Aeromonas salmonicida is an important fish pathogen that has a geographically widespread distribution, with a broad host range and an economically destructive impact on culti-vated fish, particularly salmonids (Trust et al., 1980; Fryer et al., 1988; Johnsen and Jensen, 1994; Mooney et al., 1995; No-mura et al., 2002). As a member of the family Aeromonadace-ae, A. salmonicida has been classified into typical and atypical A. salmonicida pathogroups (Wiklund and Dalsgaard, 1998; Abbott et al., 2003). The “typical” group of A. salmonicida is frequently associated with furunculosis in salmonids. Such a group strain includes A. salmonicida subsp. salmonicida. In contrast, the “atypical” group of A. salmonicida is involved in various ulcerative diseases or atypical furunculosis in salmo-nids and non-salmonids (Wiklund and Dalsgaard, 1998). Oth-er A. salmonicida subspecies such as A. salmonicida subsp. achromogenes, masoucida, smithia, and pectinolytica belong in this atypical A. salmonicida group (Wiklund and Dalsgaard, 1998). Although a classification has been created according to both genetic and biochemical characteristics of individual A. salmonicida strains, subspecies populations unable to be classified under these criteria have been increasing; thus, more reliable classification methods are needed for atypical A. sal-monicida strains (Lund et al., 2003).
Previous studies have reported that both typical and atypi-cal A. salmonicida infections occur worldwide, including in North America, Europe, and Japan (Trust et al., 1980; Fryer et al., 1988; Mooney et al., 1995; Nomura et al., 2002). Among the various A. salmonicida fish host species, salmonids are the most susceptible to furunculosis (Lund and Mikkelsen, 2004; Austin and Austin, 2007). In Korea, A. salmonicida subsp. sal-monicida was isolated from cultured masu salmon (Oncorhyn-chus masou) in 1986 (Fryer et al., 1988) and more recently detected by the PCR from rearing water on a trout farm (Lee et al., 2002). However, A. salmonicida subsp. salmonicida has not been reported in other salmonids in Korea until now. In this study, the prevalence of typical A. salmonicida was ex-amined in both migrating wild chum salmon and artificially hatched fry during 2006-2011 in Korea by PCR using typical A. salmonicida-specific vapA gene primers. Because the vapA gene encodes the surface virulence A-layer protein, the vapA-positive and A-layer-expressing typical A. salmonicida were isolated and further characterized using both biochemical and phylogenetic analyses.
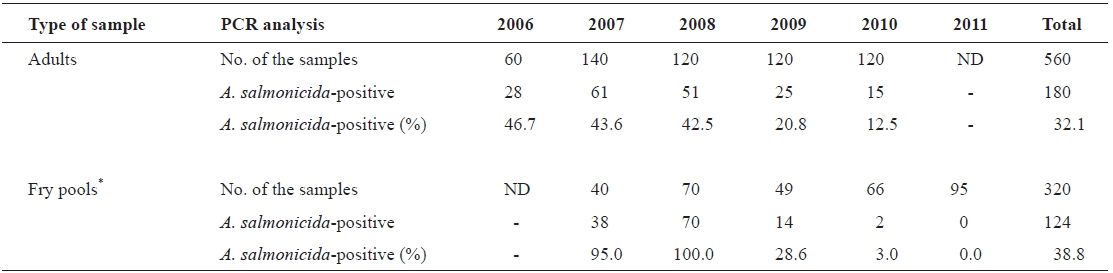
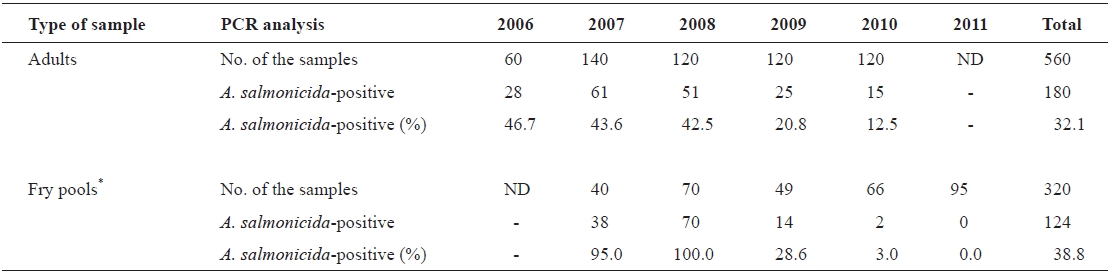
Wild adults and artificially hatched chum salmon fry sam-ples were randomly collected from the Namdae River basin and hatcheries at Yangyang City (located on the east coast of South Korea) during 2006-2011. The number of chum salmon samples in this study is listed in Table 1. Both kidney and spleen from individual wild adult chum salmon were col-lected after artificial spawning and used for the PCR analysis. Artificially hatched fry samples were randomly selected and euthanized with excess MS-222 (tricaine methane sulfonate; Sigma; St. Louis, MO, USA). Five individual fry were pooled
and considered as 1 fry sample for the PCR analysis. Subse-quent bacterial isolation was conducted for all PCR-positive adult samples as described below.
