Various clinical and epidemiological studies have associated increases in morbidity with ambient air concentrations of particulate matter (PM) [1]. Ambient air pollution includes soluble or free transition metals, such as Fe, Cu, Ni, V, Co, and Cr in different amounts, forms, and oxidative states [2]. It is known that the metal-mediated generation of reactive oxygen species (ROS) can cause severe oxidative stress within cells or tissues through the oxidation of nucleic acids, proteins, and lipids [2,3]. Especially, Fe can be involved in the generation of highly reactive hydroxyl radical (OH·) in the presence of O2 (or H2O2) and/or a reductant, (such as ascorbate or cysteine) [3]. Other metals (Cu, Ni, Co, Cr, and V) have been known to become redox-active under certain conditions, causing oxidative damage to biomolecules [2,4].
It is widely believed that H2O2-induced damages to cellular DNA are mediated by transition metal ions, mainly iron and/or copper, which are able to catalyze the formation of OH· by Fenton- type reactions [5]. There are also ample evidences to support the notion that the most important mechanism of oxidative damage to proteins is metal-catalyzed oxidation [5,6]. This highly reactive free radical attacks neighboring amino acid residues, some of which are converted to carbonyl-containing derivatives [7,8], which have been established to accumulate on tissue proteins during aging and disease development [8]. Indeed, exposure of humans to PM has been associated with acute lung injury and development of lung diseases, as well as cardiovascular diseases [9,10]. Further epidemiological studies showed a stronger correlation between respiratory diseases and smaller particulates, such as those with an aerodynamic diameter of less than 2.5 μm (PM2.5) [11]. However, there has been no specific study on how particle size influences the manner in which oxidative damage affects biomolecules, such as DNA and proteins.
The goal of this study was to compare the metal-induced oxidative damages incurred by biomolecules through PM2.5 exposure with those of PM10, each of which was collected in Seoul metropolitan area. The oxidative damage caused by a standard reference material (SRM) 1648 was also investigated and compared with the results from PM2.5 and PM10 [12].
2.1. Particulate Matter and the Determination of Mobilized Amounts of Metals
SRM1648 was purchased from the National Institute of Standard & Technology. SRM1648 is an urban air particulate matter sample, in which approximately 50% of the particulates are less than 10 μm in diameter (and the other 50%). Collection of ambient particulate matter within Seoul metropolitan area and particle preparation was described in our previous study [13], which showed the amount of mobilized (and thus bioavailable) metals from both PM2.5 and PM10. Briefly, ambient PM was collected within Seoul metropolitan area using a high-volume air sampler (Andersen Inc., Idaho Falls, ID, USA), designed to collect particles into two fractions, PM10 and PM2.5, at a flow rate of 1.2 m3/min, using a quartz microfiber filter (20.3?25.4 cm; Whatmann, Maidstone, Kent, UK). The collection filter was soaked in Chelex-treated ultrapure water and was sonicated 10 times at 3 A for 30 sec with ultrasonic generator (Sun-il Co., Hwaseong, Korea) prior to use. The suspended solution was centrifuged at 50 g for 10 min to remove filter fiber. The supernatant was then dried with a speed vacuum-dryer (Savant Co., Cumbria, UK), and was thoroughly blended with a bead beater before use. Total and mobilized amounts of 9 metals (Co, Cr, Cu, Fe, Mn, Ni, Pb, V, and Zn) in particulate matter were determined by inductively coupled plasma mass spectrometry (Sciex ELAN 5000; PerkinElmer, Waltham, MA, USA) or inductively coupled plasma atomic emission spectrometry (Optima 3000 XL, PerkinElmer).
2.2. Effect of Mobilized Metals on the Induction of DNA Single Strand Breaks
Metals were mobilized by preincubation of particulate matter (1 mg/mL) in 50 mM NaCl (pH 7.5) with or without citrate or ethylenediaminetetraacetic acid (EDTA, 1 mM) in the presence or absence of ascorbate for 24 hr on a wrist-action shaker in the dark. The supernatants that contained only mobilized metals were examined for their abilities to catalyze the formation of DNA single strand breaks (SSBs). Supernatants containing mobilized metals from SRM1648, PM2.5, or PM10 were incubated with ?X174 replicative form I (RFI) DNA (0.5 μg) for 30 min. Because EDTA binds iron and could potentially compete with the other chelators being studied, tracking dye without EDTA was added to the samples. Samples were then loaded into a 0.6% agarose gel and were subjected to electrophoresis for approximately 2 hr to separate closed-circular, superhelical (form II) DNA from DNA with SSB (form I). The negatives were then analyzed using an AlphaImager model 1220 and AlphaEase stand-alone software (Alpha Innotech Co., San Leandro, CA, USA). The area obtained upon integration of the peaks for the representative forms of DNA (i.e., I and II) were then used to calculate the percentage of DNA with SSB in each lane. All results were reported as the percentage of SSB DNA ± standard deviation relative to the control DNA. To determine the effect of desferrioxamine (DFO), a specific chelator of iron [14,15], on the formation of DNA SSB, DFO (10 mM) was preincubated with particles (1 mg/mL) for 24 hr prior to the addition of citrate or EDTA.
