In order to understand the diversity and genetic relationships of silkworm strains preserved in Korea, we genotyped 78 Bombyx mori strains (Bombycidae: Lepidoptera) originating from Japan, using eight polymorphic microsatellite loci. We obtained per-locus allele numbers ranging from 5 to 16 (with an average value of 9.1), per-locus observed heterozygosity ranging from 0.13 to 1.00, and per-locus polymorphic information content ranging from 0.36 to 0.77, indicating that some loci are highly variable. Phylogenetic analysis with the eight concatenated microsatellite loci showed no clustering based on known strain characteristics and origin. Nineteen strain-specific apomorphic alleles, which discriminated 16 of the 78 silkworm strains, were obtained from eight loci. These strain-specific alleles can thus be utilized for routine discrimination of strains from Japan, without any further typing of other loci. Homozygotes were also observed at some loci (27 of 118 genotypes), which can also be used to discriminate several strains by typing a few loci. These results showed that eight microsatellite loci described herein were sufficiently variable to discriminate among the 78 silkworm strains we examined, and may be useful for future investigations of this economically important species.
The domesticated silkworm is the foundation of the silk industry in many countries, including China, India, and Brazil. More than 1000 inbred lines of the domesticated silkworm
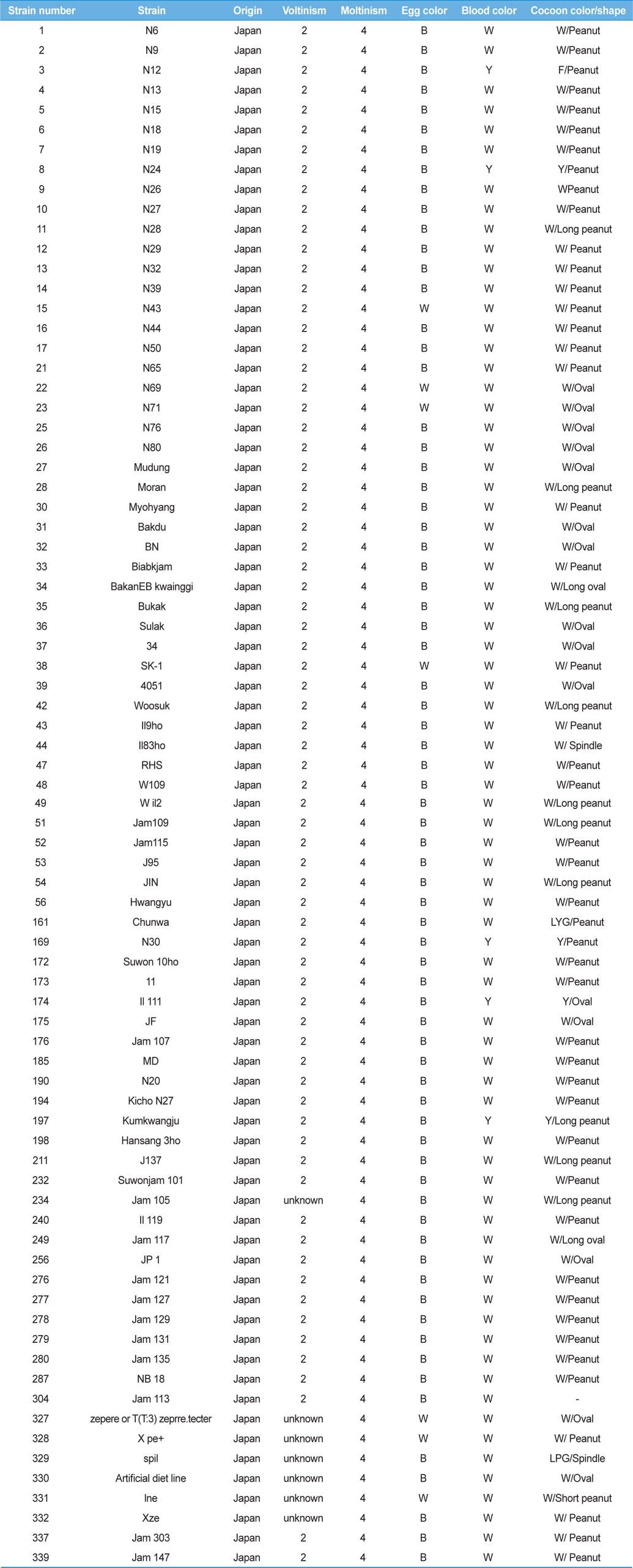
[Table 1.] General information for the Japan-origin silkworm strains utilized in this study
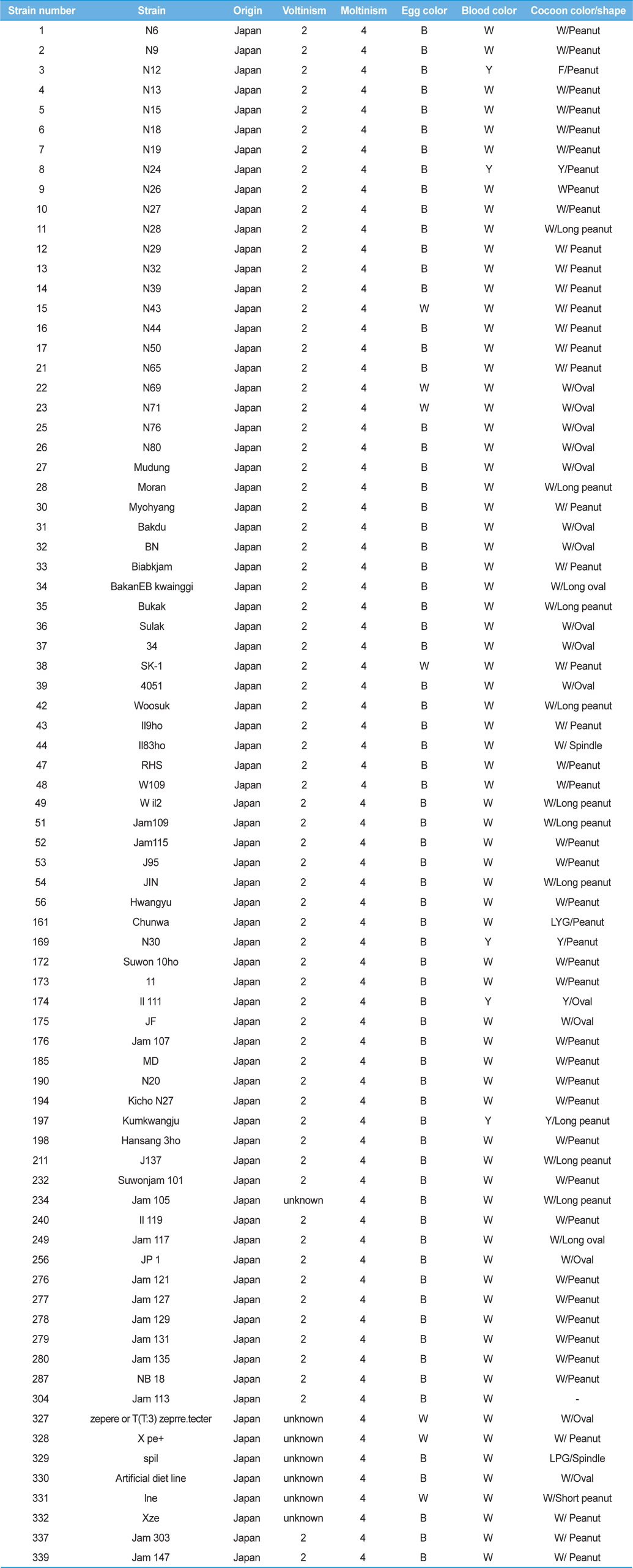
General information for the Japan-origin silkworm strains utilized in this study
Microsatellites are simple sequence repeats (SSR) of one to six bases that are abundant in both coding and non-coding regions of all eukaryotic nuclear and some prokaryotic genomes (Tautz and Renz, 1984). Due to the allelic hyper-variability of these markers, conservation in flanking sequence, and co-dominant mode of inheritance and recent advances in PCR technology, microsatellites are useful markers in several fields of sciences where detection of fine-scale genetic structure is required (Weber and May, 1989; Meglécz
Numerous microsatellite loci have been identified and described in the silkworm (e.g., Dharma Prasad
In this study, we selected eight microsatellite markers that were used in previous studies (Kim
Seventy-eight
>