One hundred mg of adult chum salmon kidney were asepti-cally sampled and homogenized with 900 μL of phosphate-buffered saline. Each homogenate (100 μL) was spread direct-ly on trypticase soy agar (TSA; Difco, Detroit, MI, USA) and furunculosis agar (FA) plates containing 1% tryptone (Difco), 0.5% yeast extract (Oxoid, Hampshire, UK), 0.1% L-tyrosine (Sigma), and 0.25% sodium chloride. If brown-pigmented colonies appeared on the plates after a 15℃ incubation for 2?7 days, they were selected and sub-cultured in trypticase soy broth (Difco) for a preliminary biochemical screening, including Gram staining and assessment of oxidase activity and catalase production, as previously described (Austin and Austin, 2007). Further biochemical characterization of the A. salmonicida isolates was performed using the API 20E kit (BioMerieux, Marcy l’Etoile, France) according to the manu-facturer’s instructions. Bacterial growth on Coomassie Bril-liant Blue agar (CBBA) plates was examined as previously described (Cipriano and Bertolini, 1988) to confirm the pres-ence of the surface A-layer.
Antimicrobial susceptibilities were tested using a standard disk diffusion assay and Mueller-Hinton agar plates (Difco). The diameters of bacterial growth inhibitory zones were mea-sured and interpreted according to NCCLS guidelines (Na-tional Committee for Clinical Laboratory Standard, 2001).
Genomic DNA was extracted from 100 mg of salmon tissue sample or 5 mL of cultured bacterial cells using the AccuPrep Genomic DNA Extraction kit (Bioneer, Seoul, Korea) accord-ing to the manufacturer's instructions. All genomic DNA was adjusted to a final concentration of 100 ng/μL for the PCR analysis.
The PCR amplification was performed to detect a partial vapA gene specific to typical A. salmonicida using the prim-ers AP1 (5′-GGC TGA TCT CTT CAT CCT CAC CC-3′) and AP2 (5′-CAG AGT GAA ATC TAC CAG CGG TGC-3′), as previously described (Gustafson et al., 1992). The target gene was amplified with 25 μM of each primer and 2 μL of genom-ic DNA using the lyophilized AccuPower PCR Premix (Bi-oneer). Thermal conditions consisted of an initial incubation at 94℃ for 2 min, 30 cycles of 15 s at 94℃, 30 s at 57℃, and 90 s at 72℃, with a final extension at 72℃ for 3 min (Byers et al., 2002). Genomic DNA from typical A. salmonicida (FPC 367 strain, Japan) was used as a positive control.
All vapA genes from the A. salmonicida isolates were am-plified using the complete A. salmonicida vapA gene-specific primers F-1 (5′-TCA ACG GAT AGG TTC AAC CC-3′) and R-1 (5′-CAG AGT GAA ATC TAC CAG CGG TGC-3′) as previously described to analyze the genetic diversity in typi-cal A. salmonicida-specific vapA genes (Lund et al., 2003). After conducting 1% agarose gel electrophoresis, the PCR products were purified using the AccuPrep gel purification kit (Bioneer) according to the manufacturer’s protocol, and the DNA nucleotide sequence was analyzed using the ABI PRISM 3130xl Genetic Analyzer (Applied Biosystems, Foster City, CA, USA) available at National Instrumentation Center for Environmental Management, Seoul National University (NICEM, Seoul, Korea).
