Micro-filamentous green algae grow on a variety of solid substrata such as wood, rock, pebbles, and plastic (Correa et al. 1994, Correa 1997). They also occur on or in other organisms, as epiphytes or endophytes that are mostly harmless, but a few algae have been reported to be pathogens of other algae (Correa et al. 1994, Correa 1997) or corals (Goldberg et al. 1984). Many filamentous algae live deeply embedded within tissues of larger algal hosts and this endophytic habit represents a type of symbiosis (Lewis 1973, Starr 1975, Goff 1982, Lewin 1982, Douglas and Smith 1989, Gauna and Parodi 2008). Some filamentous endophytes cause only minor changes in their hosts, whereas others are known to produce either degradative losses or tumoral lesions in their hosts (Andrews 1977, Garbary 1979, Yoshida and Akiyama 1979, O’Kelly and Yarish 1981, Nielsen and McLachlan 1986a, 1986b, Peters 1991, Brodie et al. 2007).
The endophytic brown alga, Streblonema aecidioides causes tissue thickening in commercial Undaria sp. (Yoshida and Akiyama 1979), and Streblonema-like endophytes are known to produce galls in some algal hosts (Andrews 1977, Apt 1988). An economically important alga, Chondrus crispus shows severe lesions and cellular damage when infected by the green algae, Acrochaete operculata and A. heteroclada (Correa et al. 1988, Correa and McLachlan 1991, 1992). Similar lesions have also been described in Mazzaella laminarioides after infection with Endophyton ramosum (Correa et al. 1994, Sánchez et al. 1996). A field population of Laminaria hyperborea was heavily infected with endophytic algae, and infection changed the host morphology and commercial values (Lein et al. 1991).
Algae Base lists 109 microfilamentous ulvophycean endophytes (Guiry and Guiry 2014). The morphology of many endophytes is relatively simple, and identification is often difficult or impossible because of the absence of diagnostic characters (Rinkel et al. 2012). The genus Ulvella was established as Acrochaete, and to date, 48 species have been identified based on the morphology and tufA gene sequences (Pringsheim 1863, Nielsen et al. 2013, Guiry and Guiry 2014). In addition, the genera of Bolbocoleon and Blastophysa currently include two and one species, respectively (Guiry and Guiry 2014). A further constraint on identification is that a single host frond or species may have many species of green and brown algal endophytes (e.g., C. crispus, Correa et al. 1987, Correa and McLachlan 1994).
The genus Grateloupia distributes from tropical to warm temperate regions of the world, and includes approximately 90 species (Lee et al. 2009, Guiry and Guiry 2014). Grateloupia has been used as food and for agglutinin medication in Korea (Kang 1968, Oh et al. 1990). Iima and Tatewaki (1987) reported an endophytic Blastophysa rhizopus in a Grateloupia host. B. rhizopus is known as a pathogenic green alga resulting in “green spot rotting,” which destroys tissue in Neodilsea yendoana host plants (Iima and Tatewaki 1987). In Korea, Ulvella viridis (formerly Entocladia viridis) was briefly reported as an epiphyte of Griffithsia japonica (Lee et al. 1998), but no data has shown that it is an endophyte in any other seaweeds. We initially observed that Grateloupia spp. were commonly associated with green endophytic filaments. The primary aims of this study were to identify these green endophytes and to characterize the abundance of the infections in four Grateloupia species from Korea.