Genomic DNA extraction, PCR amplification, and genotyping
Approximately 100 eggs of each strain were crushed in a glass grinder in liquid nitrogen, and genomic DNA was extracted using the DNA Extraction Kit, in accordance with the manufacturer’s instruction (Qiagen, USA). Eight microsatellite loci were successfully amplified using primers previously designed using published and unpublished
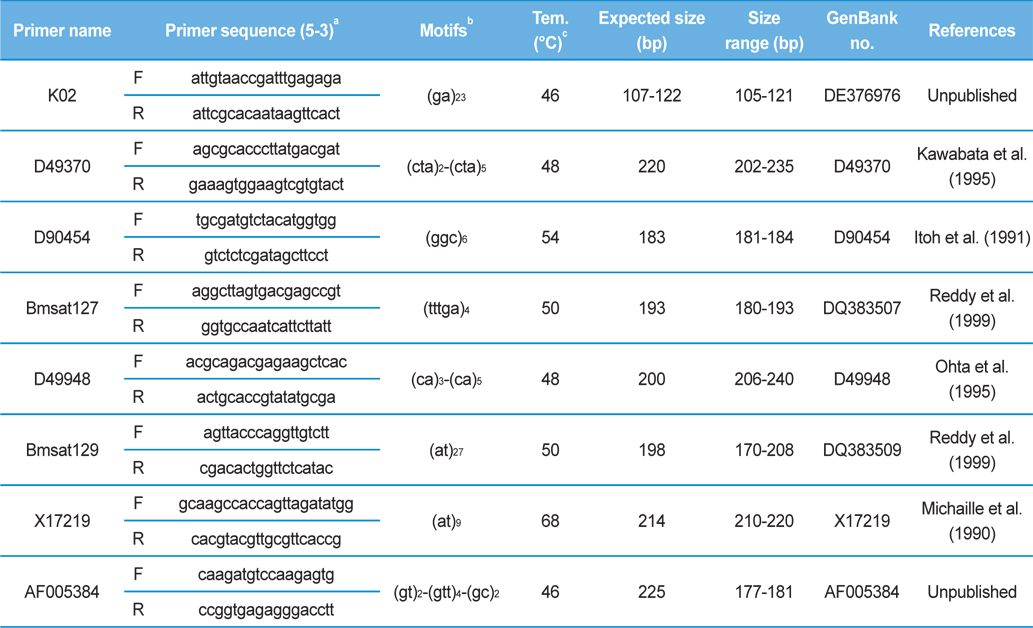
[Table 2.] Information of the 8 microsatellite loci analyzed in 78 Japan-origin silkworm strains
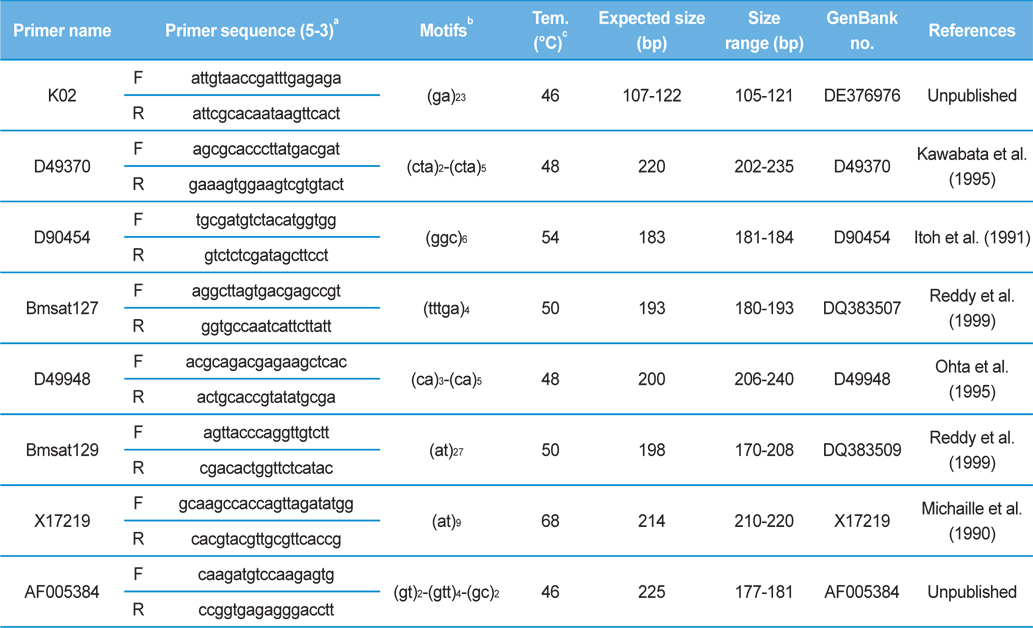
Information of the 8 microsatellite loci analyzed in 78 Japan-origin silkworm strains
PCR was carried out in a 25 μL reaction volume containing ~30 ng of genomic DNA, 200 nM of each reverse and forward primer, 200 μM of each dNTP, 2.5 μL of 10× PCR buffer [50 mM KCl, 10 mM Tris-HCl (pH 8.8), 150 nM KCl, 1.5 mM MgCl2], and 1 unit of FR-
>
Analysis of variation and phylogenetic tree construction
Observed heterozygosity (
Fragment analysis of the 78 silkworm stains in eight microsatellite loci were largely successful; however, analysis of one strain at locus D49370, five strains at locus Bmsat127, two strains at locus D49948, and four strains at locus Bmsat129, one