Lumbricidae is a relatively large holarctic family of mostly terrestrial earthworms comprising approximately 670 valid taxa from a total of 1,130 names in 63 genera (Blakemore, 2008a). Natural distribution is from North America (e.g.,
Specimens, lodged in National Institute of Biological Resources (NIBR), are described in the author’s usual style (e.g. Blakemore, 2010). Cytochrome c oxidase subunit 1 (COI barcode) sequences are appended with analyses via megaBLAST (www.blast.ncbi.nlm.nih.gov/BLAST.cgi). Abbreviations are rhs, right hand side; lhs, left hand side; TP, tuberculae pubertates.
Order Megadrilacea Benham, 1890
Family Lumbricidae Rafinesque-Schmaltz, 1815
Genus Eisenia Malm, 1877 (type-species: Enterion fetidum Savigny, 1826)
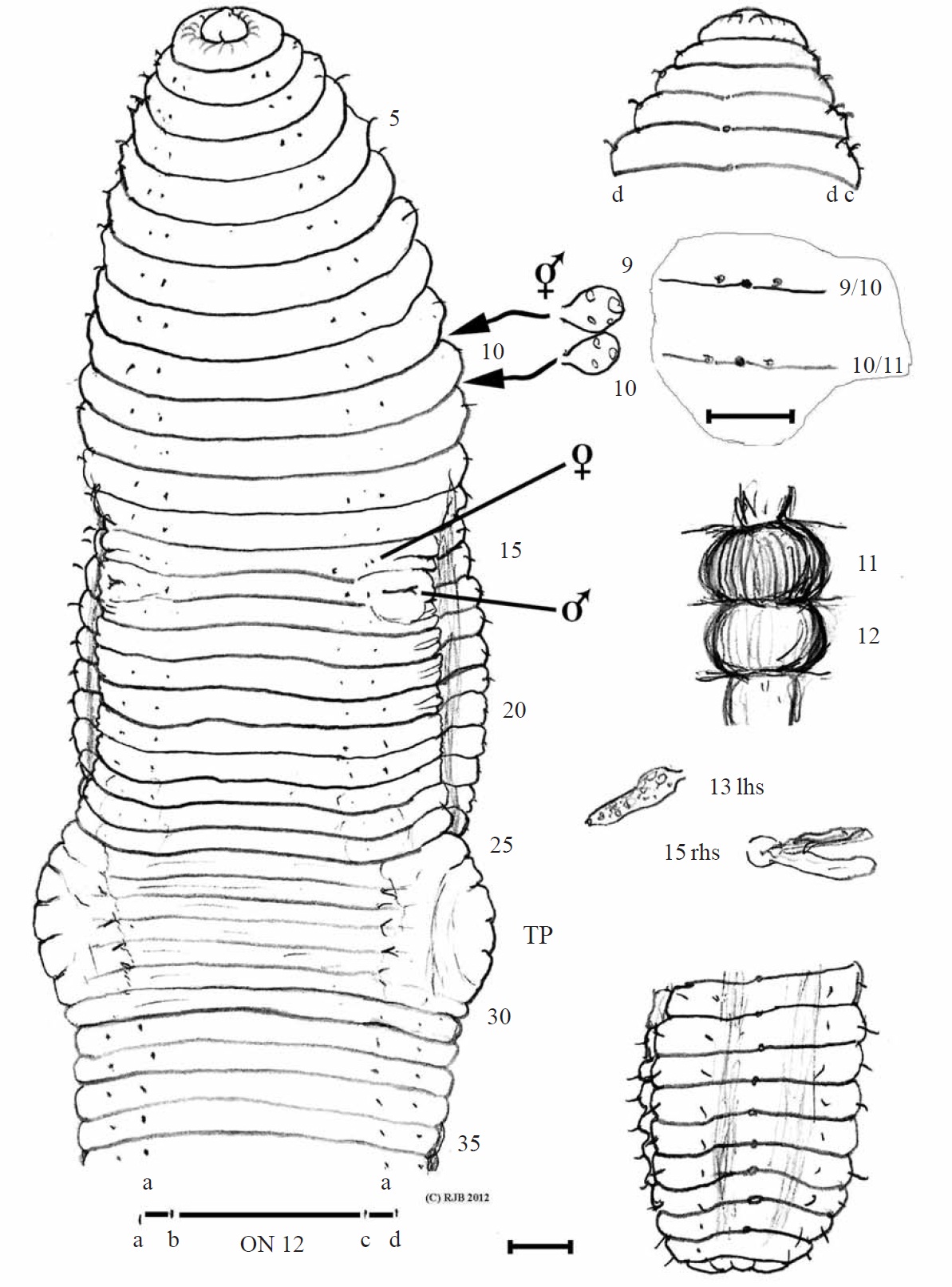
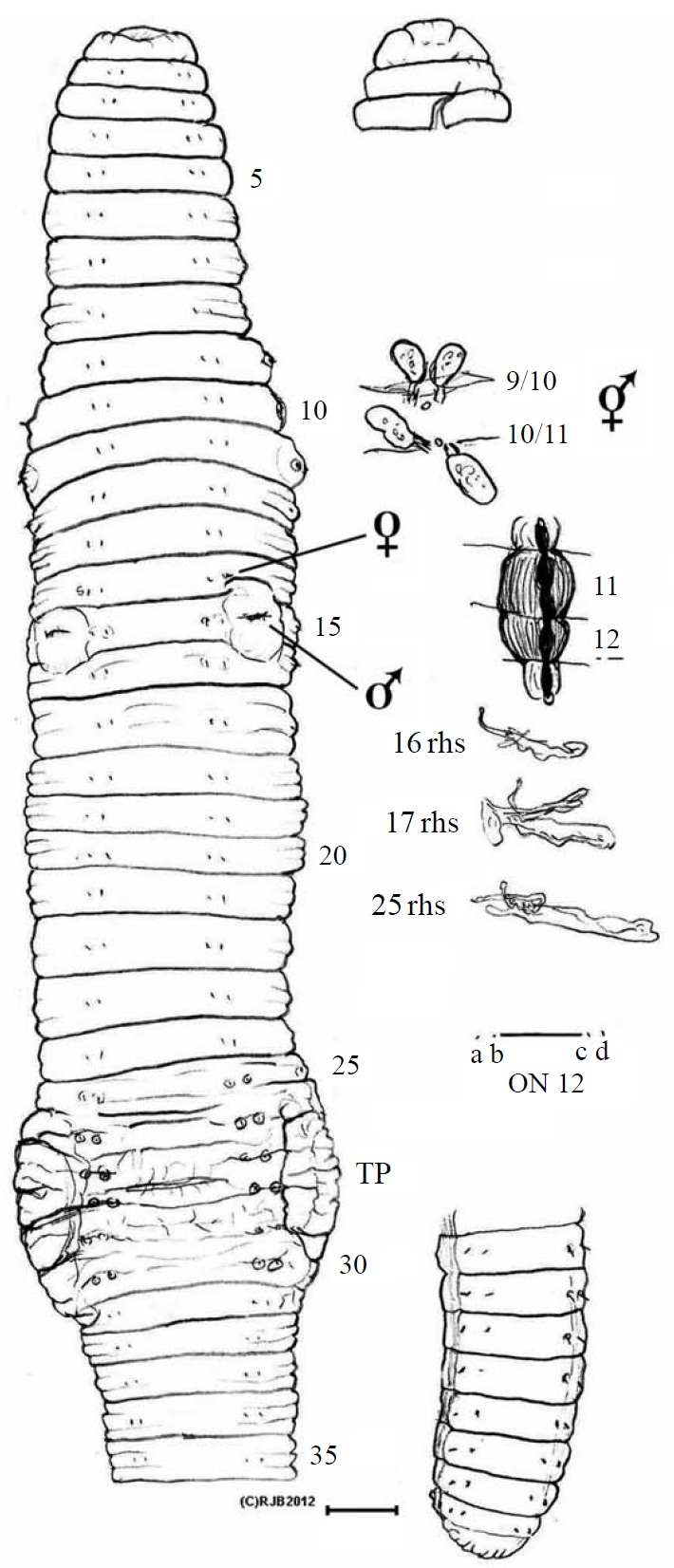
Eisenia gaga sp. nov. (Figs. 1, 2)
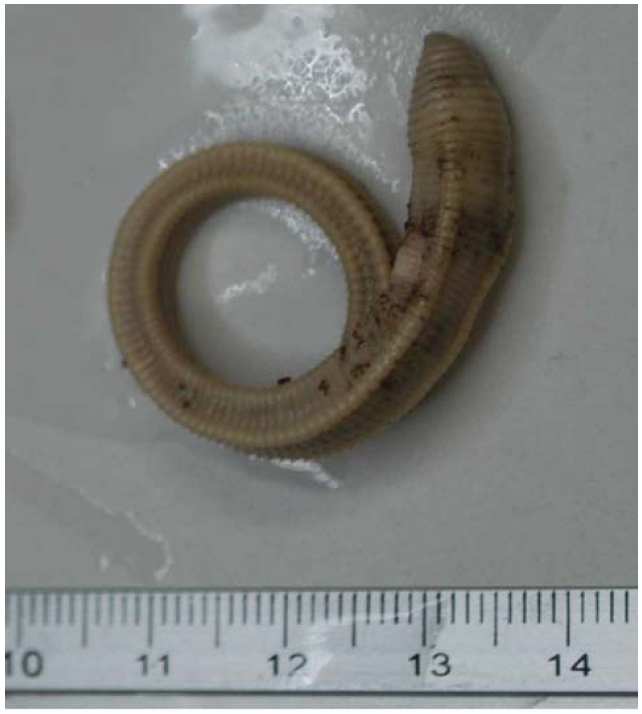
Material examined. Holotype, NIBR IV IV0000245509 (dissected and figured, providing DNA-WM1) (Fig. 1); eight paratypes, NIBR IV0000245510-IV0000245517 two dissected (specimen #2 WM2) and one photographed (#5 WM3) (Fig. 2), all fixed in 75% EtOH. Collected 26 Jan 2012 by Park TS, Seo HY from damp leaf litter on slopes of Mt. Doksil, 34°4′32.73N, 125°6′31.88E; summit 639 m, on Gageodo Island in Yellow Sea of South Korea. Found with a damaged
Etymology. After historical name for Gageodo Island, “Gaga” meaning “beautiful.”
