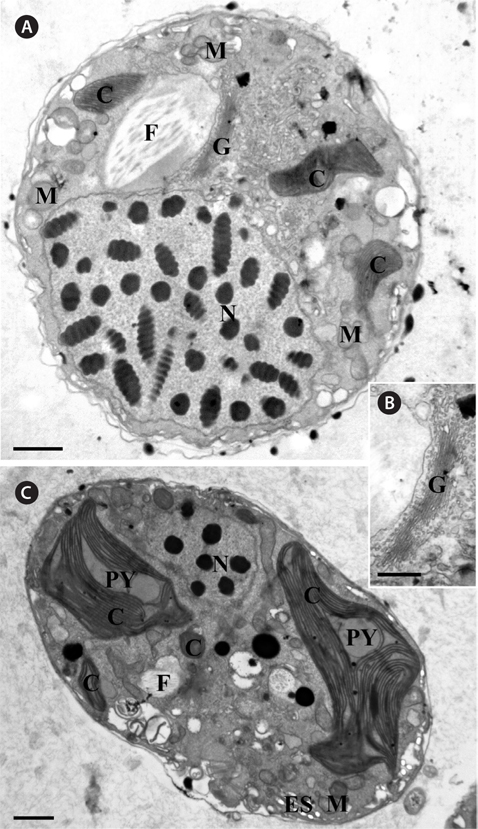
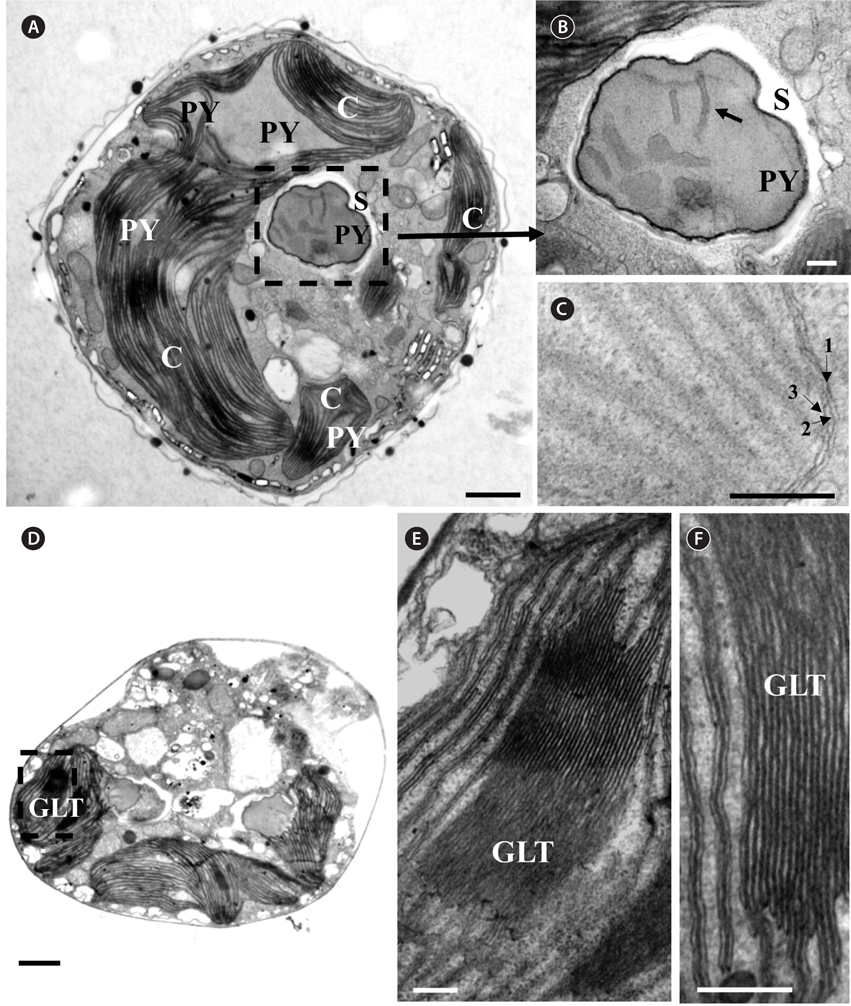
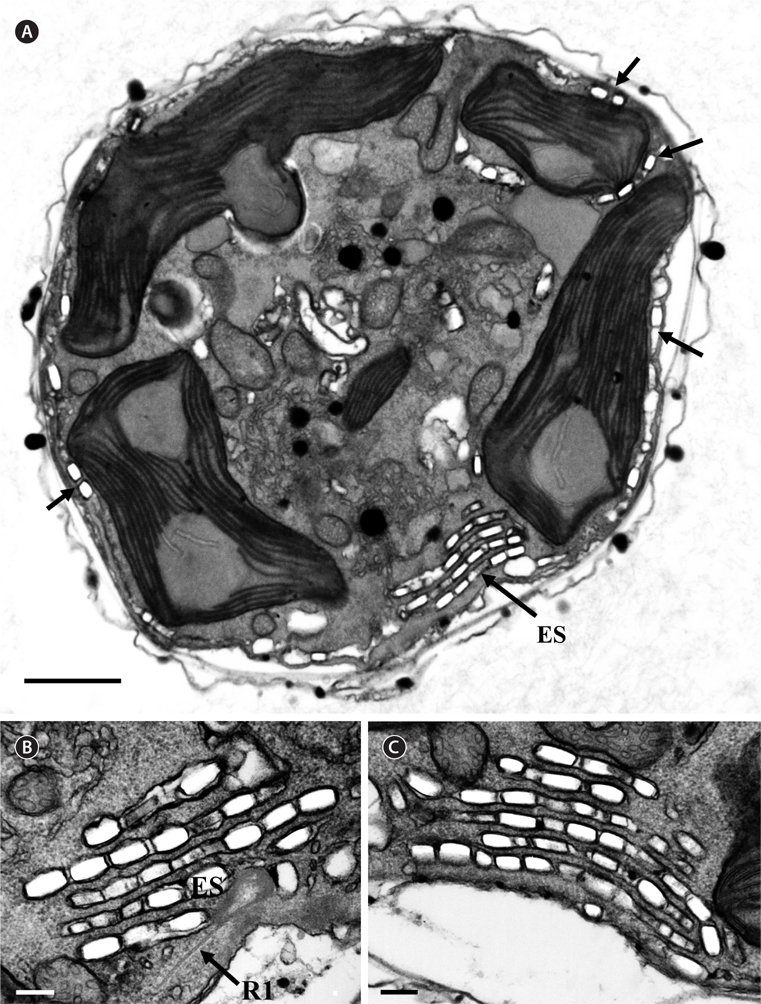
Thin TEM sections showed the main features of the cell, such as the chloroplasts, eyespot, fibrous vesicles, golgi body, mitochondria, nucleus, pyrenoid, and starch (Figs 5A-C & 6A). Longitudinal TEM serial sections showed the nucleus to contain many chromosomes, and its length and width were approximately half of the cell length (data not shown). The nucleus lacked nuclear chambers and a nuclear fibrous connective. Mitochondria were present in both central and peripheral parts (Fig. 5A-C). The golgi apparatus, composed of 6-10 stacked cisternae and fibrous vesicles, was located near the nucleus (Fig. 5A & B). Chloroplasts with complex lobes were located predominantly near the cell surface. These chloroplasts typically had three thylakoids per lamella (Fig. 6C & F), but arrangements of five, six, or more thylakoids per lamella (i.e., grana thylakoid associations) were also seen (Fig. 6D-F, Appendix 2B & C). All sectioned cells showed extensive “grana” formation (Fig. 6D-F, Appendix 2B & C). Three evenly spaced membranes enclosed each chloroplast (Fig. 6C). The chloroplasts formed a peripheral