Artificially oxidized cysteine residues in peroxiredoxin 6 (Prx6) were detected by electrospray interface capillary liquid chromatography-linear ion trap mass spectrometry after the preparation of two-dimensional gel electrophoresis (2D-GE). We used Prx6 as a model protein because it possesses only two cysteine residues at the 47th and 91st positions. The spot of Prx6 on 2D-GE undergoes a basic (isoelectric point, pI 6.6) to acidic (pI 6.2) shift by exposure to peroxide due to selective overoxidation of the active-site cysteine Cys-47 but not of Cys-91. However, we detected a tryptic peptide containing cysteine sulfonic acid at the 47th position from the basic spot and a peptide containing both oxidized Cys-47 and oxidized Cys-91 from the acidic spot of Prx6 after the separation by 2D-GE. We prepared two types of oxidized Prx6s: carrying oxidized Cys-47 (single oxidized Prx6), and other carrying both oxidized Cys-47 and Cys-91 (double oxidized Prx6). Using these oxidized Prx6s, the single oxidized Prx6 and double oxidized Prx6 migrated to pIs at 6.2 and 5.9, respectively. These results suggest that oxidized Cys-47 from the basic spot and oxidized Cys-91 from the acidic spot are generated by artificial oxidation during sample handling processes after isoelectric focusing of 2D-GE. Therefore, it is important to make sure of the origin of cysteine oxidation, if it is physiological or artificial, when an oxidized cysteine residue(s) is identified.
During the past decade, proteome analysis has utilized two-dimensional gel electrophoresis (2D-GE) for protein separation and peptide mass finger printing using mass spectrometry.1 In an extensive research on proteome analysis, protein identification and post-translational modifications have been major research fields. Protein modifications caused by oxidative stress, such as oxidation of amino acid side chains, are important therapeutic targets.2-4
Cysteine is an amino acid residue sensitive to oxidative reaction. The sulfhydryl group of cysteine residue exists not only as free-SH and -S-S-, but also in an overoxidation form
Human umbilical vein endothelial cells (HUVECs) were purchased from Cambrex (East Rutherford, NJ, USA) and cultured as described previously.8 Cells were lysed in a sample buffer containing 9 M urea, 2% (w/v) 3-([3-cholamidopropyl] dimethylammonio)-1-propane sulfonate (CHAPS), 65 mM dithioerythritol, and 0.5% carrier ampholyte (pH 4-7; GE Healthcare, Tokyo, Japan). The cell lysate was applied to an immobilized pH gradient gel (13 cm Immobiline Drystrip, pH 4-7; GE Healthcare) and rehydrated at room temperature for 12 h. Isoelectric focusing (IEF) was performed for a total of 48,990 Vhrs at a maximum voltage of 8,000 V. The strips were equilibrated by two steps for 20 min in an equilibration buffer containing 6 M urea, 2% sodium dodecyl sulfate (SDS), 30% glycerol, and 50 mM Tris-HCl buffer (pH 8.8), and supplemented with 10 mg/mL dithiothreitol (DTT) for the first equilibration and 40 mg/mL iodoacetamide (Wako Pure Chemicals, Osaka, Japan) for the second equilibration. Sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) was carried out with 12.5% gel. Gels were stained with SyproRuby (Invitrogen, Carlsbad, CA, USA) and scanned with Molecular Imager FX (Bio-Rad Laboratories, Irvine, CA, USA). In-gel digestion of the protein spot was performed as described previously.8
Two peptides, Prx6(42-53) (DFTPVCTTELGR) and Prx6(85-97) (DINAYNCEEPTEK) were synthesized using a PSSM-8 automated peptide synthesizer (Shimadzu, Kyoto, Japan) according to standard 9-fluorenylmethyl-oxycarbonyl (Fmoc) chemistry and purified by reversed-phase HPLC. Cysteine residues of these two peptides were oxidized by performic acid and carbamidomethylated by iodoacetamide, respectively. To prepare performic acid, 50 μL of 30% H2O2 was added to formic acid (450 μL). Twenty-five microliter of the performic acid solution was added to 100 μL of peptide solution (0.1 mg/mL) and incubated at 4℃ for 1 h. Carbamidomethylation of the cysteine residue was conducted by adding 140 μL iodoacetamide (100 mg/mL) in 50 mM NH4HCO3 to 100 μL of peptide solution (10 mg/mL), followed by incubation at room temperature for 1 h. These reaction mixtures were purified by reversed-phase HPLC. These peptides were quantified by the ninhydrin method of amino acid analysis after acid hydrolysis at the Peptide Institute Inc. (Osaka, Japan).
