An increasing number of studies are being conducted into the role of changes in gene expression on the physiology, biology and reproduction of organisms. These gene expression studies often employ real-time quantitative polymerase chain reaction (RT-qPCR). RT-qPCR can also be used to validate other studies, including microarray data and also more frequently RNA-seq data using a measure called RPKM (Mortazavi et al. 2008). While microarray data and RNA-seq cover a large array of genes, RT-qPCR is still the gold standard for expression studies of target genes. RT-qPCR relies on an internal PCR standard for which to compare changes in gene expression of target genes between experimental conditions. These standards, so called ‘housekeeping’ genes or reference genes, transcripare from genes that are assumed to be necessary for regular physiological maintenance of the organism, and are considered to not alter during the experimental manipulations or different life history stages. Studies have shown that equal expression of ‘standard’ housekeeping genes (HKGs) can not be assumed and must be tested for each organism under study (e.g., Ransbotyn and Reusch 2006, Hibbeler et al. 2008, Wu et al. 2013).
Transcriptomics is a powerful way to study an array of genes that could be important in various biological process. To find genes that are over- or under-expressed in particular stages, and therefore could be causative, it is critical that a good internal standard is established. These internal standards (HKGs) ideally are expressed at near similar levels in all experimental conditions, so to that changes in expression of target genes can be calibrated for handling errors (normalization). HKGs have been evaluated for the red alga
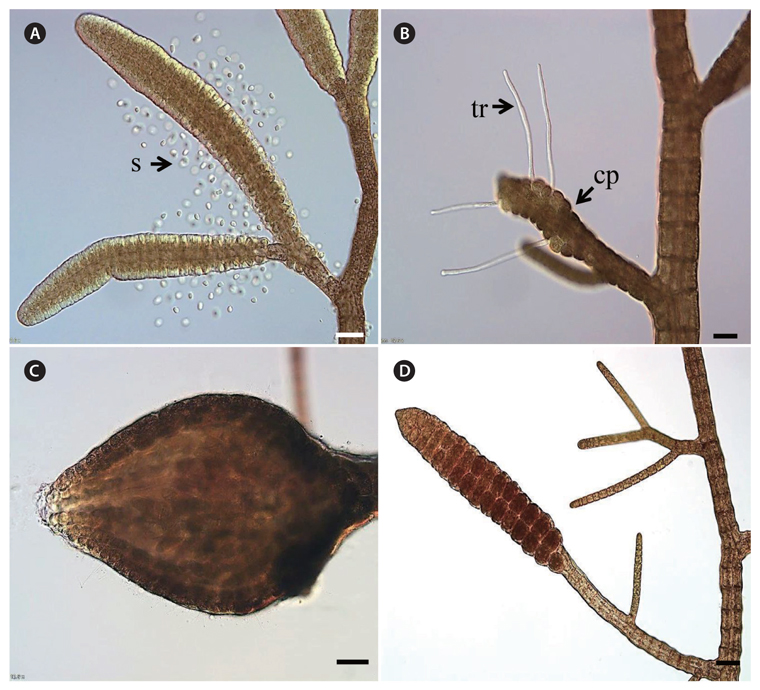
Haploid gametophytes (male and female), females containing carposporophytes and free-living diploid sporophytes of
>
RNA extraction and cDNA synthesis
Total RNA were isolated from the different life history stages using the RNeasy Plant RNA extraction kit (Qiagen, Gaithersburgh, MD, USA) according to manufacturer’s protocol. RNA isolation was performed three times for the plants of each life history stages. Genomic DNA contamination was eliminated by on-column digestion with RNase-free DNase I. RNA quality and concentration were determined using gel electrophoresis and spectrophotometrically. RNA samples with 260/280 ratio from 1.9 to 2.1, 260/230 ratio from 2.0-2.5 were used for further analysis. First-strand cDNA was synthesized from 3 μg of total RNA using the iScriptTM cDNA synthesis kit (Bio-Rad Laboratories, Hercules, CA, USA) in a final reaction volume of 20 μL with random primers following the manufacturer’s protocol. All cDNAs were diluted 10 times with autoclaved milli-Q water and stored at -20°C until use.