2.3. Effect of Mobilized Metals on the Formation of Protein Carbonyl
Treatment of PM or pretreatment of PM with DFO for the experiment of protein carbonylation were conducted with the same method as that described above. All experiments upon biomolecules were carried out at room temperature (25℃) in a darkroom lighted by incandescent red lights, so as to prevent photochemical reduction of the metals [12]. All results were reported as the percentage of carbonyl relative to that of the control (i.e., untreated) protein. The protein carbonyl content was determined by the reaction with 2,4-dinitrophenylhydrazine (DNPH) as described by [7].
For measurement of cytosolic protein carbonyl formation, each particle was suspended in sterile 14 mM NaHCO3 (pH 7.4) immediately before use, vortexed for 1 min, and diluted to a concentration of 100 μg/cm2 with the complete growth medium. The pH of the suspended particle was adjusted to 7.5. The cells were cultured in flasks until they were approximately 80% confluent, and then dislodged with trypsin-EDTA, resuspended in complete growth medium, counted using a hemocytometer, and plated at a cell density of 20,000 cells/cm2. The cells were then allowed to recover for 24 hr, after which they were treated with the suspended particle for 24 hr, as indicated above. The particle that was not associated with cells and/or endocytosed was then removed by centrifugation. The cells were rinsed once with 0.15 M phosphate-buffered saline (pH 7.4) and dislodged with 0.25% trypsin minus EDTA in this step because EDTA is known to a strong metal chelating agent. The cells were then lysed by repeating freezing in liquid nitrogen and thawing at 37℃. After three cycles, the cell lysate was centrifuged at 12,000 g for 30 min at 4℃, in a Beckman L8-80M ultracentrifuge (Beckman Coulter, Fullerton, CA, USA), to remove the particulate matter and cell debris. A portion of 12,000 g supernatant was used to determine the cytosolic protein carbonyl content. Each experiment conducted in this study was repeated 3 times or more, and the results were reported as mean values ± standard deviation.
3.1. Effect of Mobilized Metals on the Induction of DNA SSBs
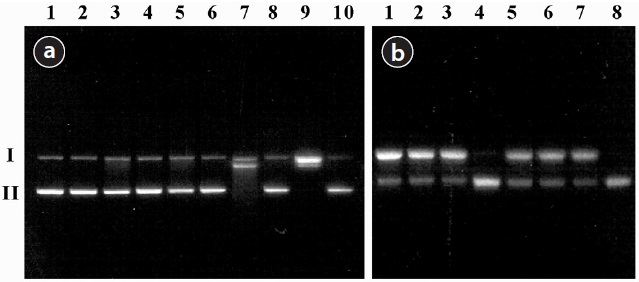
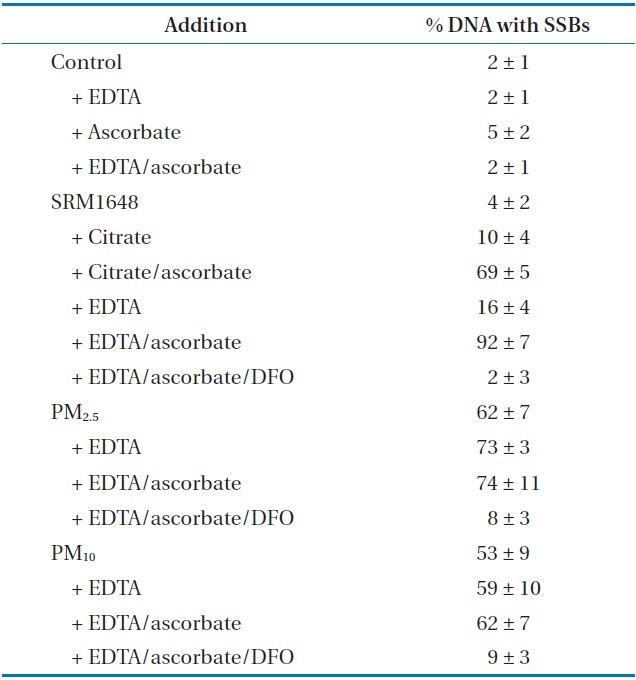