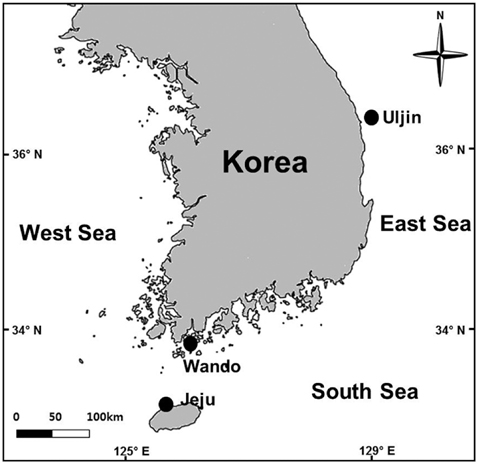
Grateloupia spp. host plants were collected from Jocheon, Jeju (33˚32′ N, 126˚38′ E), Jungdo-ri, Wando (34˚17′ N, 126˚42′ E) and Jukbyeon, Uljin (36˚59′ N, 129˚25′ E), Korea in August and September 2013 (Fig. 1). The presence or absence of infection by endophytic seaweeds was investigated under a microscope after obtaining tissue sections from 28 to 57 thalli from each study site. The infection ratio (%) was estimated by comparing the infected area with the total blade area for each plant collected from the three study sites. Each Grateloupia plant was photographed, the thallus surfaces and infected areas were measured using Image J software, and the infection ratio was calculated. Grateloupia fronds were divided into upper, middle, and lower parts in order to examine the infection process of the endophytic algae. Finally, the infection rates were also estimated independently for vegetative, tetrasporophytic and gametophytic fronds.
To identify endophytic algae living in G. lanceolata, sample plants were collected from Jeju-Island, and cultured in the laboratory. For culture, the host plants were rinsed with filtered seawater and several disks (2.5 cm in diameter) were punched out using a cork borer from infected areas of Jeju Grateloupia population. The discs were re-cleaned with running tap water, rinsed several times using sterile seawater, and incubated in Petri dishes containing 100 mL of Provasoli’s enriched seawater medium (Provasoli 1968). Culturing was carried out in a multiroom incubator (Vision VS-1203PFC-L; Vision Science Co. Ltd., Gyeongsan, Korea), at 20 ± 1℃ and a 12 : 12 h light : dark (L : D) cycle using cool-white fluorescent tubes (15 μmol m-2 s-1). Endophytic algae grew in the cultured host plants within 1-2 weeks. The endophytes were separated from the hosts, and unialgal cultures were made. The culture medium was renewed every five days during the study period.
A phylogenetic analysis based on chloroplast-encoded elongation factor tufA gene sequences was performed to identify the endophytes. Each endophyte (0.03 g in fresh mass) was crushed separately with liquid nitrogen, and genomic DNA was extracted using the DNeasy Plant Mini-Kit (Qiagen, Hilden, Germany) according to the manufacturer’s protocols. Concentrations of extracted DNA (ca. 0.3 μg mg-1) were determined using gel electrophoresis in a 1% agarose gel. During polymerase chain reaction (PCR) amplification, the chloroplast-encoded elongation factor tufA genes were amplified using published primers (Famà et al. 2002) with a final volume of 40 μL per reaction. Following manufacturer’s recommendations, the amounts of materials were used during PCR: 20 ng of DNA template, 1 unit of hot-start Taq polymerase (Genet-Bio, Daejeon, Korea), 4 μL 10× PCR buffer, 2.5 mM MgCl2, 200 μM dNTP, 5 pM of each forward and reverse primer, and sterilized distilled water. PCR was conducted using the GeneAmp PCR System 9700 Thermal Cycler (Applied Biosystems, Foster City, CA, USA) with initial denaturing at 96℃ for 2 min and a 32 denaturation cycle at 94℃ for 30 s; annealing at 58℃ for 30 s; extension at 72℃ for 50 s and final extension at 72℃ for 30 min. All PCR products were held at 4℃ following amplification. Amplification was evaluated using gel electrophoresis in a 1% agarose gel, and the PCR products were cleaned using a PCR purification kit (Solgent, Daejeon, Korea). The PCR products were then commercially sequenced (miDNA Genome Research Institute, Kunsan, Korea). The sequences were determined using the ABI 3130xl Genetic Analyzer (Applied Biosystems) and assembled using the DNASIS Max 3.0 (MiraiBio, Alameda, CA, USA).