Purified recombinant Prx6 (LabFrontier, Seoul, Korea) was oxidized by H2O2 or performic acid and digested with trypsin, as described previously.13 Matrix-assisted laser desorption ionization time-of-flight (MALDI-TOF) mass spectrometry was performed using a Voyager DE STR MALDI-TOF mass spectrometer (Applied Biosystems, Foster City, CA, USA) in negative reflector mode. α-cyano-4-hydroxycinnamic acid was used as matrix.
Capillary HPLC separation was conducted using a HPLC system (Paradigm MS4D; Michrom BioResources, Auburn, CA, USA). The system was coupled on-line to a linear ion trap mass spectrometer (Finnigan LTQ; ThermoFisher Scientific, Waltham, MA, USA). Chromatographic separation was performed on a reversed-phase capillary column (MAGIC C18, 3 μm, 50 × 0.2 mm i.d., Michrom BioResources) at a flow rate of 1.8 μL/min. The gradient condition consisted of a linear gradient from 5% solvent B (H2O/acetonitrile/formic acid, 10/90/0.1, v/v/v) to 65% B against solvent A (H2O/acetonitrile/formic acid, 98/2/0.1, v/v/v) in 20 min. Electrospray ionization was performed using an electrospray emitter (FortisTip, 50 mm × 20 μm i.d.; OmniSeparo-TJ, Hyogo, Japan).14 The automatic gain control was tuned with target values of 5.0 × 104 and 1.0 × 104 for the full scan and tandem mass spectrometry (MS/MS) scan, respectively. Maximum ion injection time was set at 200 ms for the full scan and 400 ms for the MS/MS scan. For protein identification, data-dependent MS/MS data were submitted to the MASCOT (Matrix Science, London, UK) search software using SWISS-PROT as a database. For peptide quantification, MS/MS was performed using an automated data-dependent MS/MS procedure and parent mass list. Masses of doubly charged ions of carbamidomethylated and oxidized peptides were as in the followings: carbamidomethylated Prx(42-53), 698.3; oxidized Prx(42-53), 693.8; carbamidomethylated Prx(85-97), 791.8; and oxidized Prx(85-97), 787.3. This procedure consisted of three scan events: a full scan with a mass range of
>
Identification and structure of oxidatively modified cysteine residues in Prx6 on 2D-GE
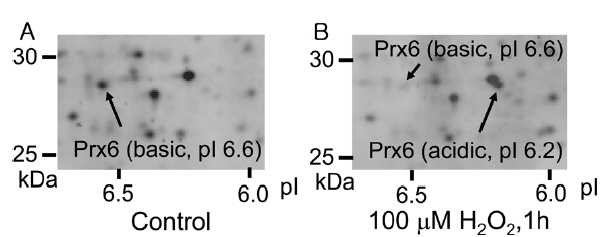
By the addition of peroxide, such as H2O2, in a cell culture medium, some cellular proteins undergo an acidic shift in their isoelectric points (pIs). In case of peroxiredoxins, their pIs decrease by 0.2-0.4 on 2D-GE due to oxidation of active-site cysteine upon exposure of cells to H2O2. 12 Figure 1 shows 2D-GE mapping of cultured HUVEC (A) and HUVEC exposed to 100 μM H2O2 for 1 h (B). The spots of basic Prx6 and acidic Prx6 were identified by capillary HPLC-MS/MS and MASCOT sequence database search (data not shown). Under control conditions, the basic Prx6 appeared at a molecular mass of 28.7 kDa and a pI of 6.6 (Figure 1A). However, upon H2O2 treatment, the basic spot of Prx6 disappeared and an acidic spot of Prx6 appeared at 28.7 kDa (molecular mass) and a pI of 6.2 (Figure 1B). To determine the modified structure of basic and acidic spots
