Mycobacteria are pleomorphic, aerobic, Gram-positive, acid-fast, non-motile rods, 0.2-0.6 μm in diameter and 1-10 μm long in the genus Mycobacterium, the only genus in the family Mycobacteriaceae (Gauthier and Rhodes, 2009). The Mycobacterium comprises obligate pathogens that cause serious human and animal diseases, opportunistic pathogens, and saprophytic species (Bland et al., 2005).
Piscine mycobacteriosis is a systemic infectious disease that is typically a subacute to chronically progressive condition caused by different Mycobacterium species, including Mycobacterium marinum, M. fortuitum, M. chelonae, and M. abscessus (Kusunoki and Ezaki, 1992; Decostere et al., 2004). More recently, additional mycobacterial species have been implicated as the causes of mycobacterial infections, including M. shottsii, M. montefiorense, and M. haemophilum (Levi et al., 2003; Rhodes et al., 2003; Whipps et al., 2007). Although mycobacteriosis is usually a chronic disease and may not produce external lesions, the internal lesions typically include enlargement and typical grey or white nodules of the spleen, kidneys, and liver (Chinabut, 1999; Gauthier and Rhodes, 2009).
This study reports on mycobacteriosis in an imported tropical ornamental fish, Macropodus opercularis, commonly known as the paradise fish.
Forty paradise fish (body weight, 1.8 ± 0.4 g) were bought from an ornamental fish wholesaler in Korea in June 2008. After 2 days of acclimation, the first fish death was recorded, and all of the fish died by July (cumulative mortality rate, approximately 100% per month). Few of the diseased fish showed any external signs, but enlargement of the spleen and discoloration of the liver were observed after postmortem examination. Moribund and freshly killed fish were examined using histopathology and microbial cultures after euthanasia with excessive 2-phenoxyethanol (Sigma, St Louis, MO, USA). The gill and body surfaces were examined microscopically for the presence of parasites.
The spleen was collected from five moribund fish and fixed immediately in 10% neutral buffered formalin. After fixation, standard histological procedures were used for tissue dehydration and paraffin embedding. Tissue sections were stained with hematoxylin and eosin (H&E) and Ziehl-Neelsen (ZN).
The liver, kidney, and spleen of moribund and freshly killed fish were inoculated into tryptone soy agar (TSA; Difco, Sparks, MD, USA) and incubated at 30℃ for 72 h. Frozen kidney and spleen tissues were homogenized in sterile 0.85% saline (w/v), decontaminated with 0.35% hexadecylpyridinium chloride for 30 min, and then centrifuged at 2,800 g for 30 min (Corner and Trajstman, 1988). The supernatant was removed, and the sediment was washed twice with 1 mL of sterile phosphate-buffered saline (PBS). The sediments were inoculated on Middlebrook 7H10 (Difco), and the medium was incubated aerobically at 30℃ and checked daily for 2 months. Before decontamination, samples were also cultured in TSA (Difco) to detect common bacterial fish pathogens and incubated for 72 h at 30℃.
For PCR, genomic DNA was extracted from frozen spleen stored at ?80℃ using an AccuPrep Genomic DNA purification kit (Bioneer, Daejeon, Korea). Molecular detection of Mycobacterium species was based on 16S ribosomal DNA (16S rRNA) and 65-kDa heat-shock protein-encoding gene (hsp65) sequences. The primers for the 16S rRNA (T39 and T13; PCR product size, 924 bp) (Talaat et al., 1997) and hsp65 (Tb11 and Tb12; PCR product size, 441 bp) (Aranaz et al., 2008) genes were used for amplification and sequencing. The DNA was amplified with an initial 5-min denaturation at 95℃, followed 166by 35 cycles of 94℃ for 30 s, 55℃ (for 16S rRNA) or 58℃ (for hsp65) for 30 s, and 72℃ for 1 min, with a final 5-min extension at 72℃. The sequencing reactions were carried out with a BigDye terminator ver.1 cycle sequencing kit (Applied Biosystems, Foster City, CA, USA), and the products were sequenced on an ABI PRISM 3130xl Genetic Analyzer (Applied Biosystems). 16S rRNA and hsp65 DNA sequences for Mycobacterium species available from GenBank plus our sequences were aligned using ClustalX ver. 1.83 (Thompson et al., 1997). Phylogenetic trees were obtained using the neighbor-joining method with Kimura’s two-parameter distance correction model with 1,000 bootstrap replications using MEGA ver. 4 (Tamura et al., 2007).
