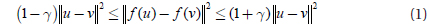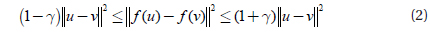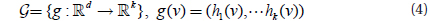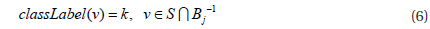A dataset can be clustered by merging the bucket indices that come from the random projection of locality sensitive hashing functions. It should be noted that for this to work the merging interval must be calculated first. To improve the feasibility of large scale data clustering in high dimensional space we propose an enhanced Locality Sensitive Hashing Clustering Method. Firstly, multiple hashing functions are generated. Secondly, data points are projected to bucket indices. Thirdly, bucket indices are clustered to get class labels. Experimental results showed that on synthetic datasets this method achieves high accuracy at much improved cluster speeds. These attributes make it well suited to clustering data in high dimensional space.
Data clustering is an important task in many areas, as such many clustering algorithms have been developed. However, many of these are neither effective nor efficient for data points in high dimensional spaces. There are several main reasons for this. Firstly, the inherent sparsity of high dimensional data hinders conventional clustering algorithms. Secondly, the distance between any two points becomes almost the same in high dimensional space [1], therefore it is difficult to differentiate similar data points from dissimilar ones. Thirdly, clusters are often embedded in subspaces of high dimensional space, and different clusters may exist in different subspaces [2].
Image clustering is a typical application of high dimensional clustering algorithms, this is because for image features nearly all the dimensions are high. So, the problem of high dimensional clustering is very apparent in image clustering applications. What’s more, the scale of an image dataset is generally large, and this scale may also change in some online applications. The performance of conventional clustering algorithms deteriorates significantly when used on this kind of dataset. For example, k-means is the mainstream cluster method in the image visual bag of words model for visual dictionary construction. However, when the number of images is large, the time taken to cluster is very long and may even be unacceptable for practical purposes. On top of this, when the image data is ever increasing, re-clustering is needed for k-means as it is not a dynamic cluster method. These limitations of k-means seriously damage its feasibility for use with large incremental image datasets. Some improvements to k-means, such as hierarchal k-means [3] and approximate k-means [4], have been presented, however, these have been shown not to support dynamic clustering [5]. Currently, widely used clustering methods such as spectrum clustering [6] and Affinity Propagation [7] methods incur high memory and computation costs due to the matrix factorization that they require. Therefore, a more efficient, new cluster method is needed.
Random projection is used in many areas including fast approximate nearest-neighbor applications [8,9], clustering [10], signal processing [11], anomaly detection [12] and dimension reduction [13]. This is largely due to the fact that distances are preserved under such transformations in certain circumstances [14]. Moreover, random projections have also been applied to classifications for a variety of purposes [15, 16].
E2LSH(Exact Euclidean Locality Sensitive Hashing) is a special case of random projection, and it was first introduced as approximate near neighbor algorithm [17]. E2LSH has attracted much attention recently, and has mainly been used in retrieval [18, 19]. In fact, the data points projected into the same buckets were found to be much more similar than those projected into different buckets. So, if we take a dataset and divide into groups according to its bucket indices, the task of data clustering has already be achieved to a certain approximation. What’s more, E2LSH is a data independent method, so it can create a dynamic index for an incremental dataset. In this way, E2LSH can be used as a dynamic clustering method. In fact, the introducer of LSH has pointed out that LSH can serve as a fast clustering algorithm, but he did this without going on to validate the claim. In fact, E2LSH has even been applied to noun clustering by Ravichandran D [20]. However, clustering image data is more difficult than text word data because of its added complexity.
Therefore, we proposed an enhanced Locality Sensitive Clustering (LSC) method for use in high dimensional space based on E2LSH. This method takes advantage of Locality Sensitive Hashing (LSH) and can cluster high dimension data at a high speed.
2. ENHANCED LOCALITY SENSITIVE CLUSTERING BASED ON RANDOM PROJECTION
The main property of random projection is dimension reduction. Compared with the classical dimension reduction algorithm PCA (Principal Component Analysis), random projection offers many benefits [21]. For example, generally PCA can’t be used to reduce the dimension of a mixture of
As for data clustering, the Johnson-Lindenstrauss Lemma shows that the distances to data points are preserved after projection. This makes it capable for use in approximate nearest neighbor search when performing information retrieval.