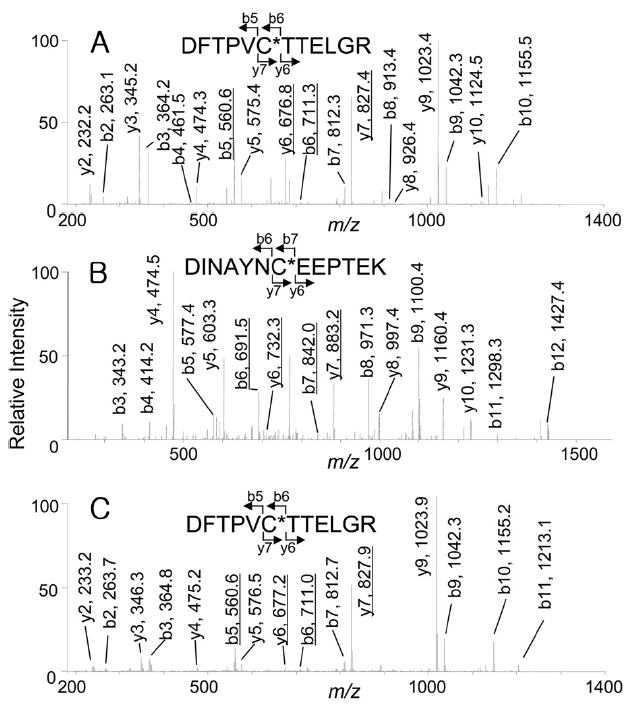
of Prx6, these spots were excised, digested with trypsin, and analyzed by capillary HPLC-MS/MS. The expected peptides containing Cys-47 or Cys-91, were Prx6 (42-53) (DFTPVCTTELGR, monoisotopic mass: 1337.5) or Prx6 (85-97) (DINAYNCEEPTEK, monoisotopic mass: 1524.6). As shown in Figure 2A, when the
indicate that Cys-47 was oxidized to cysteine sulfonic acid (oxCys-47). However, from the acidic spot of Prx6, doubly charged ions corresponding to peptides carrying cysteine sulfonic acid were selected for the precursor ions, 151 Da differences were observed in both Figure 2B (b6 and b7, y6 and y7) and Figure 2C (b5 and b6, y6 and y7) that showed the presence of both oxCys-47 and oxCys-91 in the acidic spot of Prx6.
>
pI shifts on oxidation of cysteine residues on 2D-GE
Cys-47, but not Cys-91, in Prx6 was selectively oxidized by peroxide treatment to form an acidic spot on 2D-GE, and in the basic spot, the Cys-47 remained in a reduced form.7,11,12 However, our results obtained by 2D-GE implied that both cysteine residues in the acidic spot were possibly oxidized by peroxide treatment, and oxidized cysteine residue was found even in the basic spot. If one or two cysteine residues are selectively oxidized, how much is the pI shift? To answer this question and explain the correlation between a pI shift and the number of oxidized cysteine, we prepared two types of oxidized Prx6s. The first was oxidized Prx6 (oxPrx6) containing oxCys-47 and a reduced form of Cys-91 (single oxidized Prx6). The other was oxPrx6 containing oxCys-47 and oxCys-91 (double oxidized Prx6). To prepare two types of oxPrx6s, purified recombinant Prx6 was oxidized by H2O2 or performic acid and, yielded single oxidized Prx6 or double oxidized Prx6. To confirm the structures, these two oxPrx6s were treated with iodoacetamide to protect from further oxidation of free cysteine, digested with trypsin, and measured by a MALDI-TOF mass spectrometer in negative ion mode. In case of
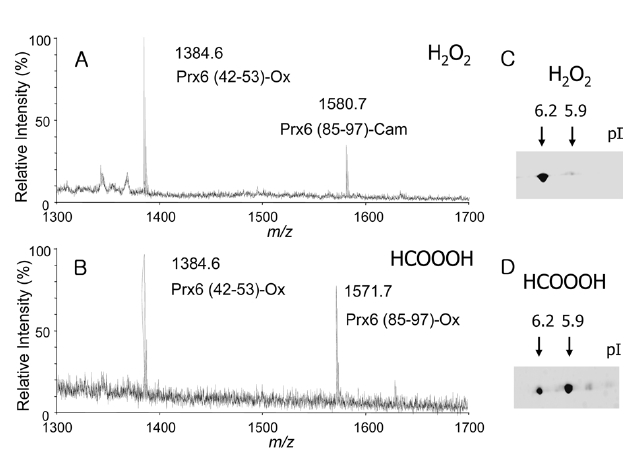
H2O2 treatment, observed masses of 1384.6 and 1580.7, corresponding to the deprotonated ion of Prx6(42-53) carrying oxCys-47 (Prx6 (42-53)-Ox) and the Prx6 (85-97) containing carbamidomethyl cysteine (Prx6 (85-97)-Cam), were detected (Figure 3A). In case of performic acid treatment, masses of 1384.6 and 1571.7, corresponding to the deprotonated ions of Prx (42-53)-Ox and Prx6 (85-97)-Ox, were detected (Figure 3B). These results clearly indicated that the H2O2-treated Prx6 possessed one cysteine sulfonic acid at the 47th