Description. Body square in section after segment 12 with setae at each rounded corner; colour bleached in alcohol. Lengths 80-105 mm (holotype H 100, paratype P1 80, P2 100, P3 105). Segments 124-162 (H 162, P1 124, P2 145, P3 120). Prostomium epilobic, narrow. Setae closely paired. Tu-
midity obscures ventral setae in some of 11, 13, 14, 16, 17 and 22-31 and on 10 & 11 laterally; ab on 14 & 15 are slightly displaced. Clitellum pale 24,25-30,31, typically 25-31. Tubercula pubertatis on ca. 27-29 wide of setal b line on each side. Dorsal pores from 3/4 (minute), open from 4/5. Nephropores sporadically visible above d setae. Spermathecal pores in 9/10/11 close to mid-dorsal line. Female pores 14; male pores 15 with tumid mounds impinging slightly into adjacent segments.
Internally, septa 5/6-14/15 slightly thickened. Spermathecae round in 9 and 10. Testis and funnels free, iridescent in 10 and 11. Seminal vesicles in 9-12 those in (9 and) 10 smaller and filled with brown bodies. Ovaries small, flattened and tapering (like a folded hand) in 13. Ovisacs small anteriorly in 14 on 13/14. Hearts in 7-11. Nephridial bladders simple, sausage-shaped (in all segments inspected). Calciferous glands large and moniliform in 11 (vascularized) and in 12. Crop in 15 and, less dilated, in 16; muscular gizzard in 17- 18. Low but broad intestinal typhlosole develops from around 20-22. Gut contents, fine silty soil and organic matter and gizzard had a few largish ‘crop stone’ grits. Apart from ‘brown bodies’, no evidence of parasites was observed in the coelom, blood vessels or other organs.
Remarks.
Eisenia koreana (Zicsi, 1972)
Type material. Budapest University, holotype Ei-8 and eleven paratypes registered as Nr. 7000, from a brook bed behind Pyongyang Zoo (Mt. Taesong), North Korea.
Diagnosis. Size 45-52 mm by ca. 3 mm wide with 87-123 segments. Grey green in life, pale laterally in 10-11. Pro-epilobic. Posterior squarish. Dorsal pores from 4/5. Setal distances stated as “ab=bc” in Zicsi (1972: 129) was likely a typing error for ab=cd, with aa>bc and dd<1/2U. Setae ab tumid on 10, 16, 17, 23, 26-30 plus on cd in 10 and 11. Spermathecal pores in 9/10/11 dorsally but spermatophores also present. Clitellum 25-31, TP 27-29. Holandric with seminal vesicles in 9-12 (smaller in 10). Calciferous glands in 11 & 12. Crop in 15-16, gizzard in 17-18.
Remarks. The stated reason Zicsi (1972: 131) attributed his species to the semi-aquatic genus
Nearby at De-sang San, Zicsi (1972: 130) reported
Eisenia sindo sp. nov. (Fig. 3)
Material examined. Holotype, H IV0000246435 (sample #3 mature, dissected; DNA WO25) (Fig. 3); paratype P1 IV00 00246436 (sample #4 WO26) and paratypes IV0000246437 (12 matures, 5 juveniles, 5 immatures, and two hatchlings plus three fragments), all fixed in 75% EtOH. Collected 4
May 2012 by Blakemore RJ from mud beside creek between paddy fields just 200 m North of Sido bridge on Sindo Island, 37°53045′N, 126°43974′E, Incheon.
