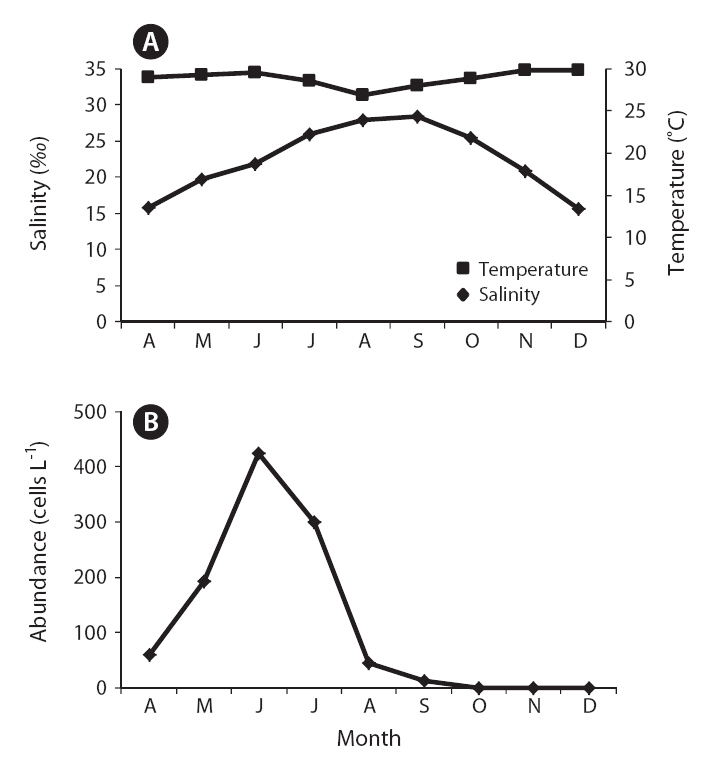
The toxic dinoflagellate Karlodinium veneficum has been implicated in numerous fish kill events around the world.Since this species commonly co-occurs with other morphologically similar dinoflagellates, field monitoring of this species in natural waters via light microscopy only has been problematic. In this study, we investigated temporal changes in K. veneficum’s abundance in the waters of Obido, Tongyeong, using a species-specific real-time polymerase chain reaction (PCR) assay. The field survey, from April to December 2010, revealed K. veneficum occurred at low densities (12 to 425 cells L?1) during this time and that cell numbers peaked in June (early summer in Korea), indicating this species generally occurs in the warmer season (mostly at 16.9-22.3°C and 33.4-34.5‰) in the Obido area.
>
Culture and analysis of environmental factors
We obtained
>
Water sample collection and DNA extraction
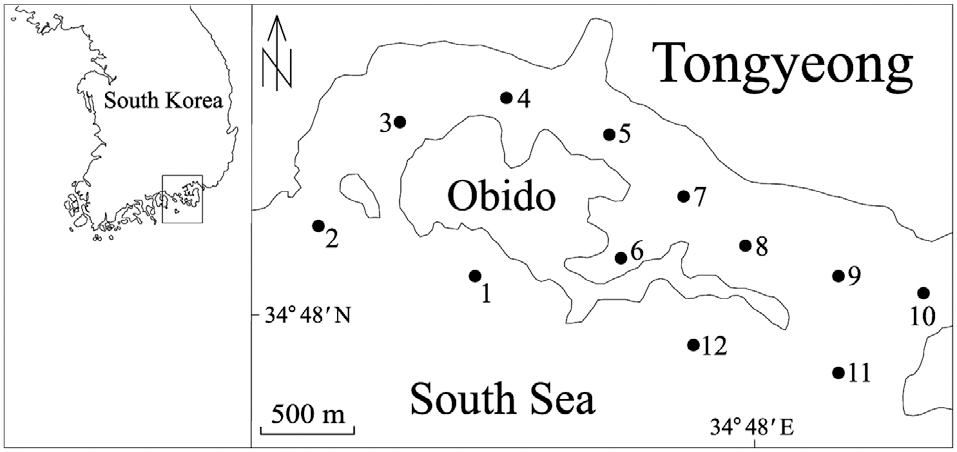
From April through December 2010, we collected 250 mL surface water samples, at 1 month intervals, from each of 12 stations in Obido, Tongyeong, Korea (Fig. 1). We filtered the water samples onto a 1.2 ㎛ pore, 25 mm diameter glass microfiber GF/C filter (Whatman Ltd., Maidstone, England), placed each filtered sample in a 2 mL microcentrifuge tube, and stored it at -70°C until DNA extraction. To prevent target DNA degradation, we processed these filtering samples upon a research vessel and extracted the samples genomic DNA from the samples within 2 months, using a phenol-chloroform extraction protocol (Hosoi-Tanabe and Sako 2005).