network that comprised 3-4 pyrenoids, each surrounded by a hemispherical starch grain (Fig. 6A & B, Appendix 2A). Underneath the sulcus, there was a large and conspicuous eyespot comprising approximately ~7 cisternae with brick-like material (Fig. 7A-C). The length and width of the crystalline bricks were 100-230 nm and 50-120 nm, respectively. The eyespot was located near the flagellar apparatus (Fig. 7B). A single row of microtubules (i.e., Root 1, R1) was observed in the narrow space between the outermost cisterna and the cell membrane of the sulcal region (Fig. 7B). The flagellar apparatus of several cells was examined in TEM serial sections (Figs 8-12), and a diagrammatic 3D-reconstruction was generated (Fig. 13). The transverse basal body (TB) and longitudinal basal body (LB) formed an angle of approximately 135˚ (Fig. 8A), as estimated from the serial sections. A striated collar surrounded the exit aperture of each flagellar canal (longitudinal striated collar and transverse striated collar [TSC]) (Figs 8 & 12). In the flagellar root system, a large multi-membered microtubular root 1 (R1, longitudinal microtubular root) was located to the left of the LB (Figs 8-10 & 12) and extended underneath the sulcus toward the antapex (not shown). A single-stranded microtubular root (R2) was associated obliquely with the right side of the LB (Figs 8D, 9C, 10C & D). The transverse microtubular root (R3) was attached to the right side of the TB (Fig. 10). The transverse microtubular root extension structure was located near the TB and R3 (Fig. 10C & D). The transverse striated root and the transverse striated root microtubule of root 4 were located near the TB (Figs 8E, 11 & 12A). The two basal bodies were connected by a small striated connective, the basal body connectives (Fig. 8D). A number of fibrous connectives or fibres were associated with the various roots. Thus, the R1 root was attached to the LB by two fibrillar connectives, the C1LB/R1 and C2LB/R1, respectively (Fig. 9). The C1LB/R1 was situated slightly posterior to C2LB/R1 and attached to the two rightmost microtubules of the R1 and one of the LB triplets. It was relatively short and only visible in two consecutive sections. The C2LB/R1 showed a distinct striation pattern and was intimately associated with the ventral surface of the R1 (Fig. 9C). The R1 and R4 roots were interlinked by a striated root connective (Fig. 8D & E). A dorsal fibre, with almost the