DNA nucleotide sequences of all vapA genes were analyzed by multiple-sequence alignment using ClustalW2 (Thompson et al., 1994). Phylogenetic trees were generated using the Mo-lecular Evolutionary Genetic Analysis software (MEGA) ver-sion 4 and the neighbor-joining method with bootstrap values of 1000 replicates (Tamura et al., 2007).
To examine the prevalence of typical A. salmonicida infec-tion among chum salmon in Korea, 880 chum salmon sam-ples, including wild adults (n = 560) and artificially hatched fry pools (n = 320) (Table 1), were randomly collected and analyzed by the previously established PCR method using
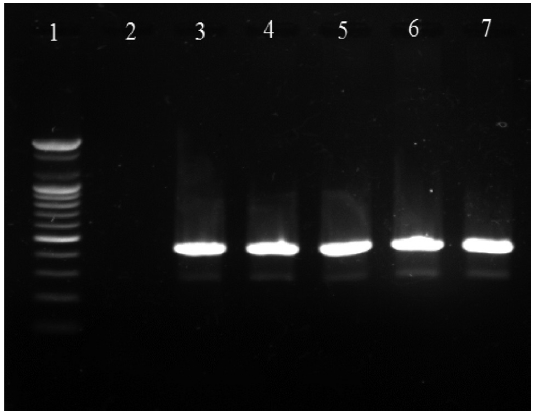
the vapA gene primers specific to typical A. salmonicida. The results demonstrated that approximately 34.5% of the samples (304/880 samples) produced the 421-bp amplicons specific to the vapA gene as expected (Fig. 1), implying that chum salmon infection by typical A. salmonicida is common in Korea (Table 1). Interestingly, the prevalence of typical A. salmonicida in the chum salmon fry pools was relatively higher in 2007 (95.0%) and 2008 (100%) than that in 2009 (28.6%), 2010 (3.0%), and 2011 (0.0%) (Table 1). A similar pattern was observed in the adult chum salmon samples. The prevalence of A. salmonicida in the wild adult chum salmon was approximately 46.7, 43.6, and 42.5% in 2006, 2007, and 2008, respectively, but decreased by 20.8% in 2009 and 12.5% in 2010 (Table 1).
Twenty typical A. salmonicida isolates were recovered from the 304 wild adult chum salmon samples that were posi-tive on the PCR using vapA-specific primers. As expected, all isolates were brown-pigmented when grown on TSA or FA plates, which is a characteristic of typical A. salmonicida. A preliminary screening confirmed that all 20 brown-pigmented isolates belonged to A. salmonicida because all were Gram-negative, oxidase- and catalase-positive, non-motile rods and possessed the ability to reduce nitrate and degrade gelatin (data not shown).