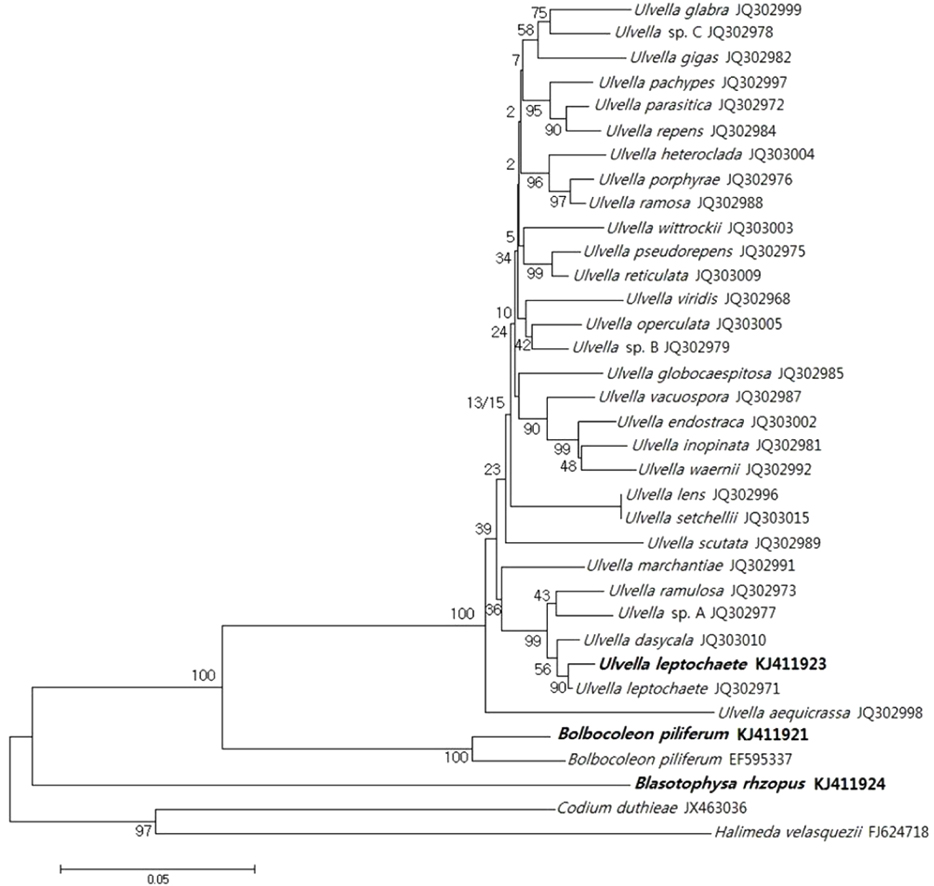
Related sequences were downloaded from GenBank according to BLAST results (Altschul et al. 1990). Phyloge\netic analyses used 1,000 bootstrap replications in neighbor-joining to evaluate the robustness of the tree topology. The data set included three new tufA sequences of endophytic algae and 32 tufA sequences from reference taxa. Codium duthieae and Halimeda velasquezii were used as outgroups.
For scanning electrom microscopy (SEM) observation, specimens were fixed with 4% glutaraldehyde in a 0.6 M phosphate buffer, rinsed in distilled water, dehydrated in a graded alcohol series, and critical point dried. Samples were then freeze-dried (ES-2030; Hitachi, Tokyo, Japan), mounted on a stub, and coated with platinum (E-1045; Hitachi). Specimens were observed using a field emission scanning electron microscope (S-4800; Hitachi).
A one-way ANOVA was used to compare significance among data. The ANOVA was followed by Duncan’s multiple range tests. All statistical analyses were performed using SPSS version 18.0 (SPSS Inc., Chicago, IL, USA) with the level of significance set at p < 0.05. All data are expressed as mean ± standard deviation.