The values of SSBs induced by the treatment of PM are shown in Table 1. The agarose gel electrophoretic pattern of DNA SSBs by SRM1648 is shown in Fig. 1(a), where no significant DNA SSB was observed with various controls without the particle (Fig. 1(a), lanes 1, 2, 3, and 5; approximately 2%). The addition of SRM1648 alone did not increase DNA SSB greatly (Fig. 1(a), lane 4; approximately 4%). Then, the addition of citrate or EDTA together with SRM1648 increased somewhat DNA SSBs (Fig. 1(a), lanes 6 and 8; approximately 10% and 16%, respectively). However, significant increases in DNA SSB were observed with the simultaneous addition of SRM1648, ascorbate, and citrate or EDTA (Fig. 1(a), lanes 7 and 9; approximately 69% and 92%, respectively). Thus, the formations of DNA SSBs in the presence of EDTA were higher than those in the presence of citrate. The percentages of DNA SSB with EDTA were about 1.5-fold higher than those of citrate. The specific chelator, DFO, was preincubated with the particle to determine whether the metals from the particles were responsible for the formation of DNA SSBs. The addition of the specific chelator resulted in an almost complete inhibition of the mobilized metal-dependent formation of DNA SSB (Fig. 1(a), lane 10; Table 1).
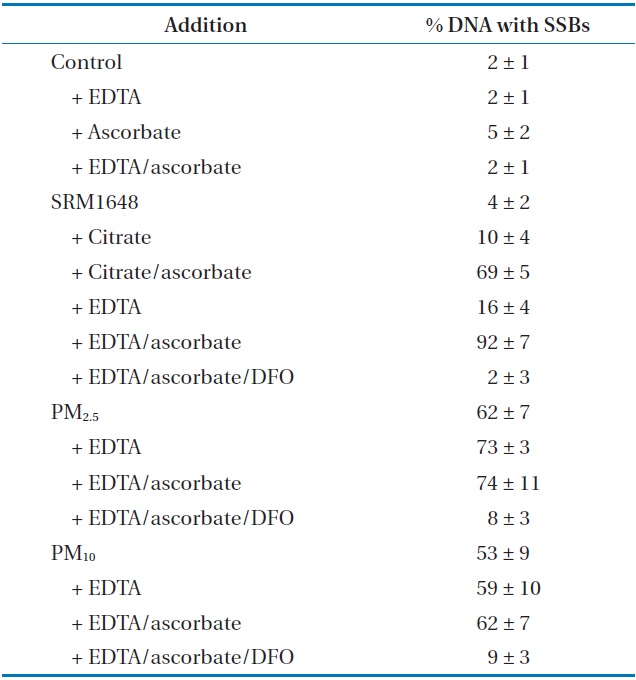
Induction of DNA SSBs catalyzed by mobilized metals from SRM1648 and PMs collected in Seoula
The DNA SSB formation results for the experiment of PM collected in Seoul, and the agarose gel electrophoretic pattern of DNA SSB, are shown in Table 1 and Fig. 1(b), respectively. The addition of PM2.5 and PM10 to the sample containing ?X174 DNA resulted in the significant formation of DNA SSB of by 2% and 53%, respectively. These results indicate that PM2.5 generated more oxidative damage to DNA than PM10. The addition of EDTA or EDTA/ascorbate to the reaction mixture containing PM2.5 or PM10 did not increase the extent of DNA damage by any substantial amount, possibly due to the high level of DNA damage which had already taken place due to contact with PM (Table 1). However, the extent of DNA SSB formation due to PM2.5 was generally higher than that due to PM10. The addition of DFO to the reaction mixture containing PMs, EDTA, and ascorbate greatly reduced the formation of DNA SSB, as has been observed with SRM1648 (Fig. 1(b), lanes 4 and 8; Table 1).