strain at locus X17219, and one strain at locus AF005384 were not successful, either due to amplification failure or multiple amplification. Thus, 76 of the 78 strains were genotyped on average at each locus (Table 3). It is likely that the unsuccessful amplification was due to null alleles, in which a mutation occurs within the primer region, resulting in a failure to amplify the complete product, or reduction of the allele number to a single allele (Kwok
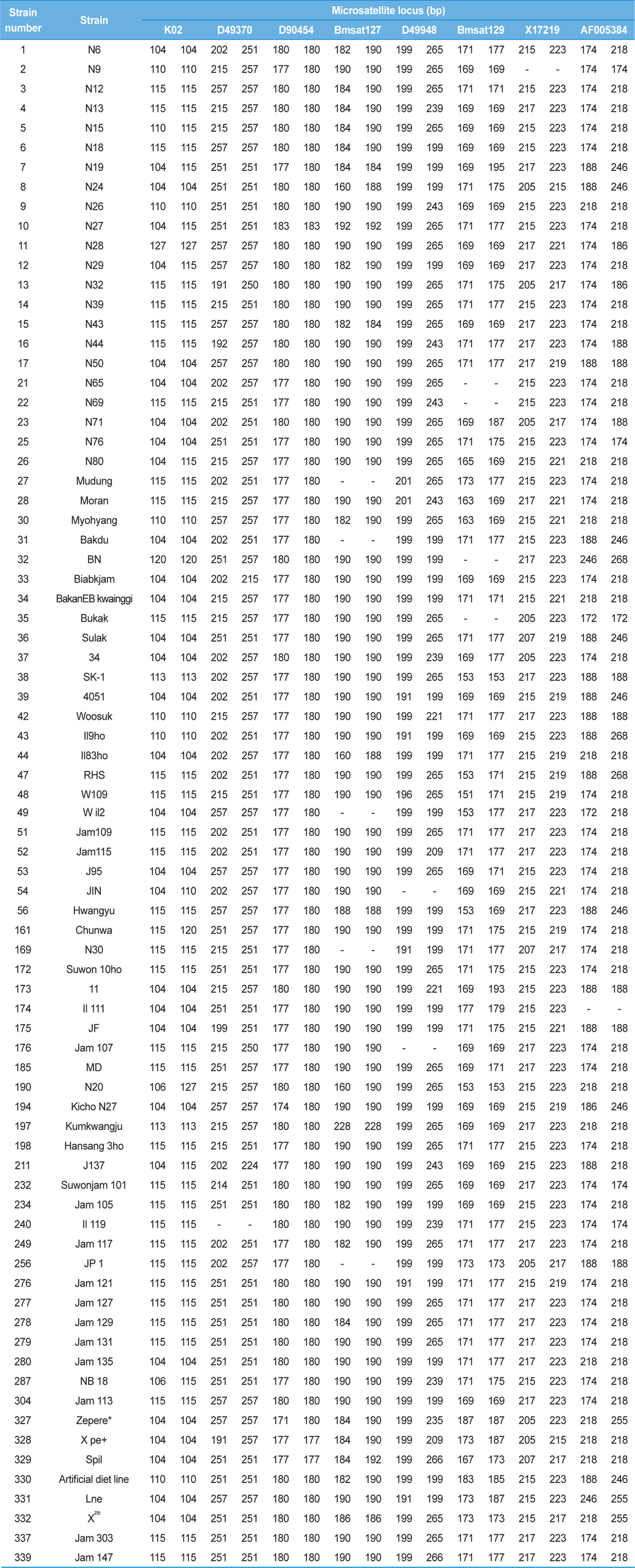
[Table 3.] Genotypes of 78 Japan-origin silkworm strains at each microsatellite locus
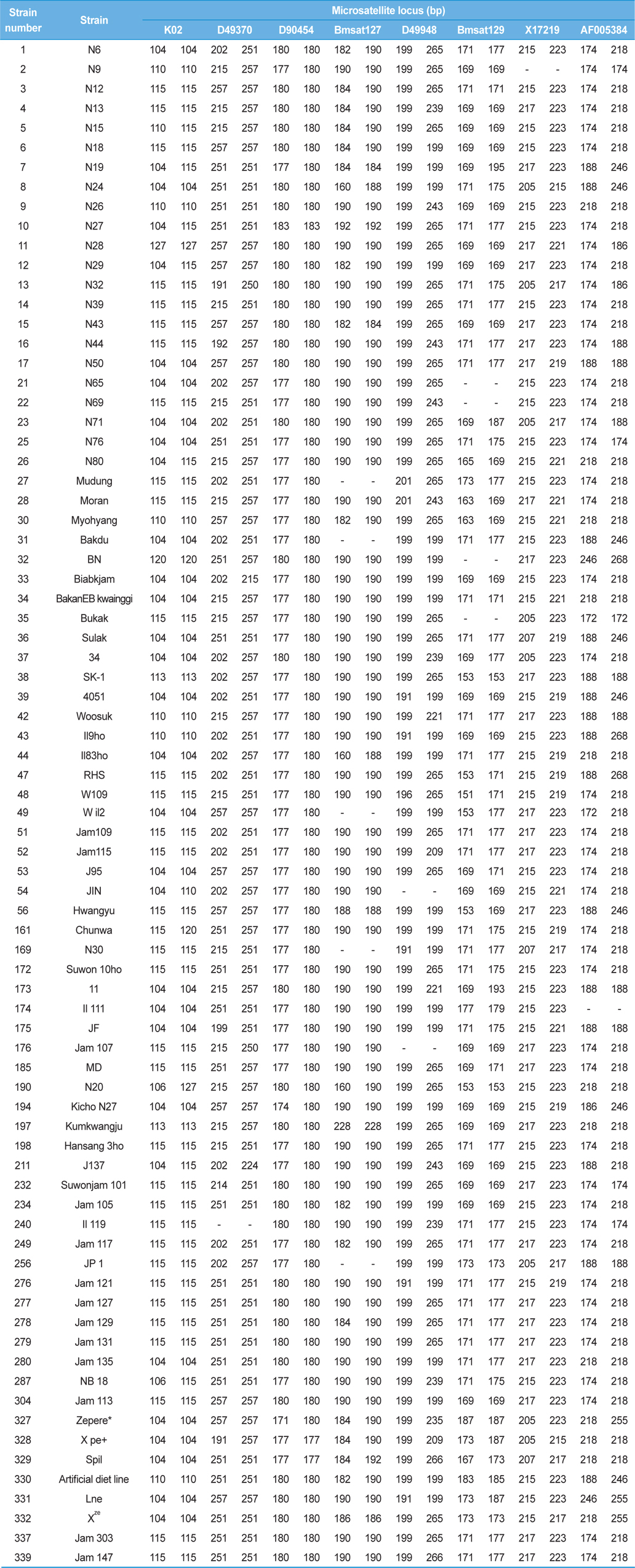
Genotypes of 78 Japan-origin silkworm strains at each microsatellite locus
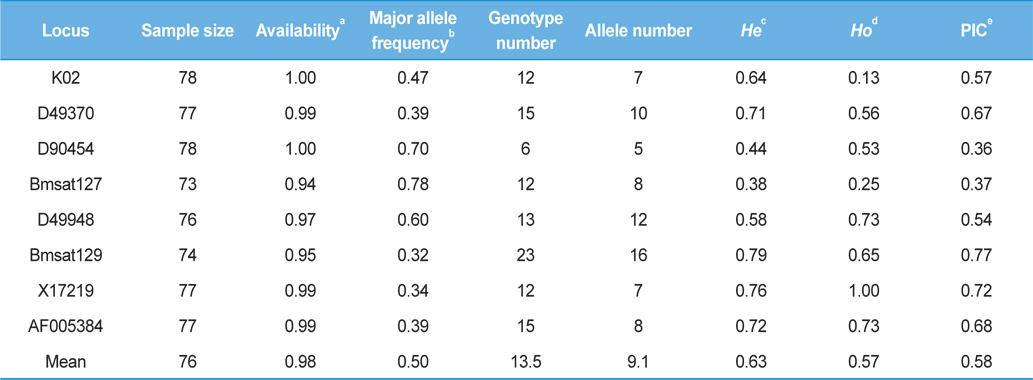
We detected 73 alleles at eight loci, and the average allele number at each locus was 9.1, ranging in number from five (locus D90454) to 16 (locus Bmsat129) (Table 4). Although the dinucleotide repeat locus Bmsat129 (Reddy
[Table 4.] Summary statistics of the 8 microsatellite loci
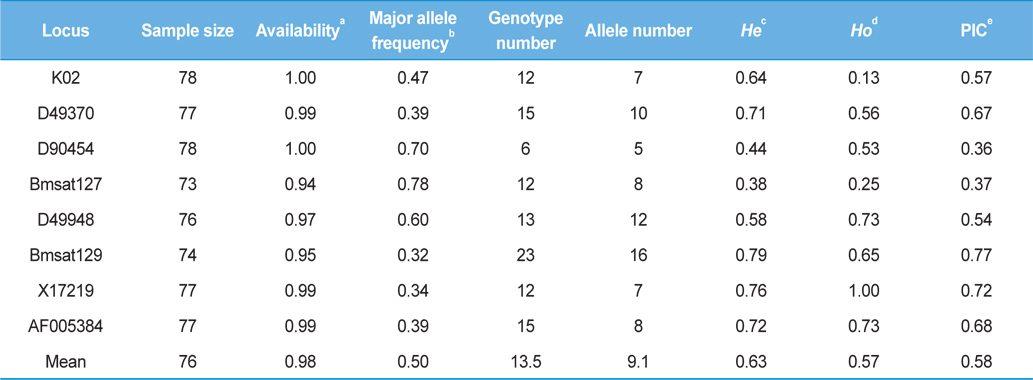
Summary statistics of the 8 microsatellite loci