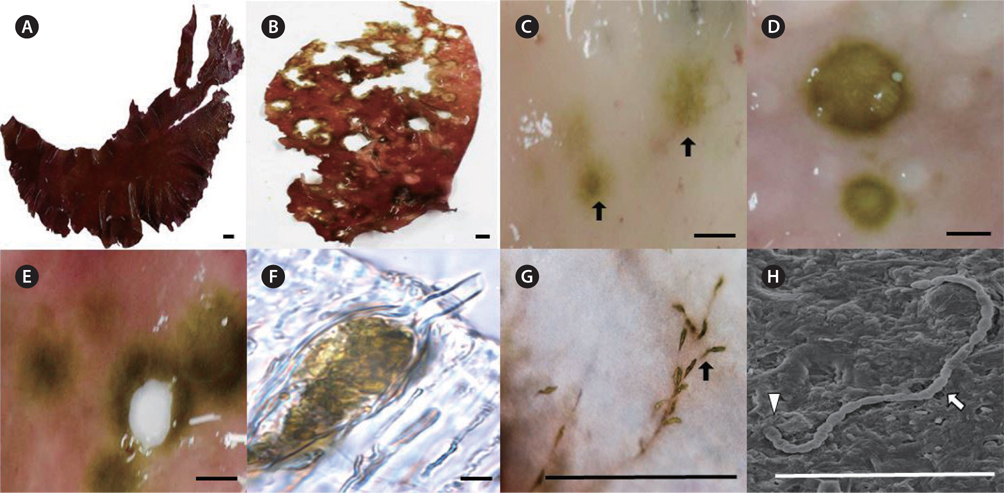
Healthy fronds of Grateloupia spp. are a dark red color with a gelatinous to cartilaginous texture (Fig. 2A). Infected Grateloupia plants had many holes and discrete light-green spots on the red fronds (Fig. 2B-E). As the endophytes became more abundant, the light-green color turned dark green, and the small spots became large holes (Fig. 2D & E). During later stages, the fronds became discolored, necrotic and torn (Fig. 2E). The endophytes grew between the cortical tissue and medulla on the host (Fig. 2F). In infected fronds, cross sections of the lesions showed green filaments embedded in the outer cell walls (Fig. 2F) that later formed a network of invasive thalli ramifying extensively into the host (Fig. 2G). The infected fronds had rough textures caused by the lesions, although no gall-like structures were observed (Fig. 2B). Endophytic filamentous algae were still found at the edge of such lesions. After infection by endophytic algae, the morphology of infected fronds differed from healthy Grateloupia thalli (Fig. 2B). During SEM observations, the endophytic cells appeared as protruding filaments from the surface of the host (Fig. 2H).
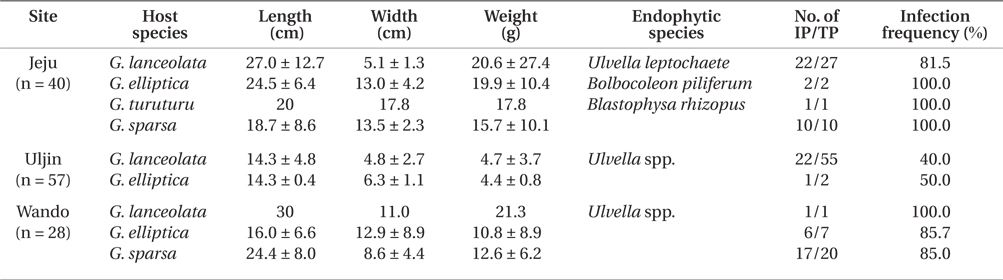