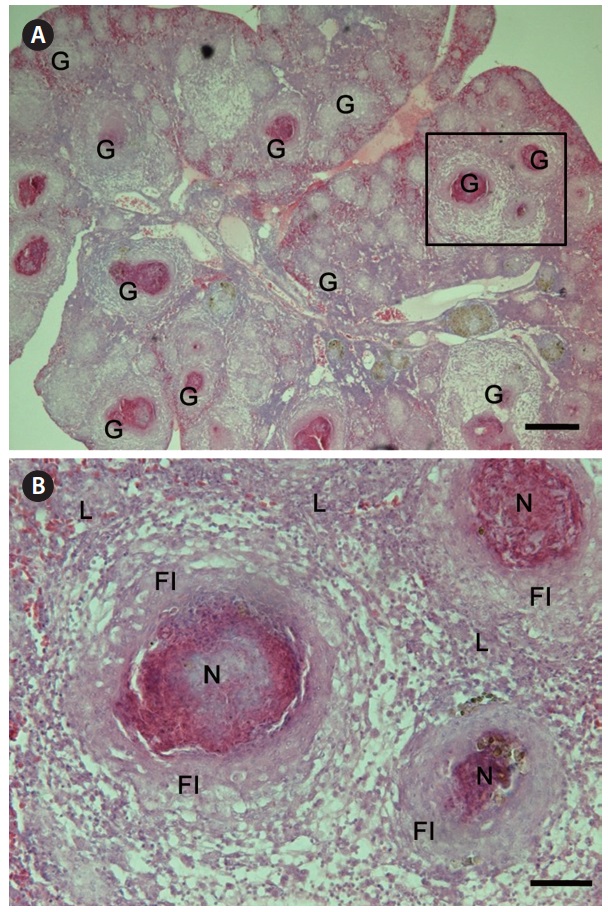
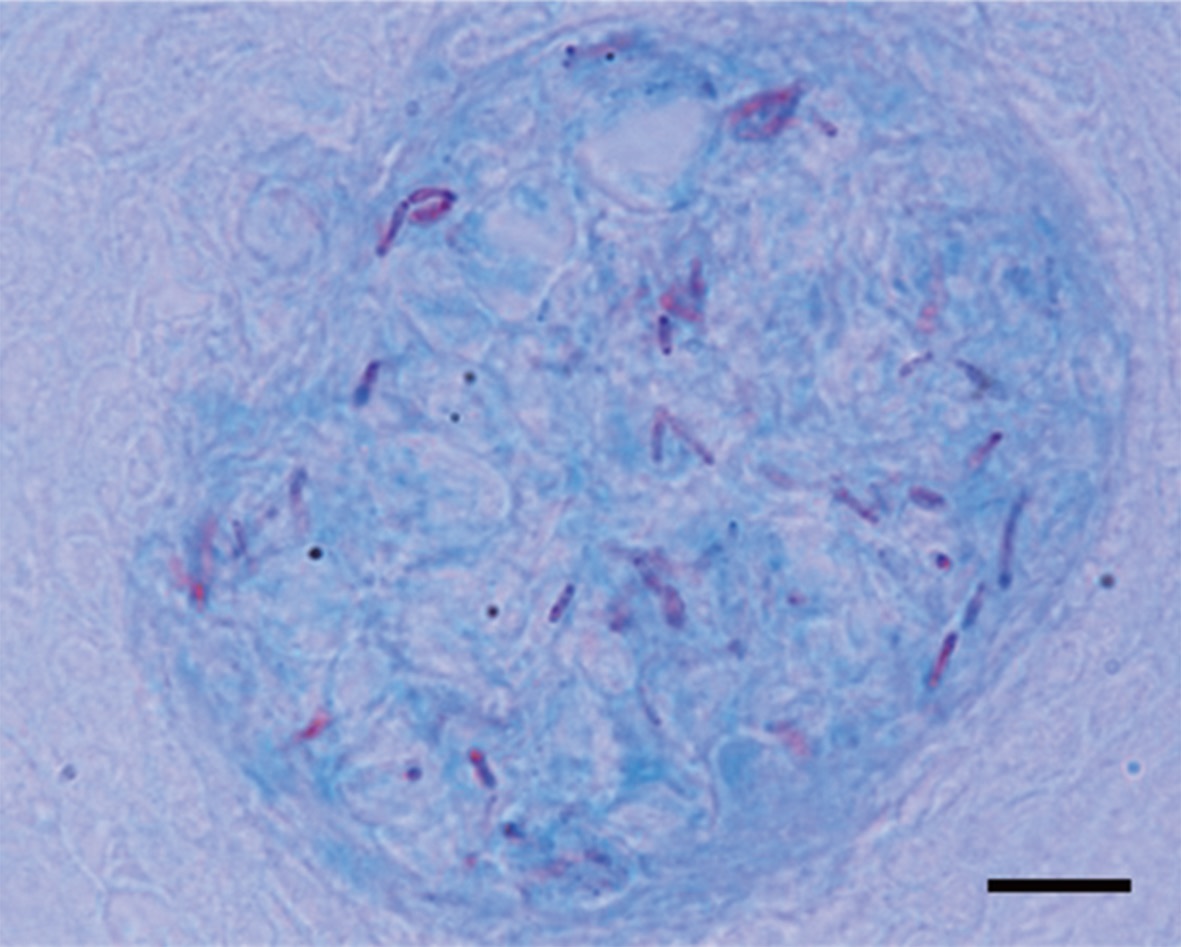
Granulomas 20-57 μm in diameter were observed in the H&E-stained spleen sections of all individuals examined (Fig. 1A). Numerous granulomas appeared in the splenic pulp and sheathed tissue (Fig. 1A and 1B). Severe necrosis was observed in the central eosinophilic area of the granulomas, surrounded by extensive fibrous infiltration containing several layers of epithelioid cells (Fig. 1B). When stained with ZN, numerous acid-fast bacilli were found in the centers of the granulomas in the spleen (Fig. 2).