3.2. Effect of Mobilized Metals on the Formation of Protein Carbonyl
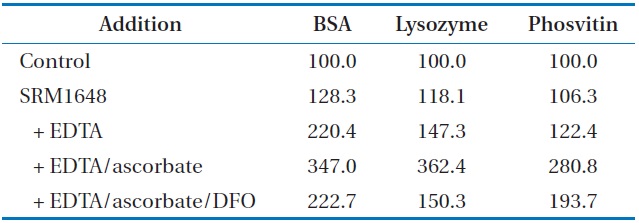
To determine the damaging effect of the metals mobilized from PMs, carbonyl formation of proteins was also investigated using 3 proteins (bovine serum albumin [BSA], lysozyme, and phosvitin). The incremental addition of SRM1648, EDTA, and ascorbate incurred increasingly higher percentages of carbonylation to the proteins (Table 2). The extents of carbonyl formation in the presence of SRM1648 or/chelator/reductant were 1.6, 2.5, and 2.3 times higher for BSA, lysozyme, and phosvitin, respectively, than for those in the presence of SRM1648/chelator, showing that each protein has different susceptibilities toward the same amount and kind of reagents. Unlike studies with DNA SSB, treatment of DFO to the reaction mixture containing SRM1648/EDTA/ascorbate did not significantly inhibit the carbonylation of each protein. By the addition of the specific chelator, the extents of protein carbonylation for BSA, lysozyme, and phosvitin were reduced by 64%, 41%, and 69%, respectively, of that which occurred due to the addition of SRM1648/EDTA/ascorbate.
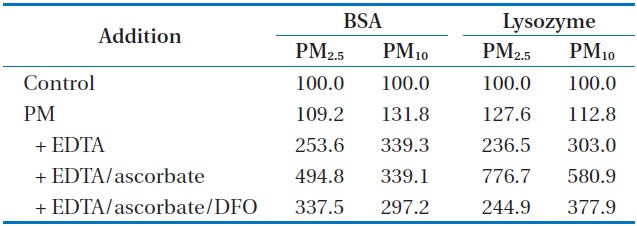
Table 3 shows how the carbonylation of BSA and lysozyme, in the presence of EDTA or EDTA/ascorbate was affected by the metals mobilized from PM2.5 and PM10. As has been observed with SRM1648, the incremental addition of PM, EDTA, and ascorbate incurred increasingly higher percentages of carbonylation to both of the proteins. The percentages of carbonylation in the presence of PM2.5/EDTA/ascorbate were 494% and 776% for BSA and lysozyme, respectively, and those in the presence of PM10/EDTA/ascorbate were 339% and 580% for BSA and lysozyme,
[Table 2.] Effect of mobilized metals from SRM1648 on the protein carbonylation (%)a
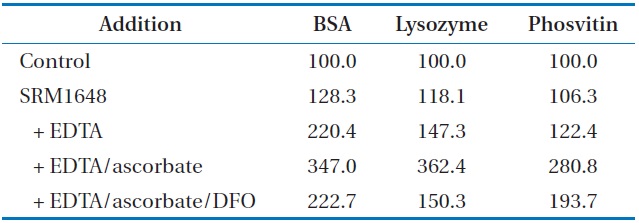
Effect of mobilized metals from SRM1648 on the protein carbonylation (%)a
respectively. This result suggests that the presence of PM2.5 incurred more oxidative damage to the proteins than that of PM10. The treatment of DFO to the reaction mixture containing PM/ EDTA/ascorbate did not significantly inhibit the carbonylation of each protein, as compared to the case with DNA SSB. The reduction in the extent of protein carbonylation was steeper with lysozyme than with BSA, as has been observed with SRM1648.
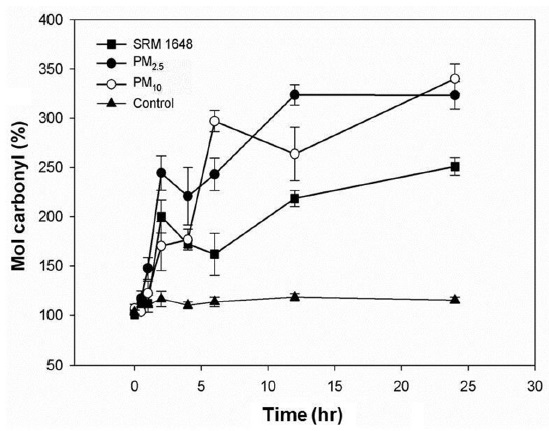
Fig. 2 shows the damaging effect of both soluble and mobilized metals from the particles phagocytized into the cells. The carbonylation of intracellular proteins were increased by 250% and 320% after 24 hr in A549 cells treated with SRM1648 and PM collected in Seoul, respectively, compared with that of the control. It was also observed that the extent of carbonylation was greater with PM2.5 than with PM10 up to 6 hr of treatment, after which data were comparable for both PM. Interestingly, with all cases, the rate of carbonylation of intracellular proteins dramatically increased initially, and then rather decreased at 4?6 hr, increasing again up to 24 hr.