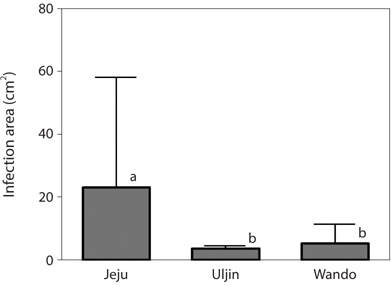
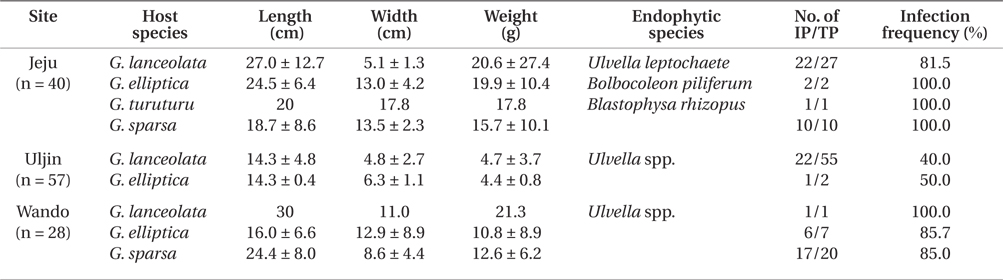
Infection by green endophytes was common in the field populations of Grateloupia spp. including G. lanceolata, G. elliptica, G. sparsa, and G. turuturu. The frequency of infected host plants was high and similar in the Jeju (87.5%) and Wando (85.7%), and that of Uljin was quite low with 40.4% (Table 1). In addition, the infected frond area was larger in the Jeju population than in the others (significant at p < 0.05) (Fig. 3). Among the 125 Grateloupia fronds, gametophytes were dominant (113 plants), followed by tetrasporophytes (9 plants), and vegetative plants (3 plants) (Table 1). The percentage of infected plants was 54.0% for gametophytes and 55.6% for tetrasporophytes.
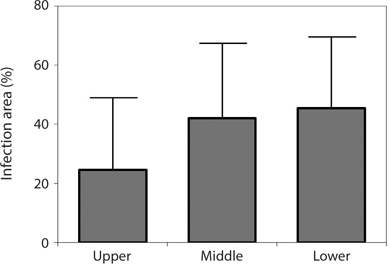
Endophytic algal infection was found on all parts of the Grateloupia fronds. There was an apparent trend towards, the lower parts of fronds having greater infection (45.35%) with values decreasing to 42.01% in the middle to 28.57% in the upper parts of fronds (Fig. 4); however, this trend was not significant (ANOVA, p > 0.05).
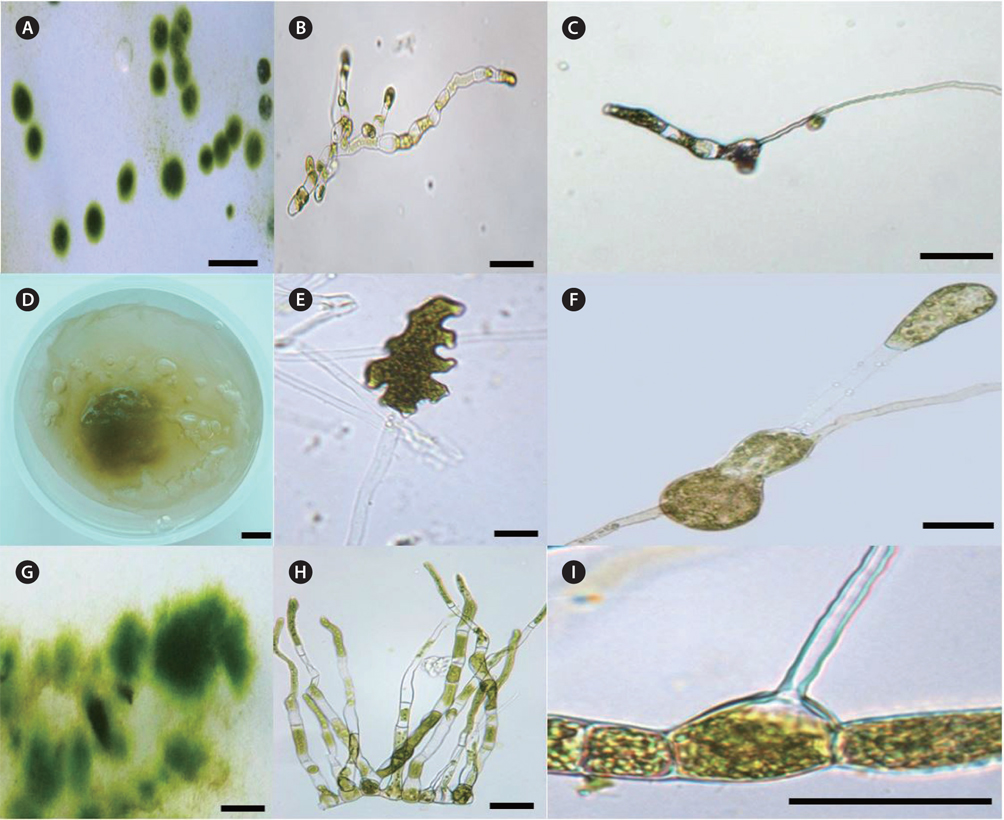