same striation pattern as the striated fibre of R4, was located on the dorsal side of the R1 (Figs 10B, C, E & 12C). Transverse TEM serial sections showed the pusule system (PU) (Fig. 11). In the right-antapical side view, the PU was located next to the TBs and TSC (Fig. 11). We serially sectioned 5 whole cells transverally and longitudinally, but peduncle, lamellar body and rhizocysts were not observed.
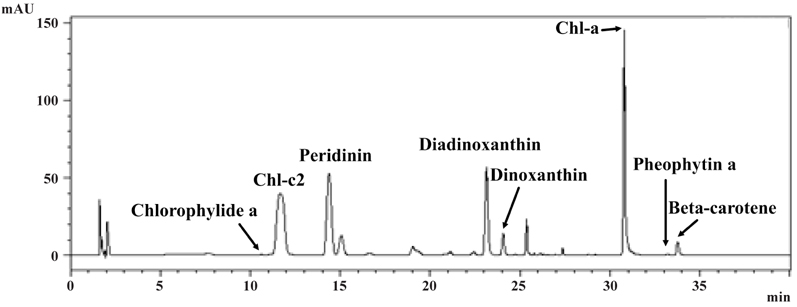
Pigment analysis of
DNA sequences of
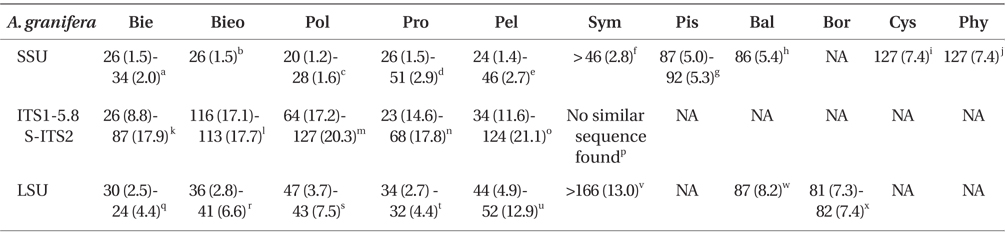
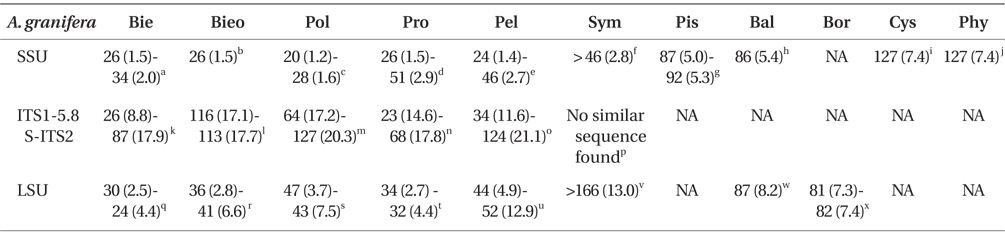
Comparison of the sequences of Ansanella granifera AGSW10 (GenBank accession Nos. HG529978-HG529980) with other genera included in the order Suessiales
When properly aligned, the SSU rDNA sequence of
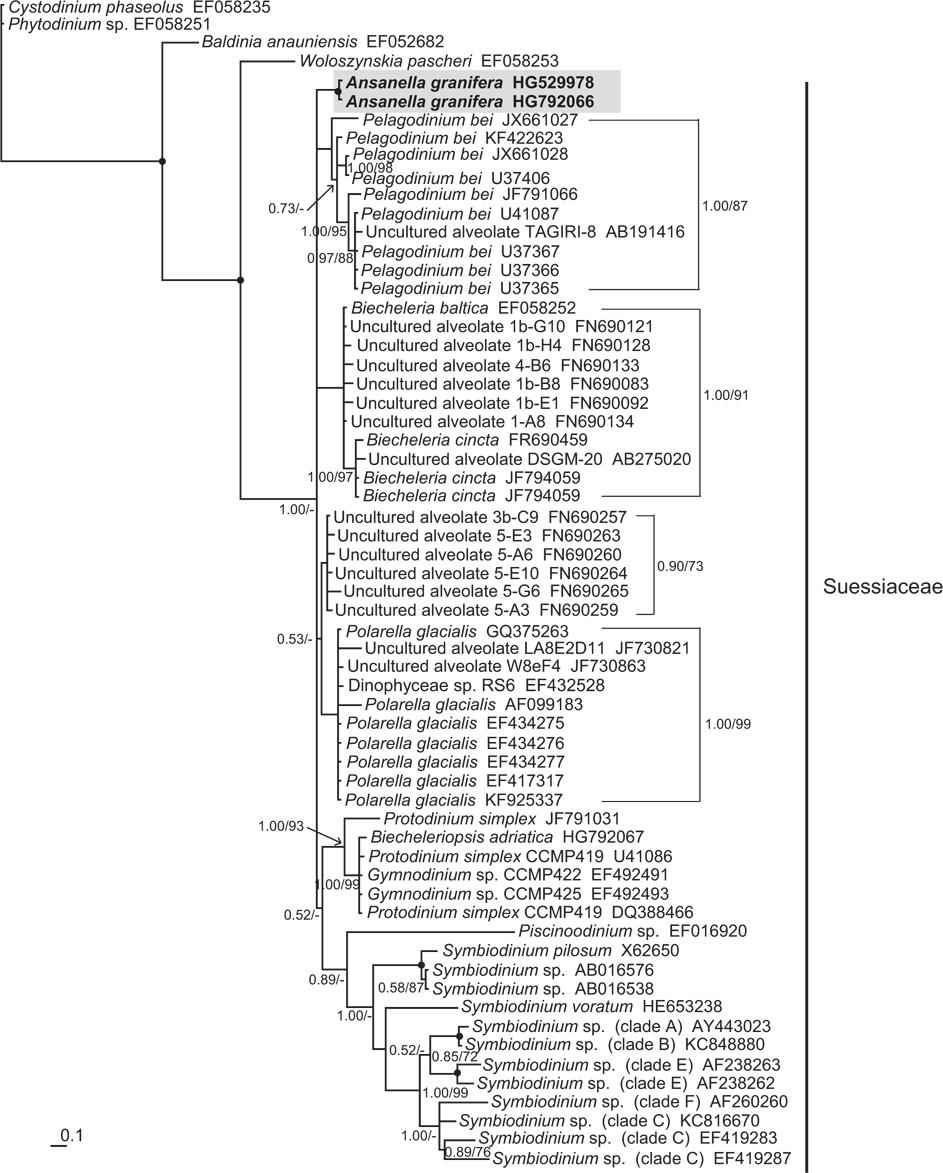
In the phylogenetic trees based on the SSU and LSU rDNA sequences of dinoflagellates,
>
Remarks on culturing and behavior
The newly isolated cells moved in straight lines or with a helicoidal mode. They sometimes stopped suddenly, and then resumed movement very quickly, with a jumplike action. They grew at salinities > 10 or <35, but died in fresh water. Thus, this dinoflagellate is an estuarine and marine species.
