The rainbow trout
The purpose of this study is to report the isolation and char-acterization of
Sixty-four juvenile rainbow trout (0.4-0.7 kg body weight) were collected from four hatcheries in Gangwon Province, South Korea, in 2010 for routine disease monitoring. Fish were packed individually in plastic bags, transported to the laboratory, and processed immediately. Gill and kidney tissues were aseptically removed from individual fish and placed in 1.5-mL microcentrifuge tubes.
>
Isolation of bacteria and biochemical character-ization
Gill and kidney tissue samples (100 mg each) were homog-enized with 0.9% (w/v) NaCl and then spread on trypticase soy agar (TSA) plates at 15℃ for 2-7 days. Among the colo-nies that appeared, yellow-pigmented colonies, characteristic of the genus
>
Extraction of DNA and amplification of 16S ribo-somal RNA (rRNA) genes
Genomic DNA was extracted from 5 mL of cultured bac-terial cells by the AccuPrep Genomic DNA Extraction Kit (Bioneer, Daejeon, Korea). Extracted nucleic acids were con-centrated to a final concentration of 100 ng/μL. PCR targeting the 16S rRNA was conducted using the universal bacterium-specific primer set, 20F (5′-AGA GTT TGA TCA TGG CTC AG-3′) and 1500R (5′-GGT TAC CTT GTT ACG ACT T-3′) (Weisburg et al., 1991). PCR was performed using the lyophi-lized AccuPower PCR PreMix tube (Bioneer) containing 20 μL of reaction mixture, which comprised 25 μM of each prim-er, 5 μL of extracted DNA and DEPC-treated water, with a DNA Engine Peltier Thermal Cycler (BioRad, Hercules, CA, USA). Amplification was carried out with an initial cycle at 95℃ for 5 min, followed by 30 cycles of 95℃ for 30 s, 51℃ for 1 min, and 72℃ for 2 min, with a final extension step at 72℃ for 5 min.
>
Sequencing and phylogenetic analysis of the 16S rRNA genes
PCR products were purified by an AccuPrep Gel Purification Kit (Bioneer). Then, 10 ng/μL of purified PCR products were directly sequenced (ABI 3730
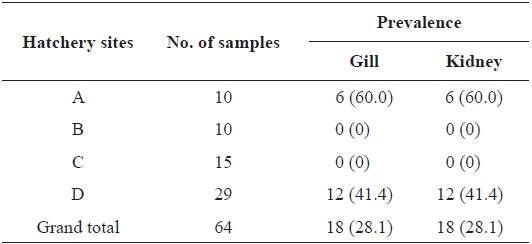