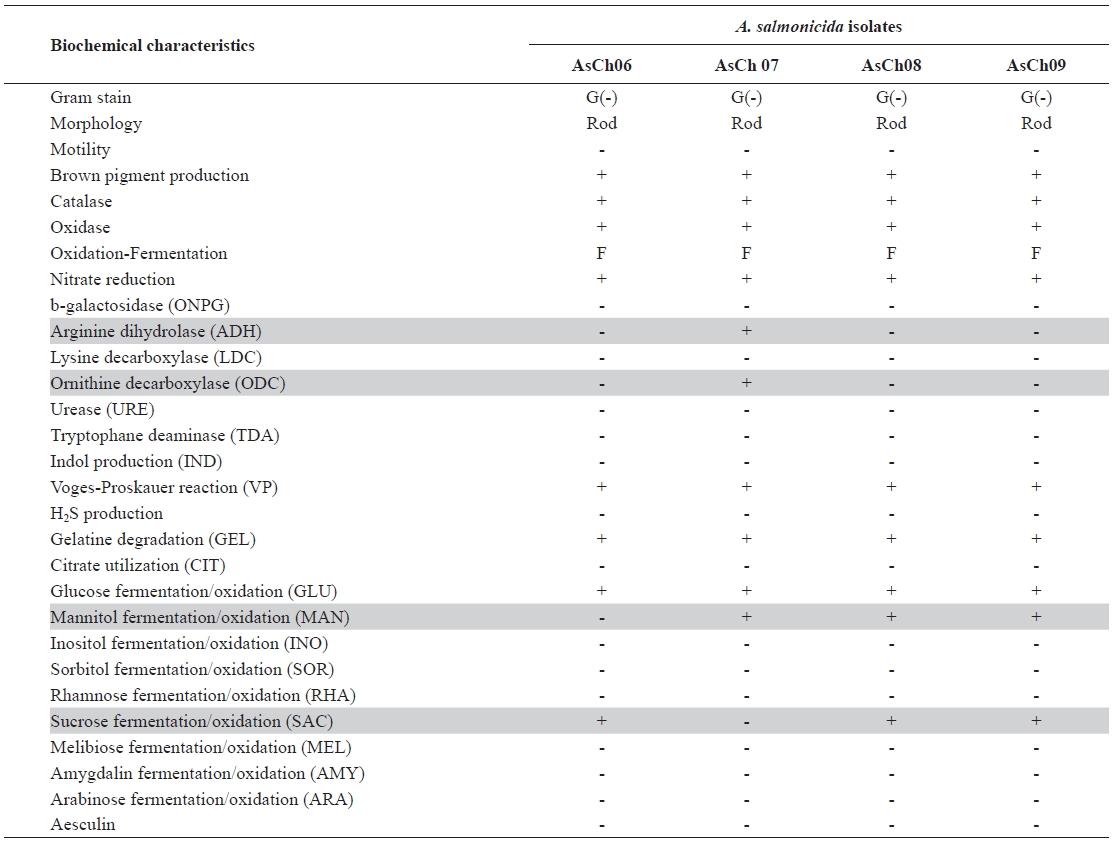
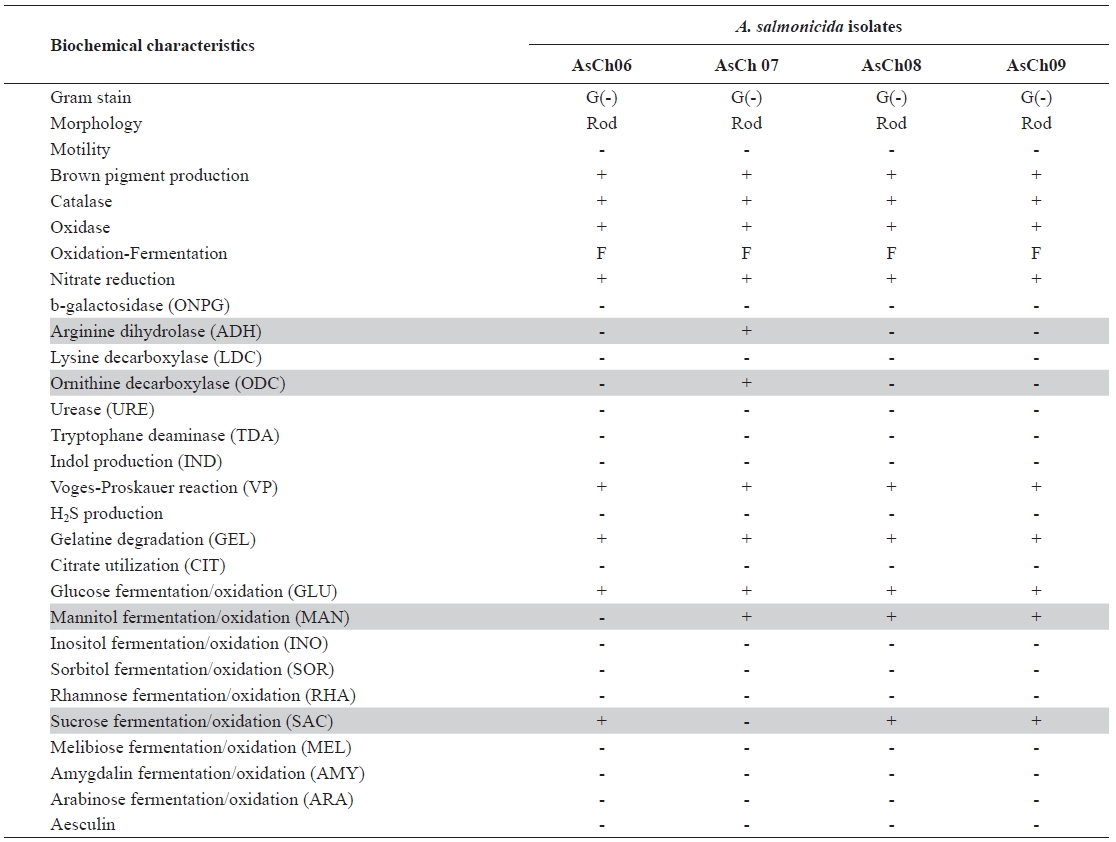
The four typical A. salmonicida isolates were randomly selected, designated AsCh06, AsCh07, AsCh08, and AsCh09, respectively, and further characterized by biochemical tests (Table 2). The results showed that the AsCh08 and AsCh09 strains had the same biochemical phenotypes (API interpre-tation profile no. 0007124). However, they slightly differed from AsCh06 (API interpretation profile no. 0007024) and AsCh07 (API interpretation profile no. 2107124) in their biochemical activity, particularly for arginine