The morphology of
In the family Suessiaceae, the Type E eyespot has been reported previously in
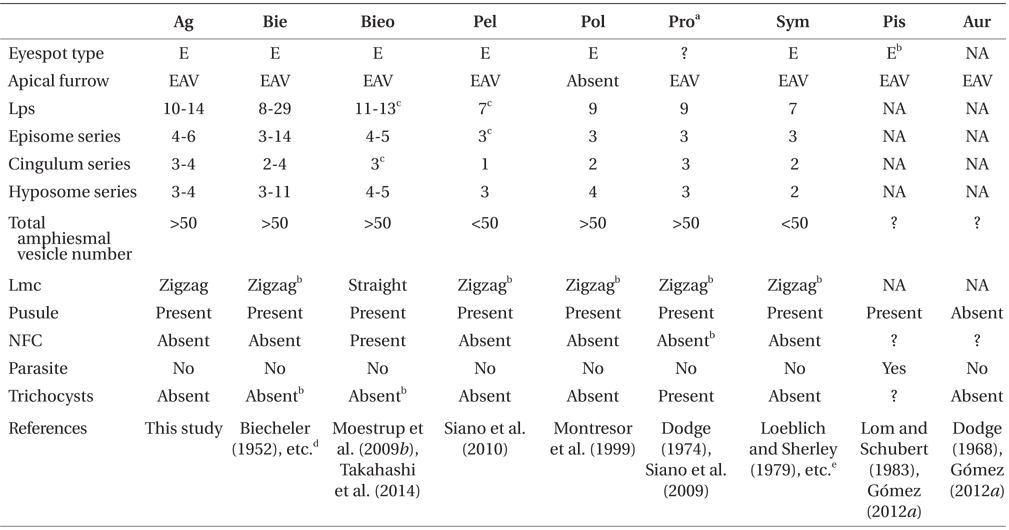
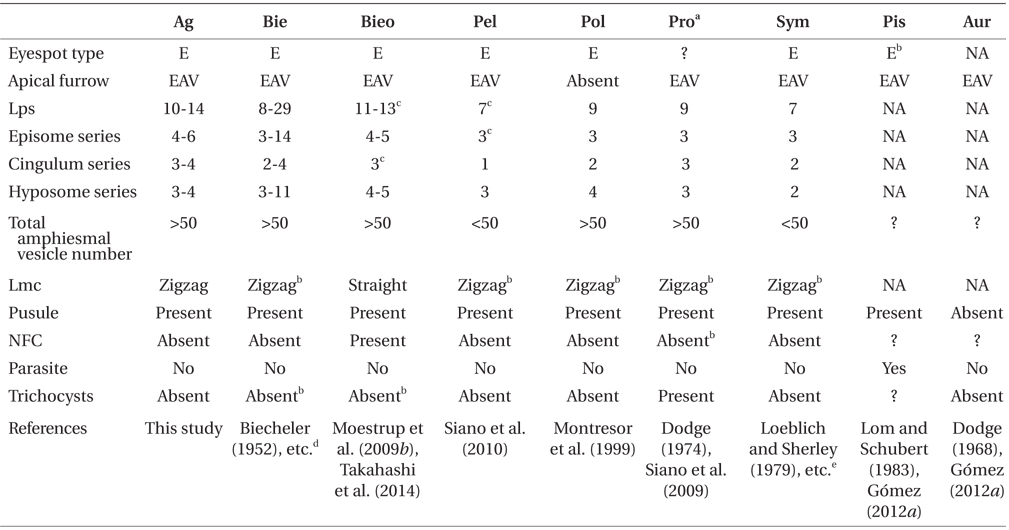
Comparison of the morphology of Ansanella granifera and other genera included in the family Suessiaceae
The flagellar apparatus of
The SSU, ITS1, 5.8S, ITS2, and D1-D3 LSU rDNA sequences of the genus
There are indications that the order Suessiales may comprise not only the fish parasite




![Ansanella granifera AGSW10 taken by transmission electron microscopy. (A) Longitudinal section of the cell showing the basal bodies (longitudinal basal body [LB], transverse basal body [TB]) and the eyespot (ES). (B-E) Flagellar apparatus. Non-adjacent, nearly longitudinal serial sections proceeding from left to right. The encircled numbers are section numbers. Micrograph showing relative positions of the LB, TB, Root 1 (R1), putative Root 