In accordance with the allelic diversity, the number of genotypes was generally proportional to the allele number (Table 4). For example, the locus D90454, which exhibited only five alleles, resulted in only six genotypes in 78 strains, whereas the locus Bmsat129, which exhibited 16 alleles, resulted in 23 genotypes in 74 strains (Table 3). The frequency of the most common allele at each locus ranged from 0.32 (Bmsat129) to 0.78 (Bmsat127) (Table 4). Thus, some loci exhibited particular alleles at very high frequencies, but others did not. For example, allele 180 found at locus D90454 occurred in 75 of the 78 strains, either as a homozygote or heterozygote, but allele 177 was observed along with allele 180 in 42 strains, and mostly as a heterozygote,. The remaining four alleles, 171, 174, 177, and 183 were found along with allele 180 in only one strain, and either as a heterozygote or homozygote (Table 3). This pattern is identical to that observed in the strains from China, where the same loci were genotypes (Kim
The expected (
>
Relationships among silkworm strains
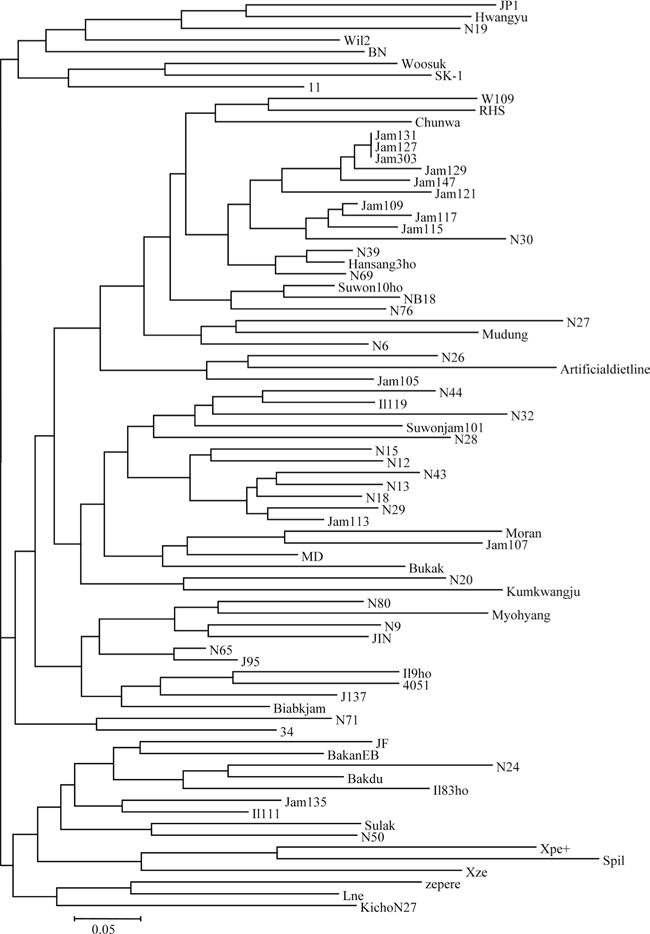
Phylogenetic analysis failed to detect any clear grouping among strains based on known characteristics, such as voltinism, moltinism, egg color, blood color, and cocoon color/shape (Fig. 1). For example, JP1, Hwangyu, N19, W