Description. Body tapering, flattened in posterior (somewhat rectangular); palid colour with light brown pigment. Lengths 100-120 mm (H and P1). Segments 148-152 (P1 and H). Prostomium epilobic wide. Setae closely paired, obscured by tumidity ventrally on 16 and around clitellum; cd setae tumid on 10 and 11 laterally. Clitellum pale 25-31. TP lateral on ca. 26,27-29. Dorsal pores from 4/5. Spermathecal pores paired in 9/10/11 near mid-dorsal line. Female pores on 14, male pores on mounds in 15.
Septa 5/6-8/9 slightly thick. Spermathecae inseminated, elongate in 9 and 10. Testis and funnels free, iridescent in 10 and 11. Seminal vesicles in 9-12 those in 10 small. Ovaries small in 13, ovisacs vestigial in 14. Hearts in 7-11. Nephridial bladders undeveloped or simple, mostly elongate sausage- shaped. Calciferous glands moniliform in 11 & 12. Crop in 16, muscular gizzard in 17-18. No typhlosole. Gut contains silty mud.
Remarks. Superficially similar to
The North American genus
Gates (1955: 10) said that seminal vesicles in segment 10 in one (or more?) of
Support for their divergent separation is decided by DNA barcode comparison of types of
>
Appendix 1. DNA cytochrome c oxidase subunit 1 data and BLAST analysis.
>WM1
TAAGTGTTGATAGAGGATTGGGTCCCCTCCCCCCGCTGGATCAAAAAATGAAGTATTAAGATTTCGATCTGTTAGGAGTATGGTAATTGCCCCTGCT AAAACTGGTAAAGAAAGGAGGAGAAGAACTACTGTGATTACTACAGCTCAAACAAATAGGGGAATTCGTTCAAGTCGTAATCCTTTTCATCGTATAT TAATAACAGTCGTAATGAAGTTGATGGCACCTAAGATTGAGGATGCACCTGCTAAGTGAAGAGAGAAAATGGCCAGATCTACTGAGGGGCCAGAG TGTGCGAGATTTCTAGATAGGGGGGGATAGACAGTTCATCCTGTTCCAGCTCCTTTTTCTACAGCAGCCGAGGATACTAATAAAATAAGAGATGGT GGAAGTAATCAAAATCTTATATTATTTAATCGGGGGAATGCCATGTCAGGGGCACCAAGTATTAGTGGGAGAAGCCAGTTTCCAAATCCTCCGATA AATACAGGTATCACAAGAAAAAAAATTATTACAAATGCATGCGCTGTAACGATAGTGTTATATAGTTGATCGCTACCTAAAAATGCTCCAGGTTGGC TTAACTCAATTCGGATTAGGAGGCTTATTCCTGCACCCACTATACCAGCTCAAACTCCTAGAATAAAATATA
> WM2
AAGTGTTGATAGAGGATTGGGTCCCCTCCCCCCGCTGGATCAAAAAATGAAGTATTAAGATTTCGATCTGTTAGGAGTATGGTAATTGCCCCTGCT AAAACTGGTAAAGAAAGGAGGAGAAGAACTACTGTGATTACTACAGCTCAAACAAATAGGGGAATTCGTTCAAGTCGTAATCCTTTTCATCGTATAT TAATAACAGTAGTAATGAAGTTGATGGCACCTAAGATTGAGGATGCACCTGCTAAGTGAAGAGAGAAAATGGCCAGATCTACTGAGGGGCCAGAG TGTGCGAGATTTCTAGATAGGGGGGGATAGACAGTTCATCCTGTTCCAGCTCCTTTTTCTACAGCAGCTGAGGATACTAATAAAATAAGAGATGGT GGAAGTAATCAAAATCTTATATTATTTAATCGGGGGAATGCCATGTCAGGGGCACCAAGTATTAGTGGGAGAAGCCAGTTTCCAAATCCCCCGATA AATACAGGTATCACAAGAAAAAAAATTATTACAAATGCATGCGCTGTAACGATAGTGTTATATAGTTGATCGCTACCTAAAAATGCTCCAGGTTGGC TTAACTCAATTCGGATTAGGAGGCTTATTCCTGCACCCACTATACCAGCTCAAACTCCTAGAATAAAATA