In this study, the addition of a low molecular weight chelator (citrate or EDTA) greatly enhanced the reactivity of the particles with DNA and proteins (Tables 1?3). The increased reactivity suggests that the mobilization of metals from the particles might be directly correlated to the oxidative stress toward biomolecules, which is consistent with the report that iron plays a critical role in hydrogen peroxide-induced DNA single strand breaks
[Table 3.] Effect of mobilized metals from PM on protein carbonylation (%)a
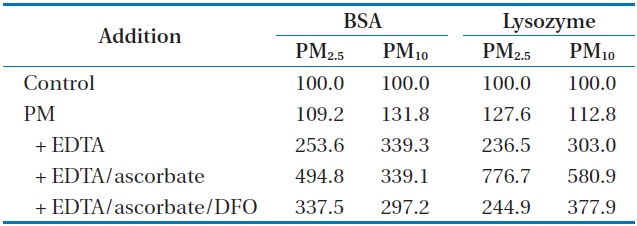
Effect of mobilized metals from PM on protein carbonylation (%)a
Although the mobilized fraction is small, some metals (Fe, Cu) are major participants in reactions which generate ROS, which causes damage to various biomolecules [5,8].
Our results also indicated that the oxidation of biomolecules was influenced by the source and size of the particles. The extent of DNA SSB was increased stepwise with the addition of SRM1648, SRM1648 plus EDTA, SRM1648 plus EDTA plus ascorbate (Table 1). However, the higher degrees of DNA SSB were obtained only by the presence of either PM2.5 or PM10 (Table 1). The overall extent of DNA SSBs was higher with PM2.5 than with PM10, and the extent of protein carbonylation was also higher with PM2.5 than with PM10 in the presence of EDTA/ascorbate (Tables 1?3). Although the amount of mobilized transition metals with several different chelators and reductants in the previous study [13] are not consistent with the results of DNA SSBs. This discrepancy between the amount of mobilized metals in the previous study and DNA SSBs in this study may be due to differing experimental microenvironments and/or other unknown factors, because mobilized matters are both inorganic and organic mixtures. The oxidation of protein in the presence of ascorbate may be of singular importance, since human lung epithelial lining fluid is estimated to contain 160 μM ascorbic acid [6]. In any case, the size of the ambient particles governs the surface area of the particles, and directly influences the oxidation of DNA and proteins. This is probably because PM2.5 has a larger surface area than PM10, and thus, metals are mobilized at a greater rate with PM2.5 than with PM10, as have been observed in our previous study [13]. Indeed, Seaton et al. [17] reported that ultrafine particles, even if present in the atmosphere at very low mass concentrations, are involved in the observed pathologies because of their large numbers per unit volume and their extremely high surface/volume ratio. Therefore, it is expected that, considering both the present results and the previously known toxicities of the transition metals, PM2.5 is more damaging to various biomolecules than PM10.
Two kinds of biomolecule were expected to have similar responses toward oxidative stress induced by mobilized metal; that is, the addition of specific chelator DFO to the reaction mixture containing EDTA/ascorbate was expected to decrease the degree of damage to both biomolecules. However, while the treatment decreased the damage of DNA to the level of the control, the treatment did not suppress much the damage of proteins (Tables 1?3). The above result indicated that the damage of DNA and proteins was caused by different groups of metals, since the same mixture of mobilized metals was employed, and DFO should have the same affinity for the same group of metals. This suggests that, for the better measurement of toxicities by the metals of ambient particles, proteins as well as DNA should be employed as a biomolecule, since the oxidative damage to both biomolecules resulted from different availabilities of the same group of metals.
The results presented here showed that metals deposited on to the ambient particles could be mobilized to become redoxreactive. Thus, the mobility of metals, as well as the total amount of metals needs to be properly assessed for the risk imposed by ambient particles. Furthermore, it might have the usefulness of