il2, BN, Woosuk, SK-1, and 11 formed a clade in the NJ tree; although these strains share characteristics such as bi-voltinism, tetra-moltinism, white-colored blood, and white-colored cocoon, the egg color is either brown (most strains) or white (SK-1). Further, the cocoon shape is either peanut shaped (N19, SK-1, Hwangyu, and 11), long and peanut-shaped (Woosuk and W il2), or oval (JP1), indicating no known character-based grouping. Similar examples can be found in many other branches of the tree. Instead, the clustering pattern indicates that the microsatellite loci typed in this study may reflect genetic differences among strains that can be utilized for the discrimination of silkworm strains. Previous studies of silkworm strains originating from China also support this notion (Kim
The eight microsatellite loci exhibited the presence of strainspecific alleles (Table 3). Locus D49370 provided the alleles 192, 199, 214, and 224, which were unique to strains N44 (no. 16), JF (no. 175), Suwonjam (no. 232), and J137 (no. 211), respectively (Table 3). Similarly, the locus D90454 exhibited alleles 171, 174, and 183, which were unique to strains zepere (no. 327), Kicho N27 (no. 194), and N27 (no. 10), respectively. The locus Bmsat127 exhibited alleles 186 and 228, unique to strains Xze (no. 332) and Kumkwangju (no. 197), respectively. Locus D49948 exhibited alleles 196 and 235, unique to strains W109 (no. 48) and Zepere (no. 327), respectively. Locus Bmsat129 exhibited alleles 151, 165, 167, 183, 193, 195, 179, and 185, unique to strains W109 (no. 48), N80 (no. 26), Spil (no. 329), Artificial diet line (no. 330), 11 (no. 173), N19 (no. 7), Il 111 (no. 174), and Artificial diet line (no. 330), respectively (Table 3). In total, 19 apomorphic alleles were observed, which discriminated 16 of the 78 silkworm strains. These strain-specific alleles can thus be utilized for the discrimination of
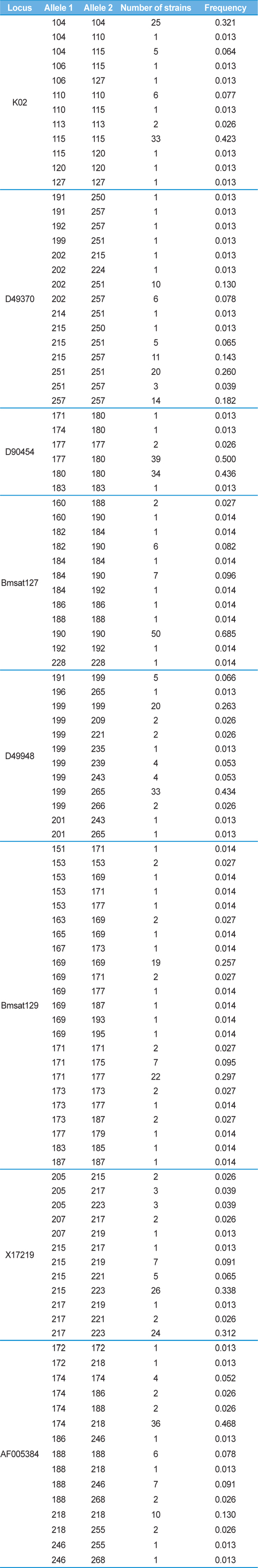
Genotyping results showed that a substantial number of strains possessed homozygotic alleles (Table 5). At locus K02, 25 strains were homozygous for allele 104, six strains were homozygous for allele 110, two strains were homozygous for allele 113, 33 strains were homozygous for allele 115, one strain was homozygous for allele 120, and one strain was homozygous for allele 127 (Table 5). Similarly, 34 strains were homozygous at two alleles for locus D49370, 37 strains were homozygous at three alleles at locus D90454, 55 