Two fast-growing bacteria, Citrobacter freundii and Pseudomonas sp., grew on TSA, but they were isolated from one fish only. Histopathological examination led us to suspect mycobacteriosis, so we attempted bacterial isolation using Middlebrook 7H10 from frozen paradise fish tissue. However, no mycobacterial growth was observed after incubation for 2 months.
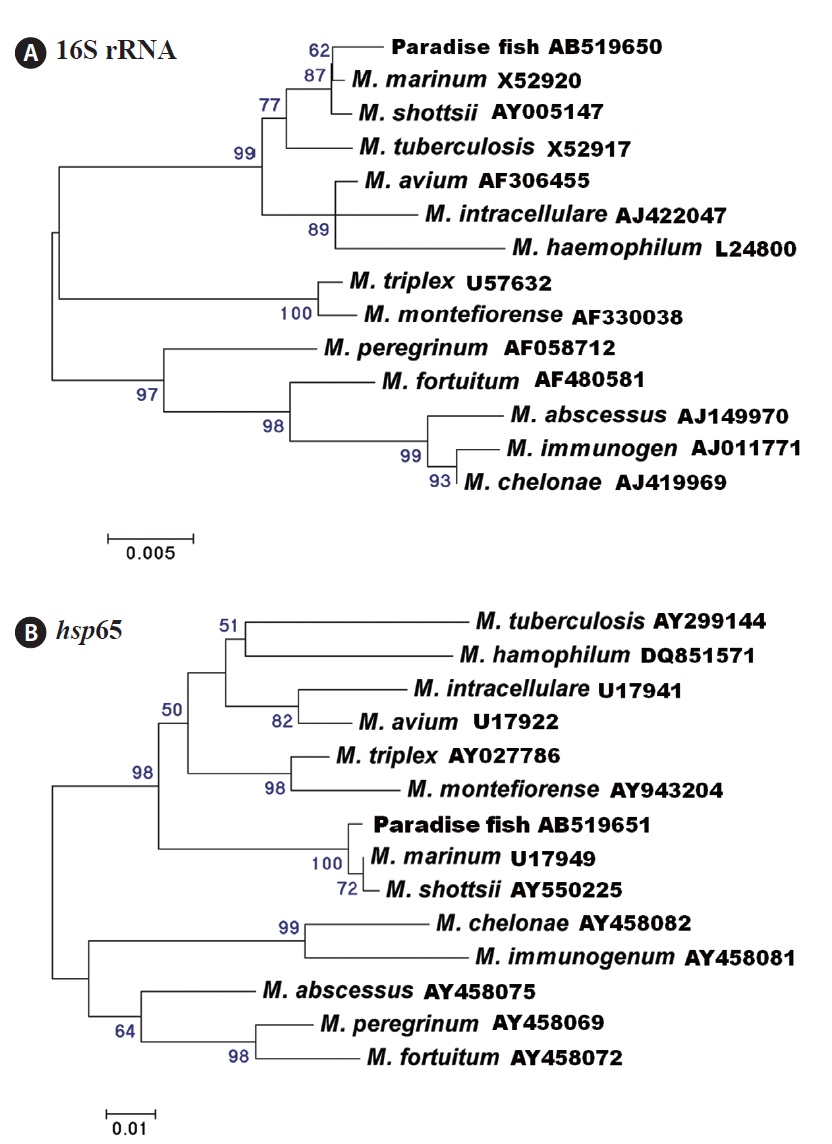
Thirteen (62%) and 14 (67%) of 21 fish were positive for mycobacteriosis, determined by PCR targeting the 16S rRNA and hsp65 genes, respectively. All of the samples positive for the 16S rRNA gene were also positive for the hsp65 gene. Analysis of the 16S rRNA (838-bp product) and hsp65 (380-bp product) DNA sequences revealed the greatest similarity with those of Mycobacterium marinum retrieved from the NCBI database (GenBank accession nos. AF456239 [16S rRNA] and AF456471 [hsp65]). Phylogenetic trees based on the 16S rRNA and hsp65 genes showed that the strains isolated in this study were related most closely to M. marinum (Fig. 3). The nucleotide identities of the 16S rRNA and hsp65 genes with those of M. marinum (M. shottsii) were 99.5% (99.4%)
and 99.4% (99.1%), respectively.