After 1-2 week culture, clumps of filamentous green endophytes were observed at the edge of the host frond disks. Some clumps had slightly different morphologies and each clump was cultured, separately. Three filamentous green endophytic species, U. leptochaete, B. piliferum, and B. rhizopus, were isolated and identified (Fig. 5). They all had cylindrical cells, irregular branches, and hairs. The three endophytes are described below.
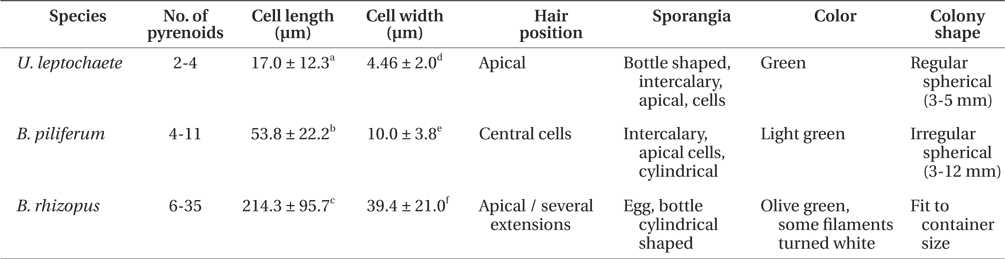
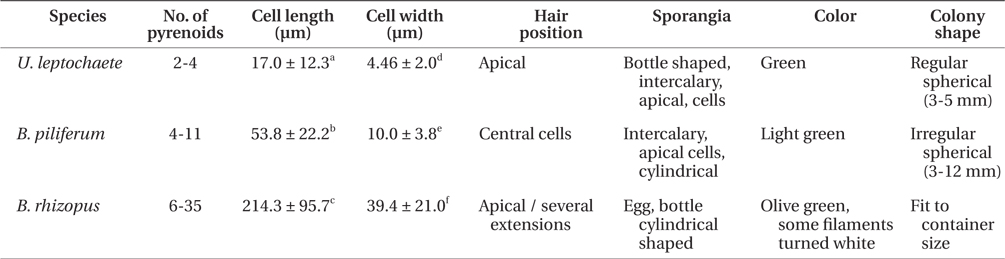
The green colonies form regular spherical masses 3-5 mm in diameter with cylindrical cells. The green filaments are 17.0 ± 12.3 μm long and 4.5 ± 2.0 μm wide with 1-4 pyrenoids per cell. Unpigmented hairs are found at the ends of filaments of the thalli and the sporangia are bottle shaped.
Colonies are of amorphous in shape. The cells are multi-nucleate coenocytes. The cells vary in shape from spherical to tubular but some possess irregular features such as the large, rounded, swollen cell-like utricles found in Codium (Fig. 5F). The filaments are 53.8 ± 22.2 ℃m in length and 10.0 ± 3.8 μm in width with 6-35 pyrenoids per cell. The hairs are found at apical sections of several extensions of thalli. Filaments are olive green with cylindrical cells and egg-, cylindricalor bottle-shaped sporangia.