strains were homozygous at six alleles at locus Bmsat127, 20 strains were homozygous at one allele at locus D49948, 26 strains were homozygous at five alleles at locus Bmsat129, and 21 strains were homozygous at four alleles at locus AF005384 (Table 5). Consequently, 27 of the 118 genotypes were homozygous at one or more of the eight loci. Because we used ~100 eggs for the extraction of DNA from each
[Table 5.] Genotype frequency in each microsatellite marker among 78 silkworm strains
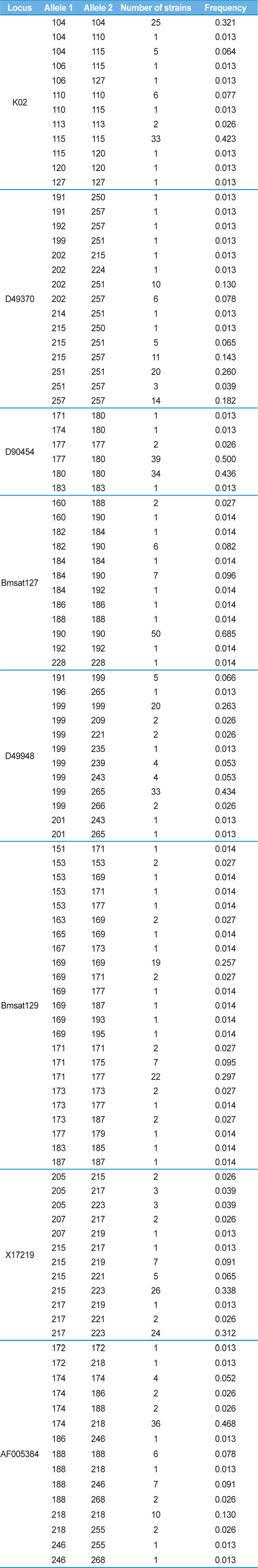
Genotype frequency in each microsatellite marker among 78 silkworm strains
Previously, 85 silkworm strains originating from China were genotyped using the same microsatellite loci, and exhibited 22 strainspecific apomorphic alleles, which discriminated 19 of the 85 strains. Furthermore, a substantial number of homozygous strains were observed. The previous and current results support the continued use of these microsatellite loci for the discrimination of silkworm strains originating from China and Japan that are under preservation in Korea. Nevertheless, the isolation of additional microsatellite loci is needed to discriminate among the >300 silkworm strains that are currently preserved in Korea.