dihydrolase, ornithine decarboxylase, mannitol fermentation, and sucrose fermentation (Table 2). The AsCh08 and AsCh09 strains were consistently able to produce all-blue colonies when cultured on CBBA plates due to surface A-layer protein expression. The antibiotic susceptibility test revealed that all typical A. salmonicida isolates were susceptible to various antibiotics including tetracyclines, aminoglycosides, β-lactam, and mac-rolides antibiotics but were resistant to nalidixic acid (data not shown).
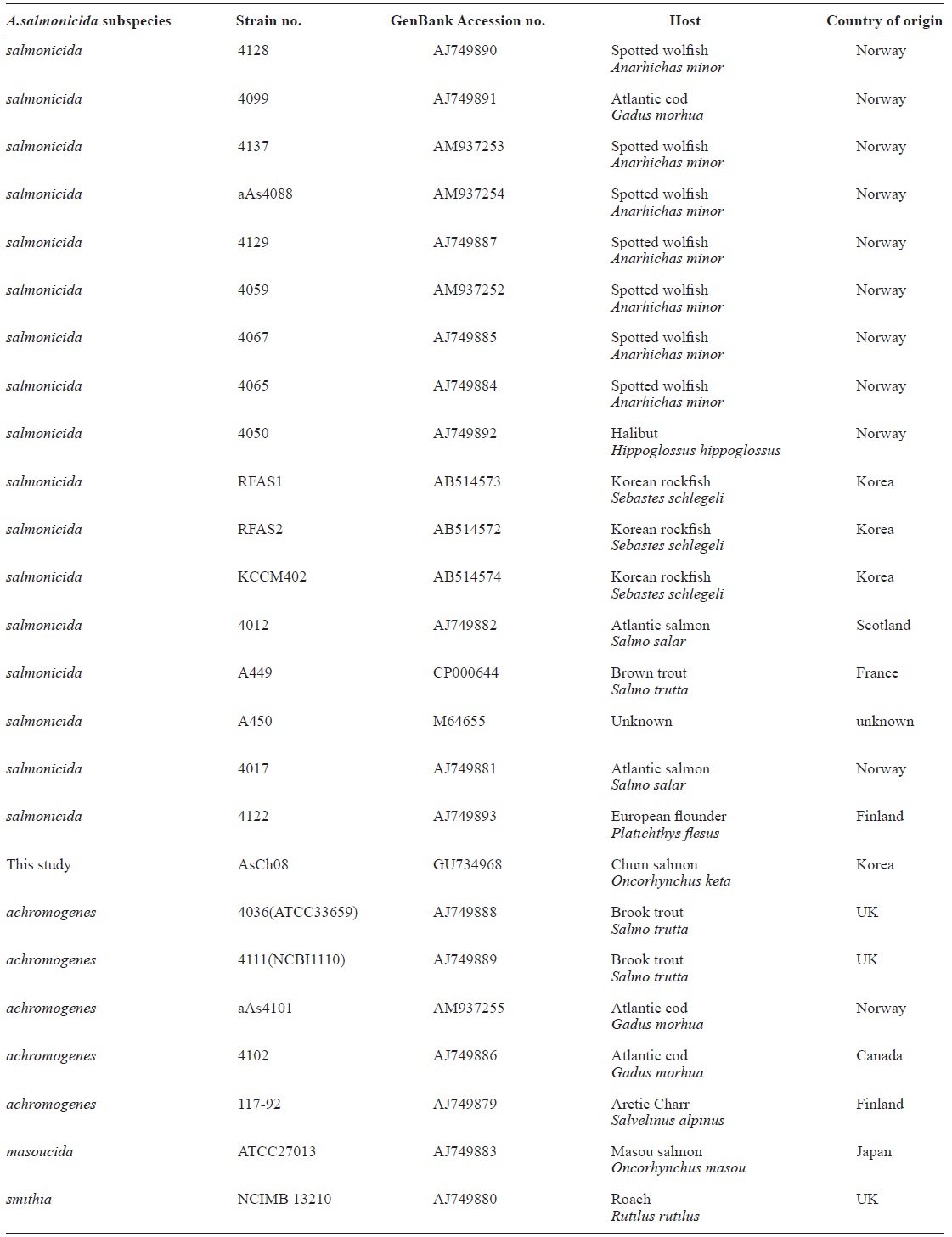
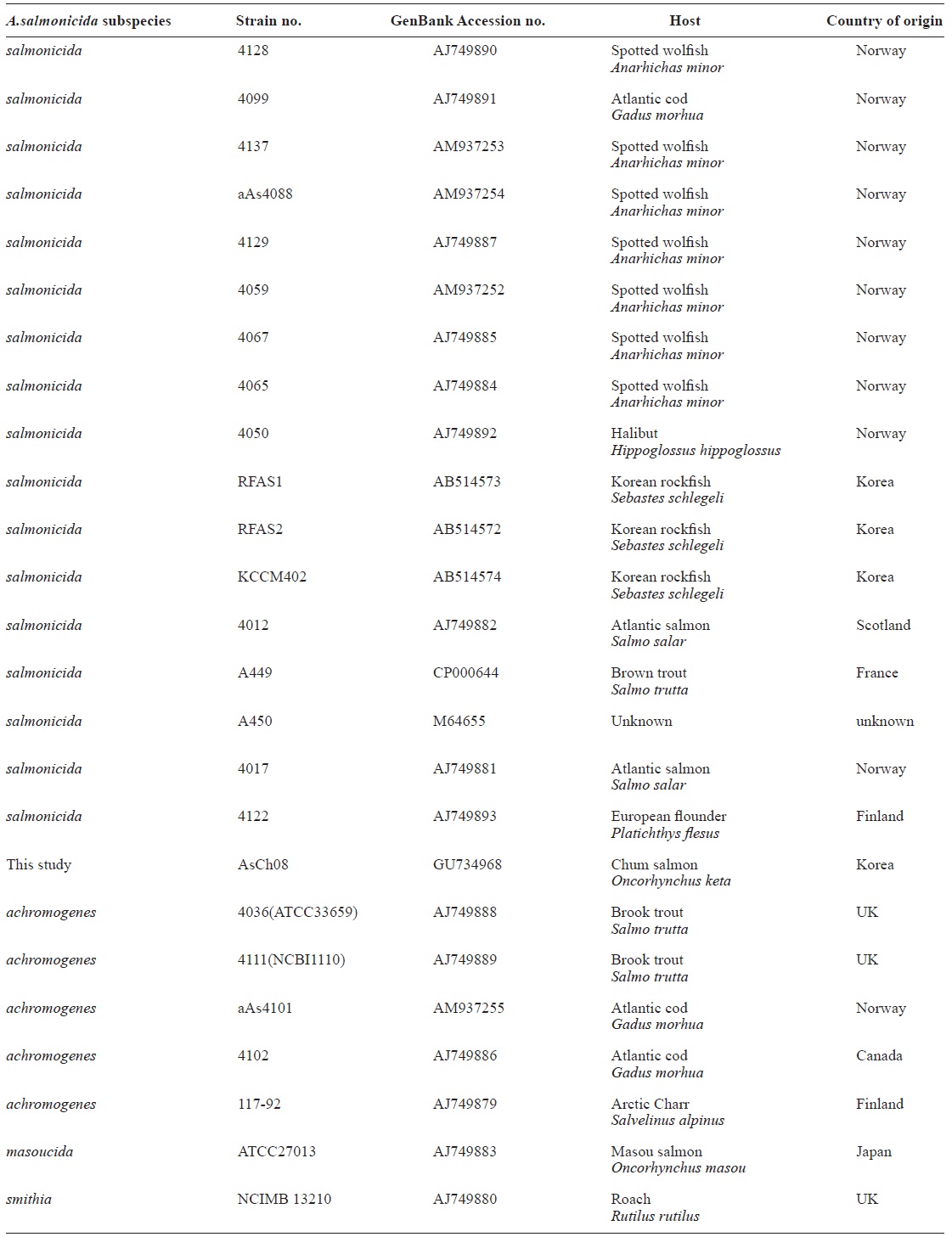