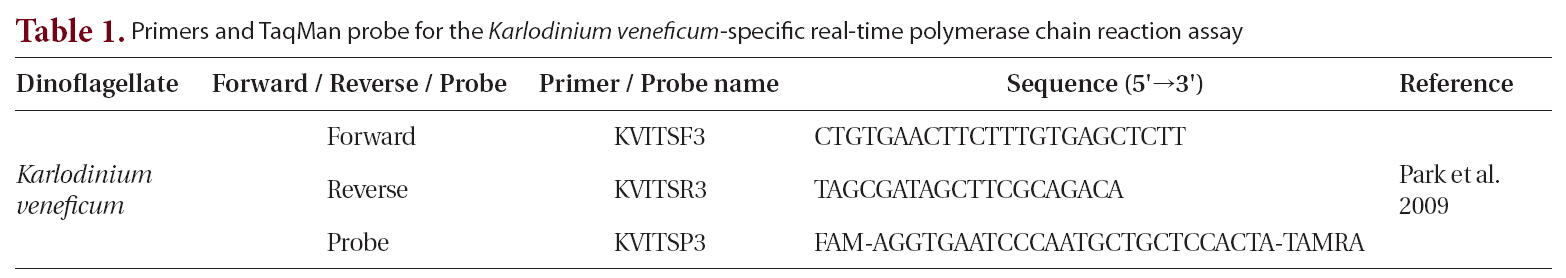
We used the following reagents to create the reaction mixture, to a final volume of 10 μL : 5 μL of platinum quantitative PCR supermix-UDG (Invitrogen, Eugene, OR, USA); forward and reverse primers, each at a final concentration of 0.3 μM; fluorogenic probe at a final concentration of 0.1 μM; 0.5 μL of template DNA (Park et al. 2009); and PCR grade water (Table 1). The thermal cycling condition comprised 2 min at 50°C and 2 min at
12
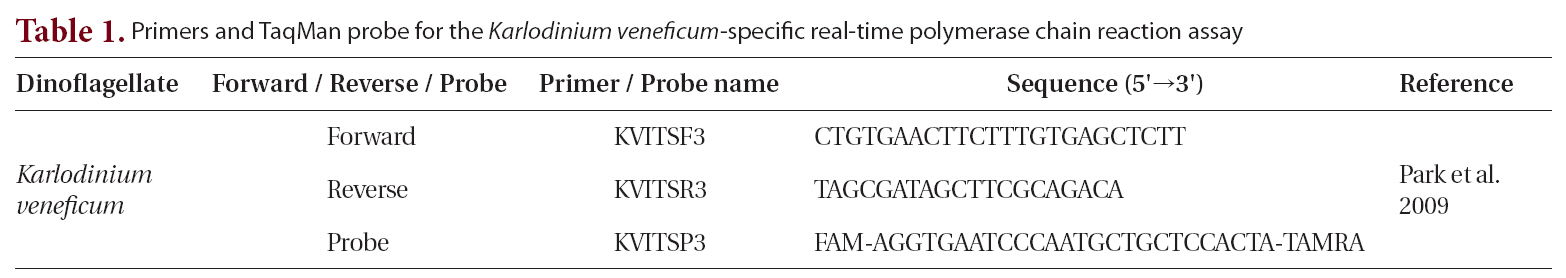

Primers and TaqMan probe for the Karlodinium veneficum-specific real-time polymerase chain reaction assay
95°C, followed by 45 ten-s cycles at 95°C and 45 s at 60°C. All samples were analyzed in triplicate. We collected the fluorescence data at the end of each cycle, while a Rotor-Gene 6000 instrument (Corbett Research, Sydney, Australia) automatically determined the cycle threshold. To remove the surface waters’ PCR inhibitors, we diluted the template DNA tenfold before use.