2 (R2), Root 4 (R4), striated root connective (SRC), basal body connectives (bbc), and longitudinal striated collar (LSC). Scale bars represent: A-E, 0.2 μm.](http://oak.go.kr/repository/journal/13610/JORHBK_2014_v29n2_75_f008.jpg)
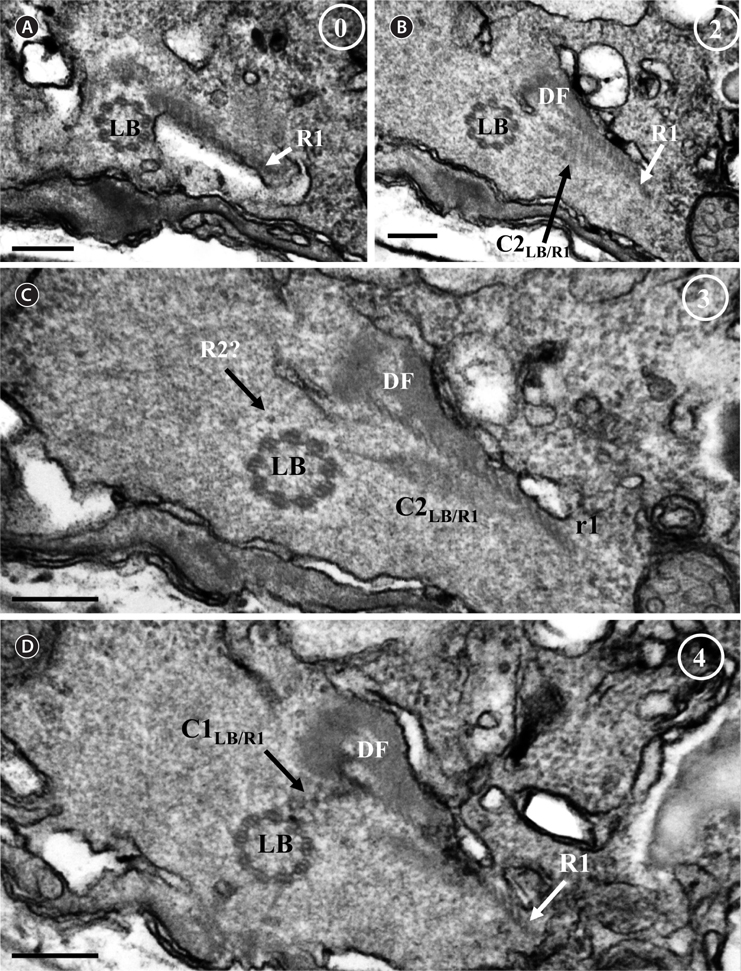
![Ansanella granifera AGSW10 taken by transmission electron microscopy. Nonadjacent longitudinal serial sections of the flagellar apparatus. The cell is seen from the outside, and the sectioning proceeds from right to left. The encircled numbers are section numbers. (A-D) Micrograph showing relative positions of the longitudinal basal body (LB), transverse basal body (TB), Root 1 (R1), putative Root 2 (R2), Root 3 (R3), Root 4 (R4), dorsal fiber (DF), and microtubular extension (transverse microtubular root extension [TMRE]). The striation pattern of the DF is very distinct. Notice also flagellar root R3 and its microtubular extension (TMRE). (E) Enlargement from Fig. 10C. Scale bars represent: A-D, 0.2 μm.](http://oak.go.kr/repository/journal/13610/JORHBK_2014_v29n2_75_f010.jpg)
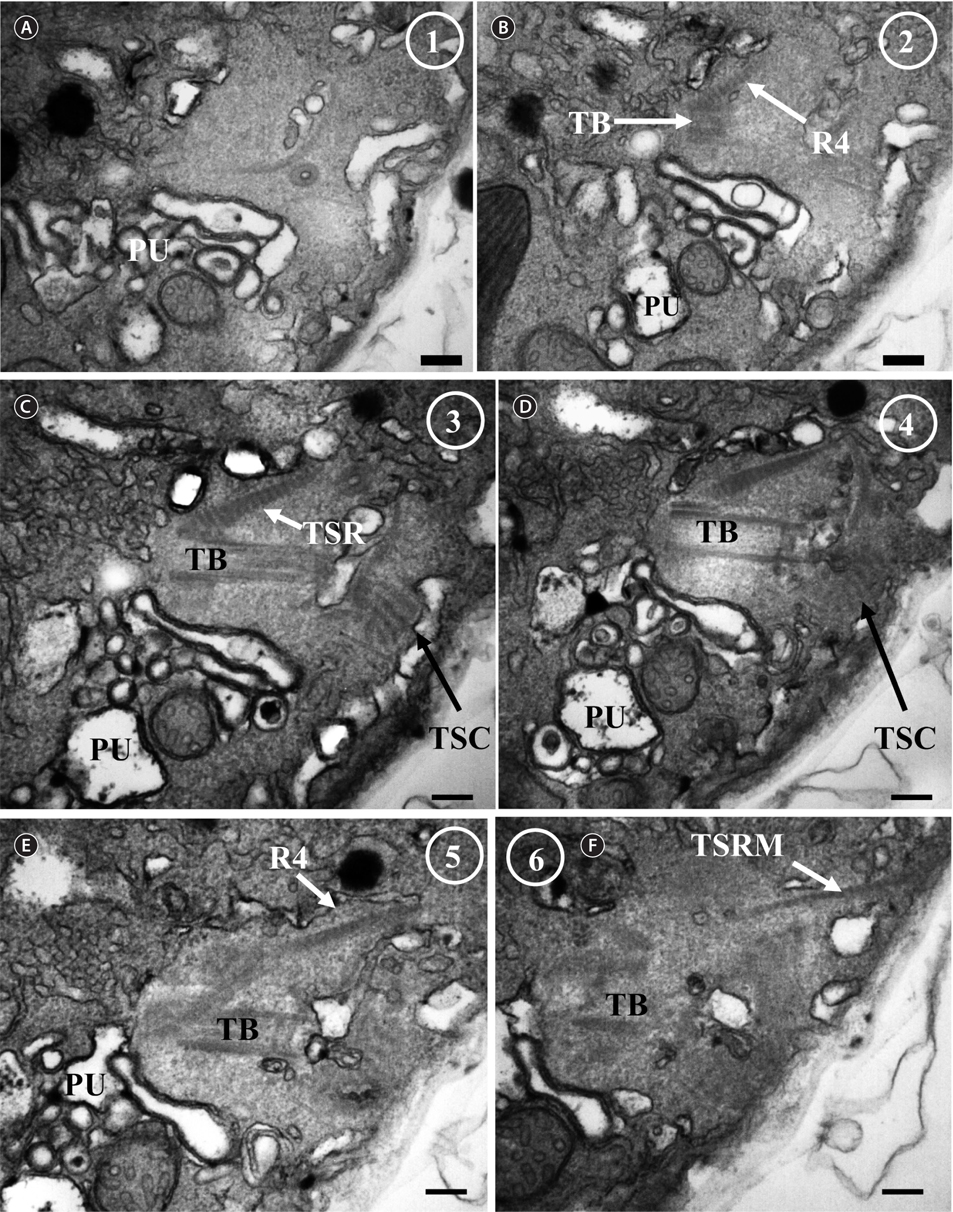
![Ansanella granifera AGSW10 taken by transmission electron microscopy. Nonadjacent serial sections. The sulcus region in transverse section, the sectioning moves from posterior to anterior. The encircled numbers are section numbers. (A-D) Micrograph showing relative positions of the longitudinal basal body (LB), transverse basal body (TB), Root 1 (R1), Root 4 (R4, transverse striated root [TSR] + transverse striated root microtubule [TSRM]), dorsal fiber (DF), transverse striated collar (TSC), and eyespot (ES). The R1 flagellar root is located in the narrow space between the eyespot and the cell surface. Scale bars represent: A-D, 0.2 μm.](http://oak.go.kr/repository/journal/13610/JORHBK_2014_v29n2_75_f012.jpg)