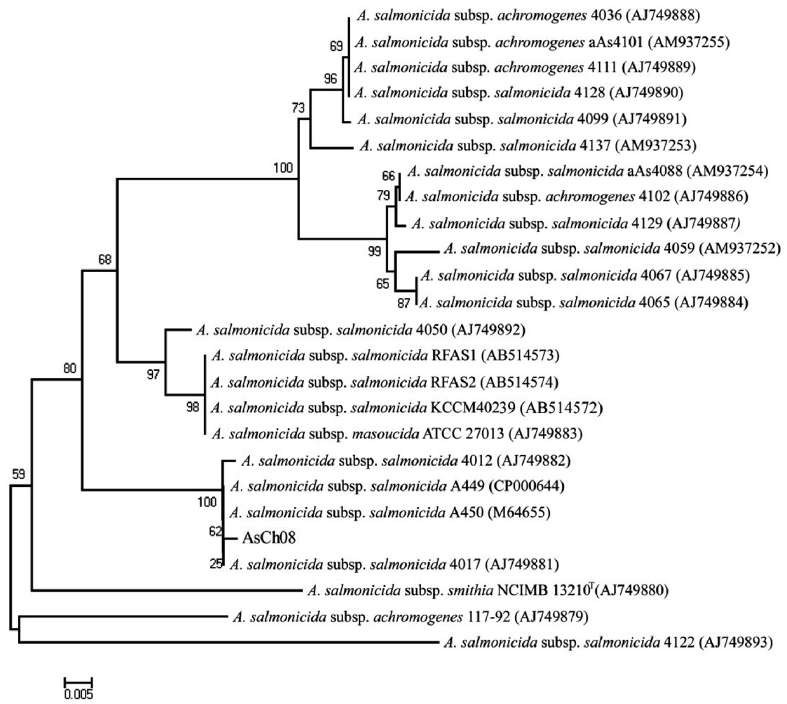
All vapA genes were amplified from the genomic DNA of AsCh06-09 by PCR and analyzed by DNA nucleotide se-quencing for the phylogenetic analysis. The DNA sequence data were deposited in the GenBank database with the acces-sion number GU734698. The analysis showed that all isolates carried identical vapA genes (data not shown). Based on the vapA gene sequences, a phylogenetic tree was produced by comparing the AsCh08 isolate with the typical A. salmonicida isolates from other countries (Table 3). Interestingly, AsCh08 showed 99.9% similarity and co-clustered with typical A. salmonicida strains previously isolated from Atlantic salmon and salmonids in Scotland and Norway such as A. salmonicida subsp. salmonicida 4012 (AJ749882), 4017 (AJ749881), A449 (CP000644), and A450 (M64655) strains (Table 4, Fig. 2). Although further detailed study is needed, it should be noted that AsCh08 showed less similarity (~97% similarity) with other typical A. salmonicida isolates from Korea such as KCCM40239 (AB514572), RFAS1 (AB514573), and RFAS2 (AB514574) than with those from Atlantic salmon and salmo-nids in Scotland and Norway (Table 4, Fig. 2).