position (single oxidized Prx6), whereas, the performic acid-treated Prx6 possessed two cysteine sulfonic acids at the 47th and 91st positions (double oxidized Prx6). The pIs of these two oxPrx6s were checked by 2D-GE. Under control conditions, purified recombinant Prx6 had pI of 6.6. After H2O2 treatment, an acidic spot with a pI of 6.2 appeared (Figure 3C). Moreover, in case of performic acid treatment, a more acidic spot with a pI of 5.9 appeared (Figure 3D). Taken together, these data indicated that oxidation of one cysteine in Prx6, oxCys-47, induced an acidic shift in pI of 0.4 (6.6→ 6.2), whereas, oxidation of both Cys-47 and Cys-91 residues resulted in a pI shift of 0.7 (6.6→ 5.9). pI shifts on 2D-GE may be proportional to the number of oxidized cysteine, in the case of phosphorylation of proteins.15
>
Difference in ionization efficiencies of peptides containing modified cysteine
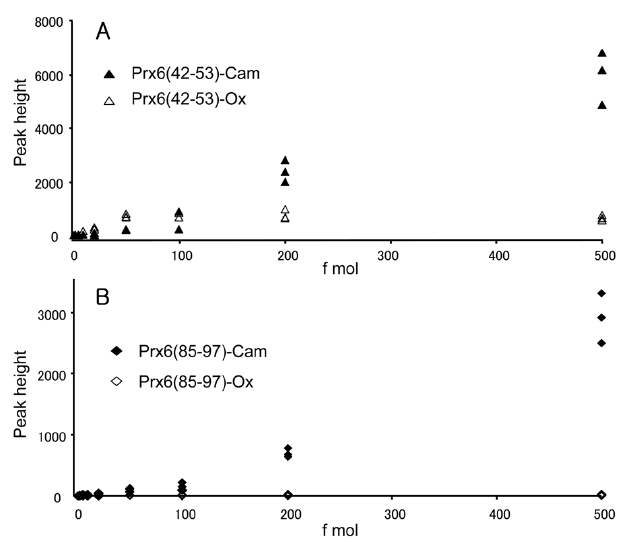
We observed one oxidized cysteine residue (oxCys-47) from the basic spot of Prx6 (pI 6.6) and two oxidized cysteine residues (oxCys-47 and oxCys-91) from the acidic spot (pI 6.2) on the 2D-GE. The presence of oxCys-91 in the acidic spot and oxCys-47 from the basic spot had not been reported previously. This is probably because of the small quantity of oxidized peptides and a lack of sensitivity for detection. To clarify the change of sensitivity in this measurement caused by oxidation of cysteine, we compared signal intensities of specific fragment ions of peptides obtained by capillary HPLC-MS/MS. This selected reaction monitoring (SRM)-like analysis enabled us to achieve a highly sensitive detection of a small quantity of peptides by lowering the background. The equimolar mixture of four synthetic peptides, Prx6 (42-53)-Ox, Prx6 (42-53)-Cam, Prx6 (85-97)-Ox and Prx6 (85-97)-Cam were prepared and subjected to capillary HPLC-MS/MS analysis in the range of 1-500 fmol. Doubly charged ions of these peptides were selected as the precursor ions, and product ions of y9 (for Prx6 (42-53)-Ox and Prx6 (42-53)-Cam) and y4 (for Prx6 (85-97)-Ox and Prx6 (85-97)-Cam) ions were used for these measurements. Correlations between peptide amounts and peak heights on this SRM measurement are shown for Prx6 (42-53)-Ox and Prx6 (42-53)-Cam (Figure 4A) and for Prx6 (85-97)-Ox and Prx6 (85-97)-Cam (Figure 4B). The carbamidomethylated peptide gave higher intensities than the oxidized peptide in both cases. In case of the carbamidomethylated peptides, a linear and positive response
was observed, whereas oxidized peptides exhibited lower slopes than those of the carbamidomethylated peptides. These results indicated that the ionization efficiency of peptides containing cysteine sulfonic acid decreased on oxidation of cysteine in this low-flow HPLC-MS.