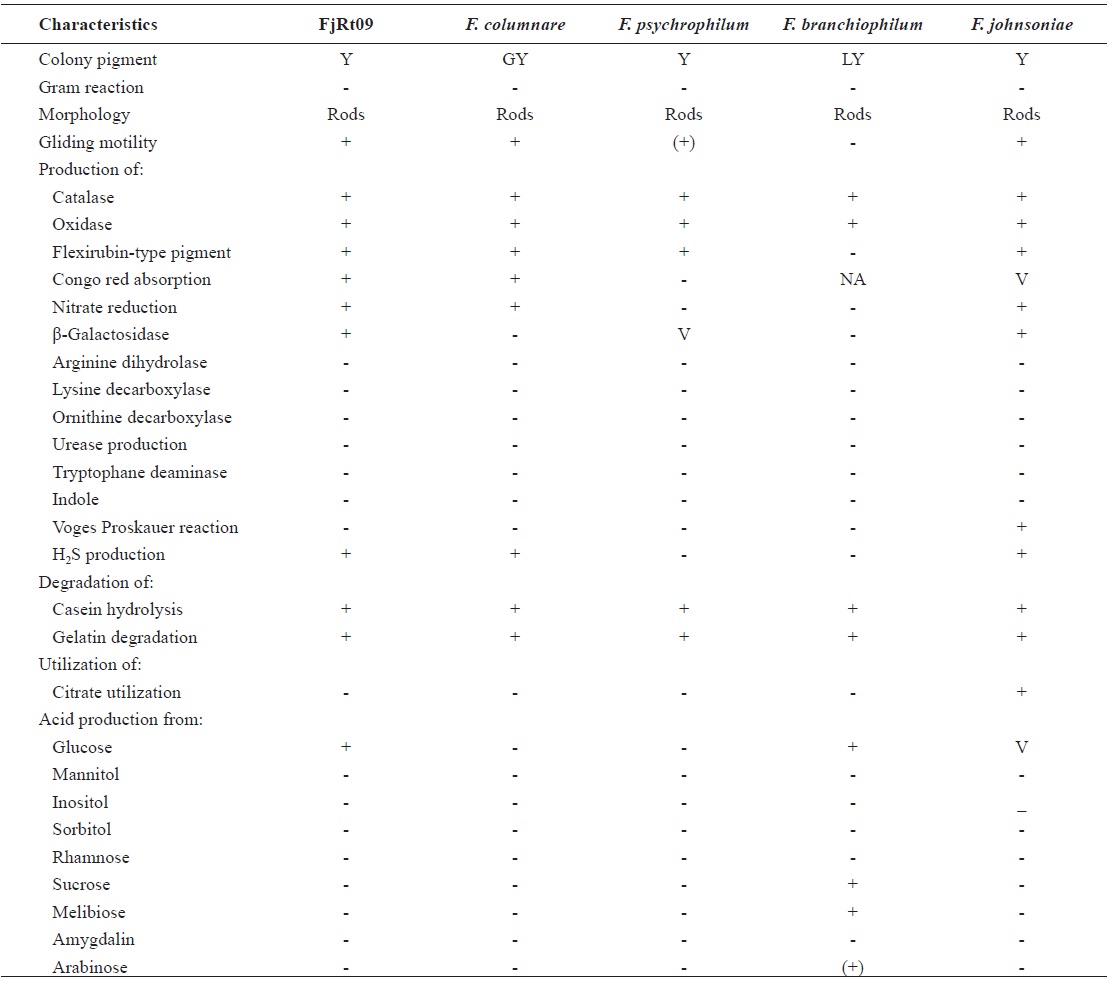
From 64 fish samples, a total of 231 colonies were observed on TSA plates after 72 h at 15℃, and 18 fish samples (28.1%) from two hatcheries had yellow-pigmented colonies, which were selected for further identification (Table 1). All 18 iso-lates showed the same biochemical types with characteristics including rapid gliding motility, Gram-negative rods, oxidase-positive, catalase-positive, flexirubin pigment, nitrate reduc-tion, β-galactosidase-positive, acid production from glucose, Congo red absorption, and hydrolysis of gelatin and casein. Growth occurred with 1% NaCl in tryptic soy broth (TSB) but not with 1.5% NaCl in TSB and no growth occurred at 4℃. The biochemical characteristics of the bacteria are sum-marized in Table 2.
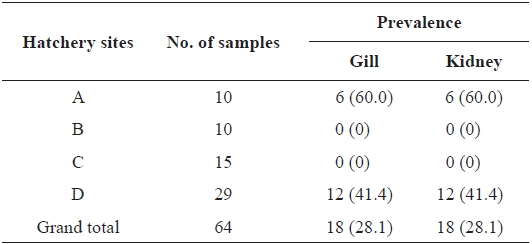
Prevalence of Flavobacterium johnsoniae in rainbow trout col-lected from four hatcheries in Gangwon Province, Korea
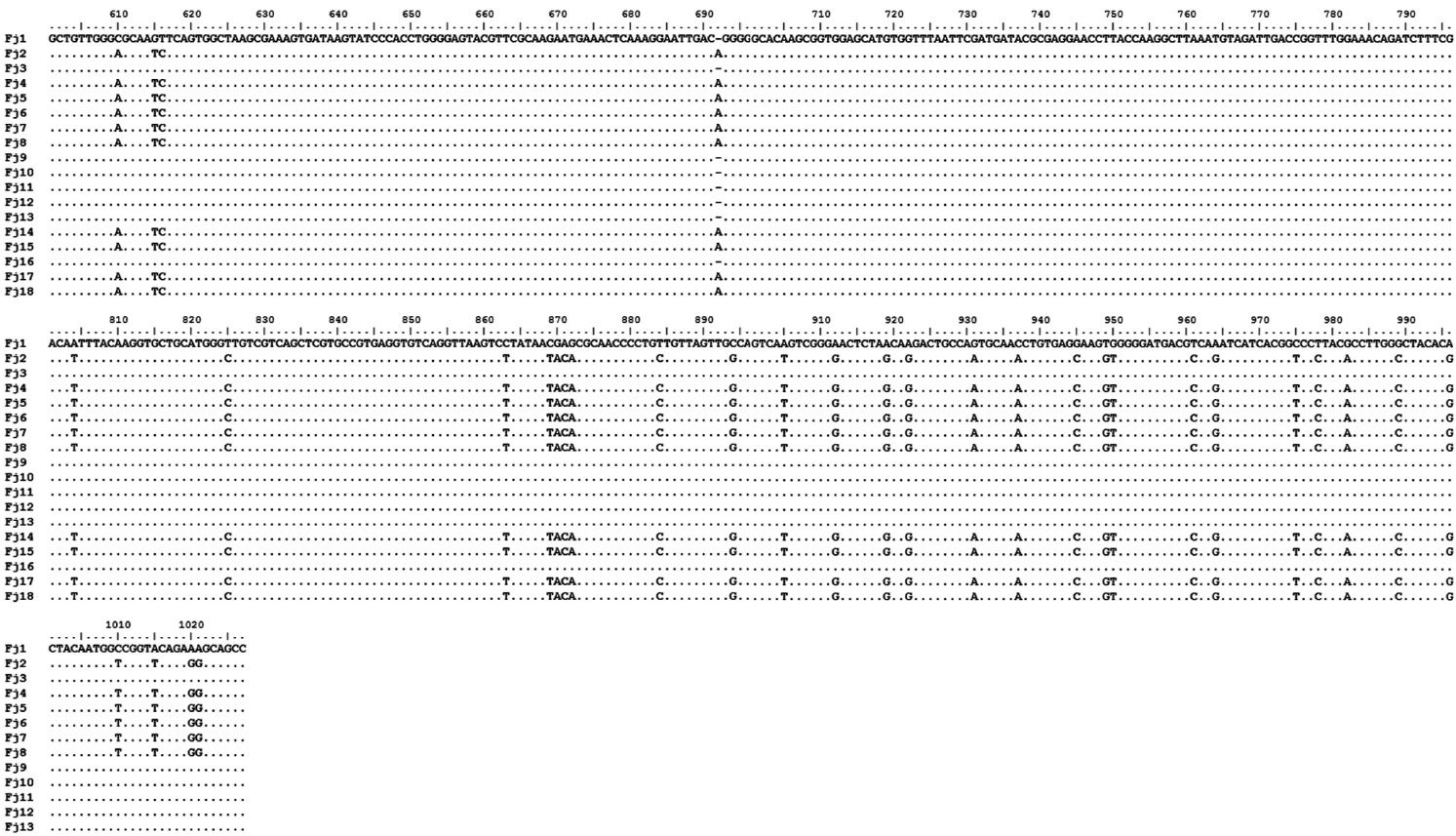
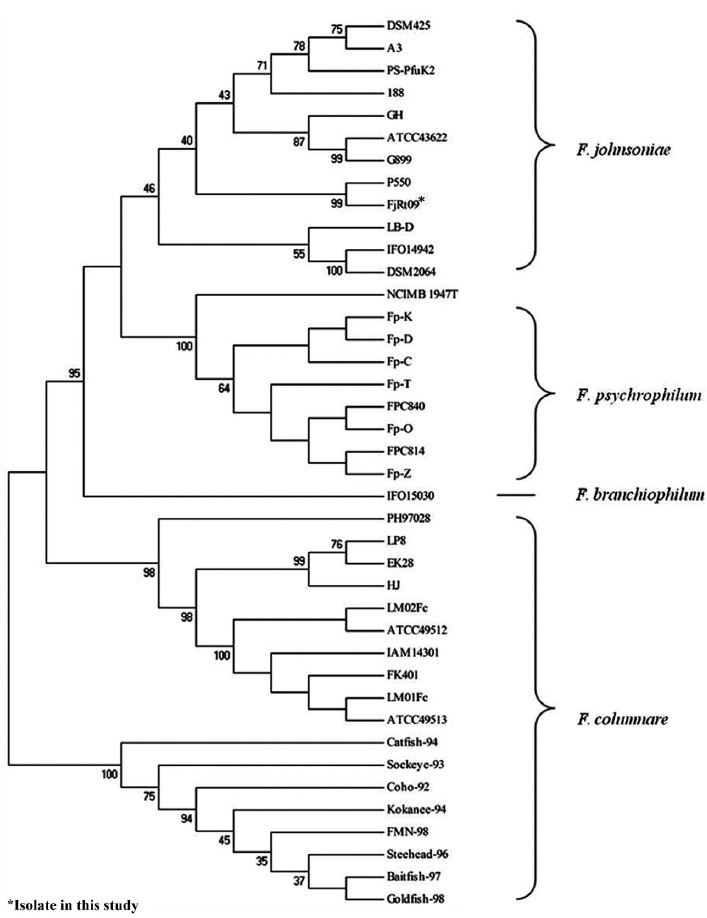