>
Construction of the standard curve for cell quantification
Before harvesting the cells (12,000 cells), we collected the laboratory-cultured
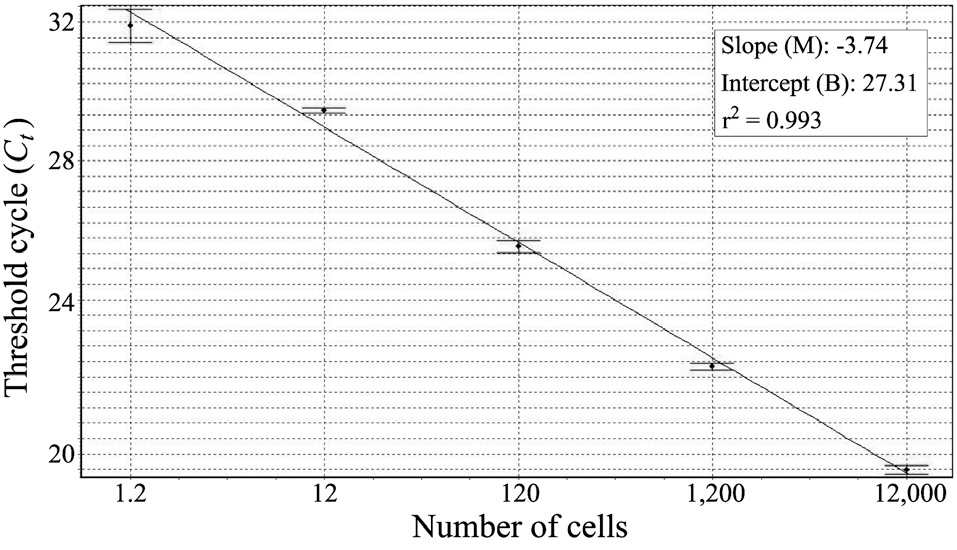
We constructed a standard curve with known cell numbers and established a strong linear relationship between the
The field survey of the Obido waters showed that
Researchers have used a number of molecular methods, including real-time PCR, fluorescent
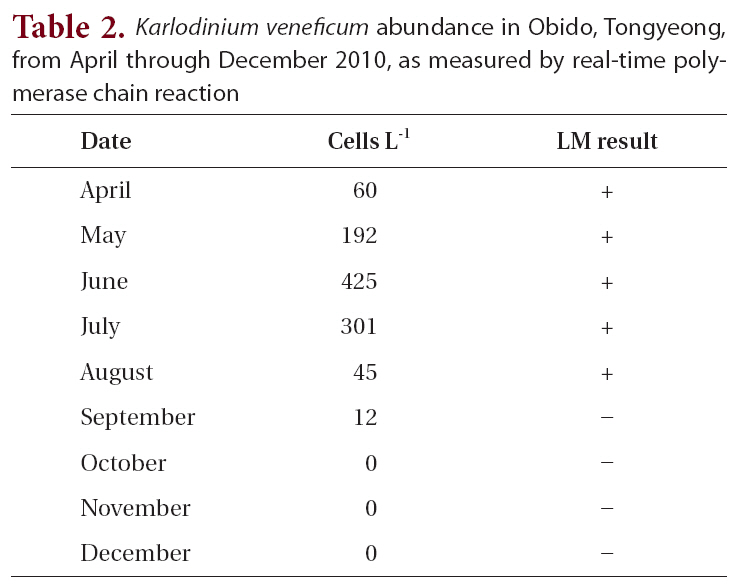
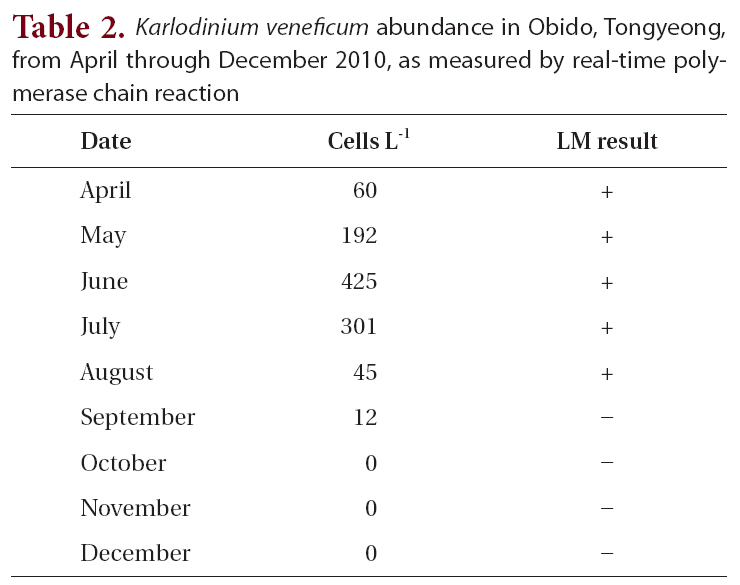
Karlodinium veneficum abundance in Obido Tongyeong from April through December 2010 as measured by real-time polymerase chain reaction
the real-time PCR could theoretically detect even one copy of the dinoflagellate’s DNA). The rDNA-based real-time PCR probe possesses such a high sensitivity because the rDNA in most eukaryotes is repeated in tandem at a high copy number. For example, one study reports 100 to 200 copies in one
In conclusion, the real-time PCR assay was highly sensitive and specific for detecting and quantifying