Several Mycobacterium species have been identified as ornamental fish pathogens (Lescenko et al., 2003). The paradise fish is also susceptible to Mycobacterium infection. For example, Gomez (2008) found granulomatous inflammation and numerous acid-fast bacteria in tissue sections of granulomas from a diseased paradise fish, but the causative agent was not confirmed at the species level. Additionally, Marci et al. (2008) reported M. fortuitum infection in severely emaciated paradise fish, with ascites and granulomas of various sizes in the internal organs. In our study, the 16S rRNA and hsp65 gene analysis show that the amplified genes were related most closely to M. marinum. However, it is difficult to distinguish among M. marinum, M. shottsii, M. pseudoshottsii, and M. ulcerans using 16S rRNA and hsp65 gene sequences (Adekambi and Drancourt, 2004). Additional research to identify the Mycobacterium sp. infecting the paradise fish is needed.
We could not culture Mycobacterium sp. from paradise fish, as freezing the spleens and kidneys affected the viability of the mycobacteria, resulting in negative cultures on Middlebrook 7H10. Although bacterial isolation is considered the gold standard for diagnosing bacterial disease, the isolation of mycobacteria from field samples is generally problematic and sometimes not even attempted because mycobacteria are slow-growing fastidious organisms, and isolation can be adversely affected by other microbial contamination (Pate et al., 2005). Instead, mycobacteria have been detected by amplifying various target genes, such as 16S rRNA, IS986, IS611, and hsp65 (Kolk et al., 1992; Telenti et al., 1993; Kox et al., 1995; Noordhoek et al., 1995), and PCR has been used to detect mycobacterial infection in several studies (Frevel et al., 1999; Astrofsky et al., 2000). In our study, PCR amplification targeting the 16S rRNA and hsp65 genes was carried out, and more than 60% of the examined fish showed positive reactions for both genes. However, the sensitivities of the 16S rRNA and hsp65 PCR primers may differ, and the hsp65 PCR primers may have been more sensitive than the 16S rRNA PCR primers in this study.
Although mycobacteriosis was confirmed in imported paradise fish, the etiological agent of the mass mortality is unclear, as Mycobacterium sp. was detected only in about 65% of the fish by PCR, and megalocytivirus (10%, four of 40 fishes) and reovirus (30%, three of 10 fishes) were also detected in the diseased paradise fish by PCR (data not shown). Further studies of the pathogenicity in paradise fish are necessary. However, our results suggest that mass mortality in paradise fish was caused by multiple infections with Mycobacterium sp., megalocytivirus, and reovirus.
Over 1 billion ornamental fish are traded internationally each year (Whittington and Chong, 2007). Paradise fish are
small freshwater labyrinth fish found in ditches and paddy fields in Southeast Asia (Man and Hodgkiss, 1981). Paradise fish are imported into Korea from Indonesia, Malaysia, and Vietnam (National Fisheries Products Quality Inspection Service, personal communication), but there are no clear guidelines or checks on paradise fish suspected of having mycobacteriosis. This is made more difficult given the lack of obvious signs of sickness in diseased fish. In this study, it is likely that the paradise fish monitored were infected before importation to Korea, as mortality occurred during the acclimation period, and mycobacteriosis is generally a chronic, slowly developing disease.
In conclusion, we confirmed the presence of mycobacteriosis in paradise fish imported from Southeast Asia into Korea. Considering the broad host spectrum of Mycobacterium sp. and increasing global international trade in ornamental fish, stricter quarantine inspection guidelines are necessary to reduce the risk of mycobacteriosis in fish in Korea.