> WM3
AATAAGTGTTGATAGAGGATTGGGTCCCCTCCCCCCGCTGGATCAAAAAATGAAGTATTAAGATTTCGATCTGTTAGGAGTATGGTAATTGCCCCT GCTAAAACTGGTAGAGAAAGGAGGAGAAGAACTACTGTGATTACTACAGCTCAAACAAATAGGGGAATTCGTTCAAGTCGTAATCCTTTTCATCGT ATATTAATAACAGTAGTAATGAAGTTGATGGCACCTAAGATTGAGGATGCACCTGCTAAGTGAAGAGAGAAAATGGCCAGATCTACTGAGGGGCCA GAGTGTGCGAGATTTCTAGATAGGGGGGGATAGACAGTTCATCCTGTTCCAGCTCCTTTTTCTACAGCAGCCGAGGATACTAATAAAATAAGAGAT GGTGGAAGTAATCAAAATCTTATATTATTTAATCGGGGGAATGCCATGTCAGGGGCACCAAGTATTAGTGGGAGAAGCCAGTTTCCAAATCCCCCG ATAAATACAGGTATCACAAGAAAAAAAATTATTACAAATGCATGCGCTGTAACGATAGTGTTATATAGTTGATCGCTACCTAAAAATGCTCCAGGTT GGCTTAACTCAATTCGGATTAGGAGGCTTATTCCTGCACCCACTATACCAGCTCAAACTCCTAGAATAAAATAT
> WO25
ACCTTATACTTTATTCTTGGGGTTTGAGCCGGAATAGTAGGCGCTGGAATAAGCCTCTTAATCCGAATCGAGCTAAGACAGCCTGGAGCATTCCTG GGAAGAGACCAGCTATATAATACCATTGTTACAGCTCATGCGTTCGTAATAATCTTTTTTCTTGTAATACCTGTATTTATTGGGGGGTTCGGCAACTG ACTTCTCCCATTAATATTGGGGGCTCCCGACATAGCATTCCCTCGTTTAAATAATATAAGATTCTGGCTACTTCCCCCTTCCCTTATTCTACTAGTCT CATCAGCAGCGGTTGAGAAAGGGGCGGGAACAGGTTGAACTGTGTACCCGCCCCTATCTAGAAATCTTGCACACGCTGGGCCATCAGTAGACCTG GCTATTTTCTCCCTTCATTTAGCGGGTGCGTCGTCTATTCTAGGAGCCATCAATTTTATCACTACAGTTATCAATATACGATGAAGAGGGTTACGTCT TGAACGAATTCCACTATTTGTGTGAGCTGTAGTAATTACTGTTGTTCTTCTTCTTCTCTCCCTACCAGTTCTAGCAGGAGCAATTACCATACTTCTAA CAGATCGAAACTTAAACACTTCATTCTTTGACCCCGCAGGAGGTGGAGATCCTATTCTTTATCAACATCTATT
>WO26
CCTTATACTTTATTCTTGGGGTTTGAGCCGGAATAGTAGGCGCTGGAATAAGCCTCTTAATCCGAATCGAGCTAAGACAGCCTGGAGCATTCCTGG GAAGAGACCAGCTATATAATACCATTGTTACAGCTCATGCGTTCGTAATAATCTTTTTTCTTGTAATACCTGTATTTATTGGGGGGTTCGGCAACTGA CTTCTCCCATTAATATTGGGGGCTCCCGACATAGCCTTCCCTCGTTTAAATAATATAAGATTCTGGCTACTTCCCCCTTCCCTTATTCTACTAGTCTC ATCAGCAGCGGTTGAGAAAGGAGCGGGAACAGGTTGAACTGTGTACCCGCCCCTATCTAGAAATCTTGCACACGCTGGGCCATCAGTAGACCTGG CTATTTTCTCCCTTCATTTAGCGGGTGCGTCATCTATTCTAGGAGCCATCAATTTTATCACTACAGTTATCAATATACGATGAAGAGGGTTACGTCTT GAACGAGTTCCACTATTTGTGTGAGCTGTAGTAATTACTGTTGTTCTTCTTCTTCTCTCCCTGCCAGTTCTAGCAGGAGCAATTACCATACTTCTAAC AGATCGAAACTTAAACACTTCATTCTTTGACCCCGCAGGAGGTGGGGATCCTATTCTTTATCAACATCTATTC
Barcode results and BLAST conclusions
1. BLASTn alignments: >99% WM1=WM2=WM3, i.e., samples of same species.
2. megaBLAST of WM1
3. BLASTn alignment WM1
4. megaBLAST of FJ214226
5. BLASTn alignment WM1
6. BLASTn alignment WO25 vs. WO26
7. megaBLAST WO25
8. BLASTn alignment WO25