2.1 The distance preservation of random projection
The Johnson-Lindenstrauss Lemma is famous for its distance preservation property. It can be described like this: Given 0<ε<1 and any set
This lemma says that all pairwise distances are preserved, with high probability, up to 1±γ after mapping.
Let
Let
Imagine a set
2.2 Locality sensitive clustering
E2LSH is a special case of LSH (Locality Sensitive Hashing), and it is a random projection based method. This can be illustrated by the definition of the hashing function. The
E2LSH can also be used for data clustering. Based on the separability description above, we can say a dataset’s distance, margin and angle could be preserved after projection. These properties make E2LSH feasible for data clustering. In fact, the bucket indices found after projection can be used to group data points. This comes from the fact that similar data are arranged into a single bucket or the adjacent several buckets. So if we group these into a cluster, and further group all similar groups into corresponding clusters, the goal of data clustering can be achieved. This is the main idea behind Locality Sensitive Clustering.
E2LSH is based on the
where a is a n-dimension vector generated by the
The projection function is similar to LSH and projects points in ℝ
The enhanced Locality Sensitive Clustering (LSC) includes several main steps. Firstly, optimal parameters
Step 1, the optimal parameters k and L are calculated for a dataset S. Step 2, k dimension vector A is generated from the Gaussian distribution, b and w are also generated according the definition of the LSH function. Step 3, all the points vi∈S are projected to a k dimension bucket index Bi, and these bucket indices constitute a matrix B. Step 4, randomly selected one column Bj from matrix B, and cluster bucket indices Bj to get n clusters where j∈[l,m], m=|S| and n is the number of clusters.
where
where
3.1 Experiments on an image dataset
To verify the effect of the new clustering method on real data, we constructed an image set from the TRECVID image dataset, the set contains four categories and 75 images total. The 4 categories are ‘compere’, ‘singer’, ‘rice’ and ‘sports’.
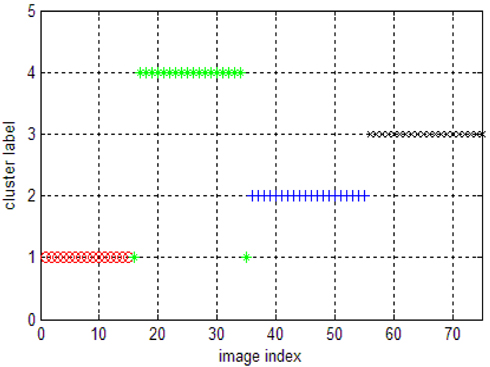
To compare the performance of our new algorithm, we first ran k-means on this image set. The results of k-means on the image dataset are showed in Fig. 1.
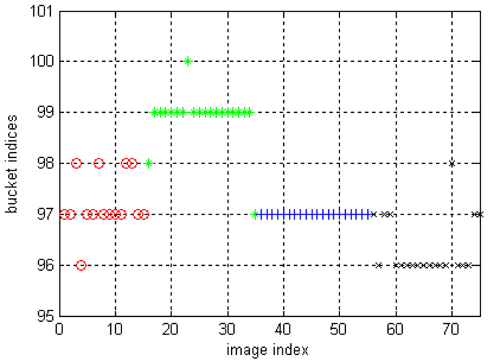
The result of LSC on the image dataset is shown in Fig. 2. Most of the cluster labels in cluster 2 and 4 are correct, the clustering labels of cluster 3 are correct while several cluster labels of cluster 1 are wrong. We can see that the accuracy of clustering labels are less than in the synthetic data, this is because the distinctness of inter-clusters are less than in the synthetic data. However, results on real data are more meaningful.
As LSC is a randomized algorithm, it is understandable that its clustering results are less accurate than k-means. On the other hand, the advantage of LSC lies in its low computation cost, fast running speed and the dynamic clustering which comes from E2LSH.
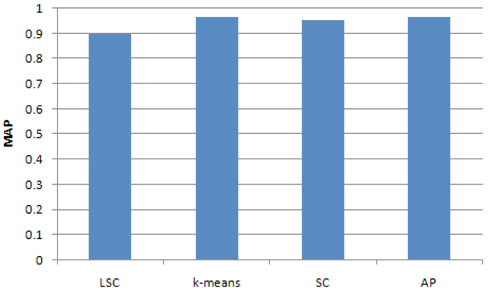
To compare the accuracy of the two methods, we computed a MAP of the clustering results for the 4 classes using LSC, k-means, Affinity Propagation (AP) cluster and Spectrum Cluster (SC) methods. The results are showed in Fig. 3. It can be seen that the accuracy of LSC is about 0.9, which is less than k-means, AP and SC.