Colonies are 3-12 mm in diameter with an irregular shape. The filaments have light-green cells, 214.3 ± 95.7 μm long and 39.4 ± 21.0 μm wide. Each cell has 4-11 pyrenoids. The hairs are in the central section of thalli, and the sporangia are a cylindrical shaped.
The three endophytes were distinguishable to the naked eye because of some morphological characteristic associated with colony formation (Table 2). Blastophysa rhizopus had multi-nucleate coenocytes, colonies 10 cm or more in diameter, and numerous pyrenoids compared to the other species. In addition, the species showed large, swollen utricle-like structures as seen in Codium. Bolbocoleon piliferum had much smaller colony sizes (3-12 mm in diameter), and prostrate filamentous plants with irregularly branched upright filaments. The hairs are found at prostrate filaments. Ulvella leptochaete had similarly small colony size; however, there were few pyrenoids per cell relative to the other species.
Although morphological differences were observed among the three endophytes, cultured specimens were definitively identified using tufA gene sequences. The tufA gene analysis for the three endophytes and published reference species is shown in Fig. 6, and this clearly identified the endophytes as members of the genera Ulvella, Blastophysa, and Bolbocoleon.
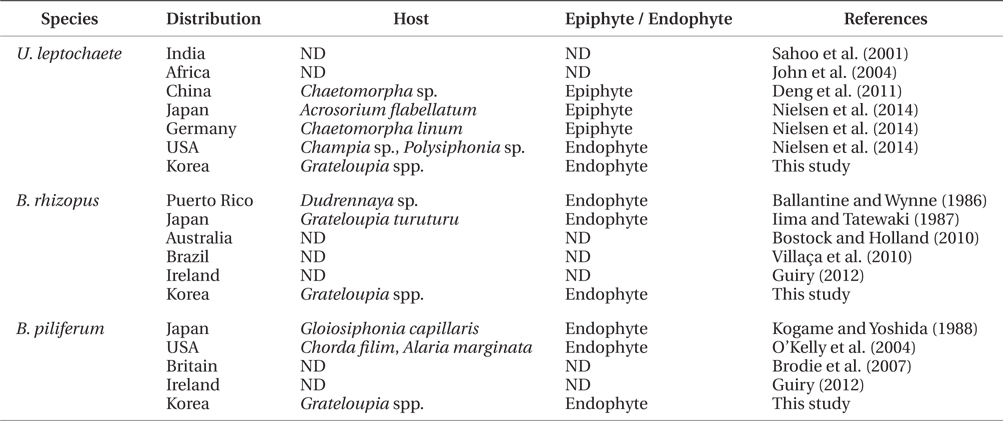
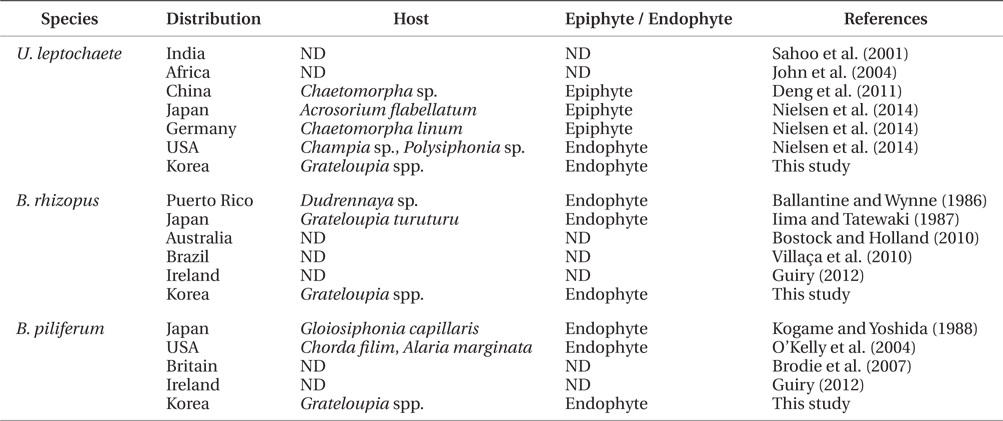
In the present study, three green endophytic filamentous algae, Ulvella leptochaete, Blastophysa rhizopus, and Bolbocoleon piliferum, were isolated and identified for the first time in Korea. The presence of endophytic Ulvella species was recently recorded in Chondrus ocellatus fronds in Korea (Lee et al. 2013), although epiphytic U. viridis was first reported on the fronds of Griffithsia japonica (Lee et al. 1998, Lee and Kang 2002). These three endophytic algae are distributed worldwide, and grow on or in diverse seaweeds as epiphytes and endophytes, which indicates that they are not host-specific (Table 3). Korean U. leptochaete was found to be similar in morphology and pyrenoid number per cell to the Chinese species, but its growth patterns were different both as an endophyte and epiphyte (Table 3). B. rhizopus exists as endophytes of various green, brown, and red algae (Iima and Tatewaki 1987, Burrows 1991). The filaments of Korean B. rhizopus are longer than those of the Japanese species. Finally, Bolbocoleon piliferum grows in marine and brackish water seaweeds. The plant length of an endophytic Korean B. piliferum is smaller than that of a Japanese one (Kogame and Yoshida 1988). In the present study, the three endophytic green algae grew all in the same frond of a Grateloupia host. This suggests that research on endophytic algae could increase information on species diversity and seaweed diseases in Korea, because many endophytes are currently regarded as pathogens of macroalgal diseases (Correa et al. 1987, 1988, Correa and McLachlan 1991, 1992, 1994).