The active-site cysteine Cys-47, but not Cys 91, independent of the enzymatic activity, is selectively oxidized by the treatment with peroxide.7, 11, 12 However, in this study, we found oxCys-47 in the basic spot (pI 6.6) and two of oxidized cysteine residues (oxCys-47 and oxCys-91) in the acidic spot (pI 6.2) on 2D-GE of cell lysate with or without H2O2 treatment. Next, we prepared non-oxidized, single oxidized (carrying cysteine sulfonic acid at the 47th position), or double oxidized Prx6 (carrying cysteine sulfonic acid at the 47th and 91st positions) using recombinant Prx6 and examined their pIs using 2D-GE. Results showed that the non-oxidized (does not contain cysteine sulfonic acid), single oxidized, and double oxidized Prx6 spots appeared at pIs of 6.6, 6.2, and 5.9, respectively. Taken together, these results strongly suggested that oxCys-47 obtained from the basic spot (pI 6.6) and oxCys-91 obtained from the acidic spot (pI 6.2) of Prx6 from cell lysate were artificially oxidized products.
If artificial oxidation occurred prior to the IEF, the oxidized Prx6 was thought to migrate to the appropriate pI corresponding to the number of oxidized cysteine. For example, when Cys-47 was artificially oxidized prior to IEF, the oxidized Prx6 should migrate to the pI of 6.2 corresponding to a single oxidized Prx6 (pI 6.2), and not show the pI of native Prx6 (6.6). In our experiment, the free sulfhydryl group is protected as carbamidomethylated cysteine by alkylation with iodoacetamide in the equilibration step after completion of IEF. However, a high concentration (6 M) of urea in IEF equilibration buffer renders certain sulfhydryl groups unable to react with iodoacetamide,16-18 therefore, not every cysteine is alkylated under certain condition. The remaining free cysteine may be oxidized during sample handling after the IEF step. Among handling processes, SDS-PAGE may be a suspect step because ammonium persulfate is used for polymerization. However, according to the results of Chiari et al., cysteine residue in protein can not be oxidized to cysteine sulfonic acid by milder oxidizing agents, such as ammonium persulfate and periodate.19 Methionine residue has been known to be easily oxidized to methionine sulfoxide. However, oxidation of methionine residue may not involve a pI shift because sulfoxide is not acidic like cysteine sulfonic acid. Methionine has been reported to be oxidized during SDS-PAGE by the remaining molecular oxygen. However, the use of photopolymerized SDS gel and thioglycolate for the cathode buffer as a reactive oxygen scavenger could eliminate artificial oxidation of methionine.20 Therefore, it is currently difficult to specify which steps significantly influence oxidation of cysteine, but the coexistence of a suitable scavenger during sample handling may make it possible to prevent artificial oxidation of cysteine residue.
To examine the relative sensitivity of peptides carrying cysteine sulfonic acid in a low-flow electrospray ionization mass spectrometry, we utilized two sets of peptides, cysteine sulfonic acid-containing peptides and carbamidomethylated cysteine-containing peptides. Low response in peptides containing cysteine sulfonic acid in positive ion mode is attributed to a lowering of the ionization efficiency because of the acidic modification of peptide. This tendency in electrospray is very similar to our previous work in MALDI.13 The extent of artificial oxidation could not be estimated because significant variations in day-to-day and run-to-run reproducibilities were observed in this low-flow capillary HPLC-MS/MS experiment.
Although we showed only an example using Prx6 as a model protein, cysteine residue may be artificially oxidized to cysteine sulfonic acid after the IEF step. Sensitivity of a recent capillary HPLC-MS system allows the analysis of a structure at less than a femto mole level. A highly sensitive analysis such as this may possibly detect an artificially oxidized cysteine and tends to confuse it with physiologically oxidized cysteine. Such non-selective oxidation appears to occur widely. Even if an oxidized cysteine residue is identified in a protein, it is problematic to conclude the physiological relevance without other evidence.