Wild adult chum salmon samples positive for the A. sal-monicida vapA gene were used for bacterial isolation, and the colonies producing a diffusible brown pigment, a characteris-tic of typical A. salmonicida isolates, were further examined. Biochemical analyses revealed that all brown-pigmented iso-lates were Gram-negative, oxidase-positive, catalase-positive, non-motile rods. They also produced dark blue colonies when cultured on CBBA plates, suggesting the presence of an A-layer (Cipriano and Bertolini, 1988). These results are consistent with the biochemical characteristics of typical A. salmonicida (Pa-van et al., 2000). The PCR results showed that typical A. sal-monicida was consistently detected at a high prevalence in both wild adult chum salmon and artificially hatched fry pools dur-ing 2006%2011. A similar long-term study reported a 1.7-50% prevalence of typical A. salmonicida infection in adult chum salmon from Japanese rivers (Nomura et al., 2002). Several other studies also showed that furunculosis is enzootic among wild salmon populations in rivers (Austin and Austin, 2007 and the references therein).
Typical A. salmonicida strains are thought to be extremely ho-mogeneous. Therefore, epidemiological discrimination among the strains based on either phenotypic or biochemical proper-ties has shown limited success in elucidating their geographical relatedness and/or host preference. Moreover, conventional ge-notypic methods such as plasmid profiling and ribotyping of A. salmonicida do not provide a clear epidemiological marker for furunculosis outbreaks (Nielsen et al., 1993, 1994). Sequencing of the 16rRNA gene is a robust tool for bacterial taxonomy, but genetic analysis of the 16S rRNA gene revealed that it is high-ly conserved in the genus Aeromonas (Martinez-Murcia et al., 2005). The vapA gene sequences of the five typical A. salmonici-da isolates have been investigated, and results showed that they were identical and were grouped into the same cluster (Lund and Mikkelsen, 2004). Nonetheless, typical A. salmonicida strains were resolved in two clusters by geographical origin; one cluster contains mainly North American strains, and the other cluster contains mainly northern European strains (O’hlci et al., 2000). In the present study, the AsCh08 vapA gene sequence (GenBank accession no. GU734968) showed 99.9% similarity with that of A. salmonicida subsp. salmonicida strains A449 (GenBank ac-cession number CP000644) and 4017 (GenBank accession no. AJ749881). In this study, AsCh08 was clustered with typical A. salmonicida type strain A450 (GenBank accession no. M64655) and type strains 4012 (GenBank acces-sion no. AJ749882), 4017 (GenBank accession no. AJ749881), and A449 (GenBank accession no. CP000644). Unexpectedly, these strains were isolated from Atlantic salmon and salmonids in Scotland and Norway, all of which are geographically distant from Korea. Although this phenomenon cannot be explained, it would be interesting to investigate the possible relationship between these isolates and their geographical origin in detail.
A phylogenetic tree of 23 typical and atypical A. salmonicida strains has been reported based on their vapA gene sequences (Lund and Mikkelsen, 2004). The results demonstrated that most of the vapA gene sequence is highly conserved and that all typical A. salmonicida strains are localized in the same phy-logenic cluster regardless of their geographic origin. Therefore, more discriminative genetic markers are required for epidemio-logical studies because it is still uncertain how this pathogen has been disseminated worldwide and how it affects wild salmonid populations.
In conclusion, typical A. salmonicida infection among chum salmon in Korea was monitored by PCR during 2006-2011, and the infection demonstrated a high prevalence. Although no infor-mation is currently available on the pathogenic potential of these pathogens and to what extent they have affected propagation of chum salmon in Korea, an appropriate control method should be considered to prevent possible clinical outbreaks in holding ponds and/or hatcheries. Affected wild adult chum salmon with typical A. salmonicida could act as carriers by transporting the pathogen to spawning areas (Nomura et al., 2002).