The amplification of 16S rRNA from all 18 yellow-pig-mented colonies showed a band of approximately 1,300 bp (data not shown). Highly conserved sequences (97-99%) were found among the 18 isolates (Fig. 1), and one isolate was selected and named FjRt09 (GenBank accession no. GU461280). Phylogenetic tree analysis showed that FjRt09 had 98% homology and grouped more closely with
Gram-negative yellow-pigmented bacteria belonging to the genus
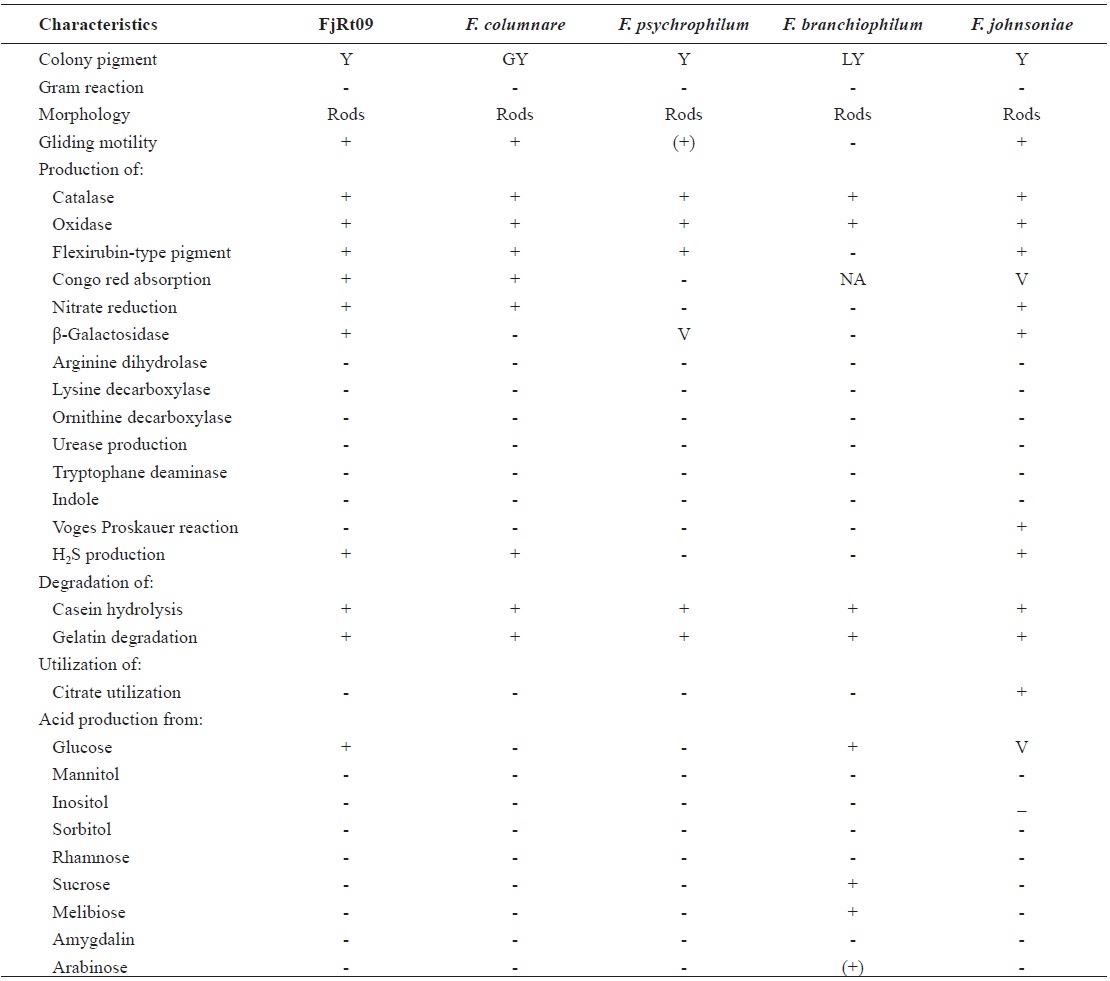
Phenotypic characteristics of Flavobacterium johnsoniae isolate from rainbow trout in Korea
lesions on fish,
and rearing density.
In this study, 18 out of 64 rainbow trout sample plates de-veloped yellow-pigmented colonies. The affected samples originated from two hatcheries, although the fish showed no external clinical signs. Biochemical characterization was conducted on the 18 yellow-pigmented colonies, and the bio-chemical characteristics of the bacterial isolates corresponded to those of