To compare clustering times, we ran the four cluster methods (LSC, k-means, AP cluster and SC) on the image dataset. The results are showed in table 1. It can be seen that LSC consumes the least time while AP and SC cost significantly more time than LSC and k-means.
[Table. 1.] Clustering times of the four clustering methods on the image dataset

Clustering times of the four clustering methods on the image dataset
3.2 Experiments on an incremental dataset
To conveniently see the running time of our new clustering method, we construct a synthetic dataset 1 and synthetic dataset 2. Synthetic dataset 1 is an incremental dataset in scale, which contains 5 clusters of 100 dimension pieces of data, the number of data points increases from 1,000 to 10,000. Synthetic dataset 2 is an incremental dataset in dimension, which contains 5 clusters of 1,000 data points, the dimension of the points increases from 10 dimension data to 100 dimension data.
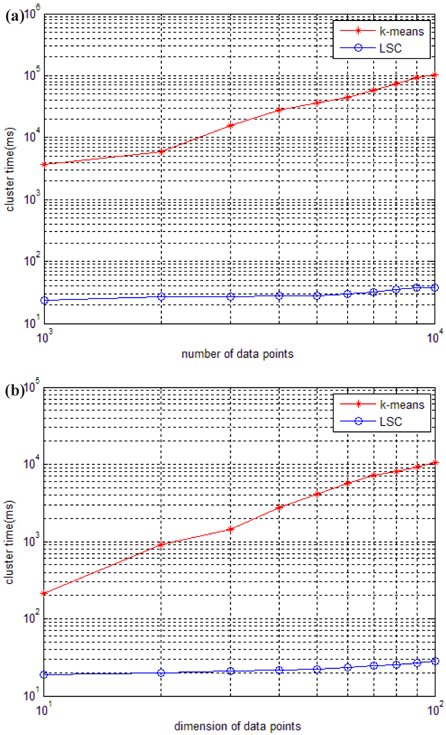
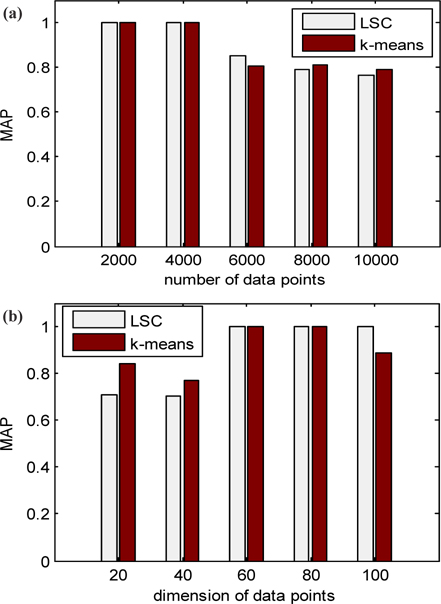
The running time of LSC and k-means for these two datasets is shown in Fig. 4. Figure 4 (a) indicates that the running time of k-means increases much more quickly than LSC when the number of data points increases from 1,000 to 10,000. When the number of data points is at 1,000, the time cost of k-means is at least 100 times more than that of LSC, and when the number of data points increases to 1,000, the time cost of k-means is at least 1,000 times more than that of LSC. Therefore, the advantage of LSC becomes more notable in larger scale datasets. In Fig. 4 (b), we can see the higher the dimensions of the data used is, the more no-table the advantage of LSC is. Even in with low dimension data of 10, the time cost of k-means is at least 10 times more than LSC. To compare the cluster accuracy, MAPs of the two methods were calculated for these two datasets. The results are shown in Fig. 5. The figure indicates that the cluster accuracy of LSC is similar to k-means, and may be better than k-means on some datasets. Therefore, LSC is more suitable for incremental datasets, and it can be a good cluster method for both high dimension and large-scale datasets.
To improve the feasibility of high dimensional data clustering, especially for use in image clustering, an enhanced Locality Sensitive Clustering method based on E2LSH is presented. This method first generates multiple hashing functions, then it projects each point using these hashing functions to get bucket indices. The bucket indices are then clustered to get class labels. In terms of clustering accuracy, experimental results show that LSC’s performance is close to k-means, AP and SC. The advantage of LSC is its fast running speed that makes it suitable for incremental clustering. This property makes it a very valuable method for large dataset clustering and especially for incremental dataset clustering.