Although the type of interaction that occurs between endophytes and hosts is not clear, endophytic algae growing in host tissue could offer advantages in protecting its host from grazers, wave action, and desiccation, as well as reducing competition for space in costal environments (Rinkel et al. 2012). Host seaweeds, however, experience negative growth and reproduction and morphological changes through the infection of endophytes, which generally do not kill their hosts (Goff 1976, Callow et al. 1979, Schoenrock et al. 2013). There is little evidence of any mutual benefit among hosts and endophytes, and endophytes change host palatability or develop thallus toughness to inhibit the grazing activities of herbivores (Amsler et al. 2009). In Korea, the endophytic Ulvella species was recently reported in the fronds of Chondrus ocellatus (Lee et al. 2013). In the present study, we reported on the necrotic process of host Grateloupia fronds involving three endophytes, which created green spots and large holes in the fronds. Similar symptoms have been observed in Chondrus crispus fronds infected by Ulvella operculata (Bown et al. 2003).
This study found that Grateloupia infection by endophytic algae was more common in the Jeju population, followed by those of Wando and Uljin. Such differences in endophytic algal infection may closely relate to seawater temperatures, which were found to be highest at Jeju (26℃), followed by Wando (21℃), and Uljin (20℃), in terms of the average for July-September 2013 (http:// www.nfrdi.re.kr). In China, the growth of epiphytic U. leptochaete increased with temperature (9-25℃) and peaked at 25℃ (Deng et al. 2011). In the present study, endophytic algal infection was greater in the lower parts and fertile fronds of Grateloupia, which could related to frond age. However, it would be premature to make this conclusion based on this study, because of a lack of sample size and planned experimental design.
The focus of this study was to isolate three green endophytes from the host Grateloupia, culture them, and identify them for the first time in Korea, and the findings of the study could now stimulate future research in this area. The infection mechanisms of endophytic algae in host seaweeds were not examined in the present study, but it is possible that interactions between bacteria and host seaweed occur before endophytic algal infection, as has been reported by many phycologists (Correa et al. 1988, 1994, Correa and McLachlan 1994, Del Campo et al. 1998). Thus, infection experiments on isolated endophytic algae and healthy Grateloupia spp. fronds should be carried out with cultures, in order to examine the endophytic infection mechanisms and the roles of bacteria. Understanding the processes of infection of host seaweeds by endophytes is very important to the study of infectious diseases in cultivated and wild populations of edible seaweeds.