>
HKG selection and primers design
Potential HKGs which are popularly used for RT-qPCR analysis were selected from a
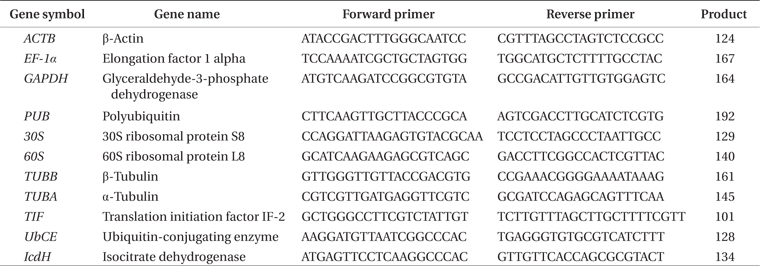
[Table 1.] Primers used for real-time PCR
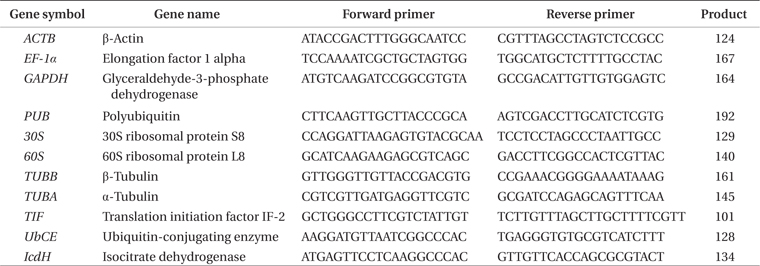
Primers used for real-time PCR
>
qPCR and statistical analysis
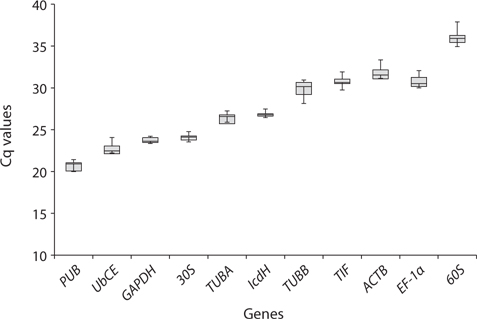
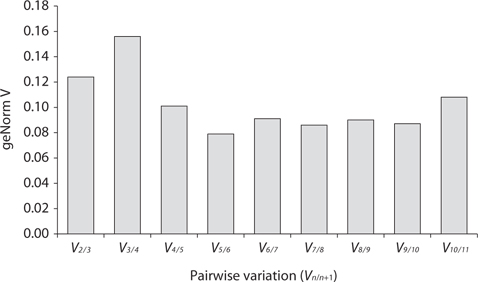
Real-time PCR was performed using iQ SYBR Green supermix (Bio-Rad Laboratories) in a real-time PCR detection system (Bio-Rad CFX96; Bio-Rad Laboratories). cDNA samples were amplified in triplicate from the same RNA preparation. The final reaction volume was 20 μL and included 5 μL of diluted cDNA, 10 μL of iQ SYBR Green supermix and 250 nM of each primers (Table 1). The reaction protocol was first set at 95°C for 3 min and then 40 cycles of 95°C for 15 s and 60°C for 20 s. For each candidate HKG, the specificity of primers and amplified single product was verified by analyzing the melting curves and gel electrophoresis. The expression of each gene was determined from Cq values (synonym, Ct value) that corresponds to a number of cycles necessary for the PCR amplification to reach a threshold. Cq variability data in HKG comparisons were expressed as whiskers-box plot for all pooled samples. Selected HKGs were tested for the stability of their expression in different life history stages following a normalization strategy using two different algorithms: geNorm using the geNorm plugin of qbase + program (Vandesompele et al. 2002, Hellemans et al. 2007) and Normfinder (Andersen et al. 2004). The expression stability of all candidate HKGs was evaluated by comparing the differences of observed Cq values across life-history stages. Pairwise variation (
The haploid gametophytes and diploid sporophytes of
The stability of expression of HKGs in
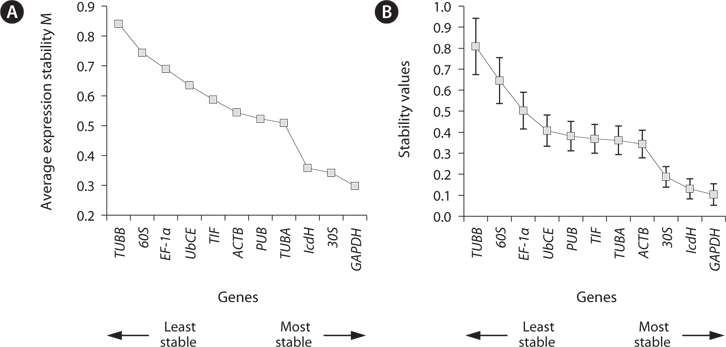
The stability of the gene expression in the different life history stages was analyzed (Fig. 3). Analysis using geNorm showed that
In
The temporal, developmental and physiological pattern of gene expression is a topic that is now amiable to scientific study. Changes in gene expression, even fairly subtle changes, could have significant effects on the physiology of the organism of study (Kim et al. 2016). To accurately quantify these gene expression changes a stable comparison is needed, to account for variation in RNA quantification and other experimental error. Accurate normalization is an absolute prerequisite for accurate measurements of gene expression changes, yet the determination of these HKGs (reference genes) could be a circular problem as the expression level of these reference genes also needs to be normalized (Hibbeler et al. 2008). The search therefore for genes that are stable over multiple conditions or life history stages is critical and several methods have been established (geNorm in Vandesompele et al. 2002, NormFinder in Andersen et al. 2004). Often normalization procedures to validate candidate genes are not done and HKGs are used that have been used in other organisms or studies, without any validation of their stability in the specific study (Chan et al. 2014). While validation methods are essential to have a stable platform from which changes can be measured between samples, in red algae, few studies have validated HKGs before measuring differences in target genes (e.g., Asamizu et al. 2003 used 60S L3 protein). This could lead to erroneous conclusions on the changes in expression level in target genes. Our results showed that, at least between life history stages of
Tests for HKG stability in life history stages of the red alga has recently been performed in
It is clear from our analysis, and several other studies, that housekeeping genes are not equally expressed and the expression of ‘standard’ housekeeping genes must be tested for each organism under study. Although RT-qPCR is expensive and time consuming a survey of potential candidate genes at the start of a study would be essential in reducing uncertainty, increasing confidence and may lessen the need for additional experiments to confirm any results.