Porcine reproductive and respiratory syndrome virus (PRRSV) is the causative agent of the most important infectious disease affecting the porcine herds worldwide. It is an enveloped virus with a single-stranded positive sense linear RNA genome of approximately 15 kb. PRRSV belongs to the family
SUMO (small ubiquitin-related modifier) is an ubiquitin-related protein that functions by covalent attachment to other proteins. SUMO and its associated enzymes are present in all eukaryotes and are highly conserved from yeast to humans (Jentsch and Pyrowolakis, 2000; Melchior, 2000; Muller
Because the PRRSV structural proteins are produced in low levels in the infected cells, a recombinant baculovirus has been often used to express these proteins. Recently, we expressed CA (Korean PRRSV strain) structural proteins using bApGOZA system in Sf21 cells; however, the expression level was very low (Koo
The CA strain (KCTC 11131BP) was isolated from the sera of field pigs at the acute stage of PRRSV infection in 2006. The virus was passaged twice on sub-confluent monolayers of MARC-145 cells in minimum essential medium supplemented with 10% fetal bovine serum (Gibco BRL, Paisley, UK), nonessential amino acids, sodium pyruvate, and antibiotics in 5% CO2 at 37℃. The virus was collected when 70% of the cells showed cytopathology. The supernatant was stored at -80℃ as the viral stock. Bm5 cells were cultured at 27℃ in TC-100 insect medium (WelGENE, Daegu, Korea) supplemented with 5% fetal bovine serum.
The larvae of the silkworm,
Viral RNA was extracted from infected cell cultures with Viral Gene-spinTM kit (iNtRON Biotechnology, Seoul, Korea) as recommended by the manufacturer. The viral RNA was used as a template for cDNA synthesis using RNA LA PCR Kit (TaKaRa, Shiga, Japan). Primers were based on the sequence of our previously sequenced PRRSV CA strain(Koo
>
Construction of transfer vector
The purified RT-PCR products were cloned into the T&A cloning vector (RBC Bioscience, Taiwan), and the plasmids were named pTRSV-ORF4, -ORF5, and -ORF6, respectively. They were digested with
>
Generation of recombinant baculoviruses
Bm5 cells were co-transfected with a mixture of purified transfer vector each containing a PRRSV gene, bBpGOZA DNA (Je
Bm5 cells were infected with wild type BmNPV or recombinant virus in a 60-mm diameter dish (2×106 cells) at a multiplicity of infection (MOI) of 5. At 3 d after inoculation, the culture supernatant was harvested and the infected cells washed with PBS, and then used to prepare SDS-PAGE sample. Individual silkworm larvae on the first day of the 5th instar were injected with 1×105 pfu of wild type BmNPV or recombinant virus. At 3 to 5 d post-injection, the hemolymph and fat body were collected by cutting a caudal leg and dissection, respectively. A few crystals of phenylthiourea were added to the tubes to prevent melanization. The fat body was homogenized in 10 volumes of lysis buffer (20 mM Tris-HCl pH 7.5, 50 mM NaCl, 5% Glycerol, 0.1% Triton X-100 containing protein inhibitor cocktail (Sigma-Aldrich, USA)) for 3 min and incubated in an ice bath for 30 min. The homogenate was centrifuged at 13,000 rpm for 10 min. The supernatant contained the total protein extract. The hemolymph was centrifuged at 10,000×g for 10 min to remove hemocytes and cell debris, and the supernatant was stored at -70℃ until further use.
>
Purification of 6xHis-tagged proteins
Lysis buffer was added to the fat body of
>
SDS-PAGE and western blot analysis
The prepared protein samples from cells and larvae were mixed with sample buffer, boiled for 5 min, and subjected to 12% SDS-PAGE gel. For Coomassie stains, gels were washed with deionized water and stained with BioSafe Coomassie. For Western blot analysis, the proteins were transferred to a nitrocellulose membrane (Pall Corp., NY, USA). After blotting, the membrane was blocked by incubation in 5% (w/v) non-fat dry milk in TBST buffer (20 mM Tris-HCl pH 7.5, 150 mM NaCl, 0.05% Tween 20) for 1 h with gentle agitation. The blot was incubated with 6xHis tag (1:1000) or porcine anti-PRRSV (1:100) antibodies (Choongang Vaccine Lab., Daejeon, Korea) in TBST for 1 h and washed. Subsequently, the membrane was incubated with anti-mouse and anti-pig IgG horseradish peroxidase conjugate for 30 min at room temperature. After repeated washing, the immunoreactive bands were visualized using the ECL Western Blotting Detection System (Elpis Biotech, Daejeon, Korea).
>
Expression of the PRRSV structural proteins in Bm5 cells
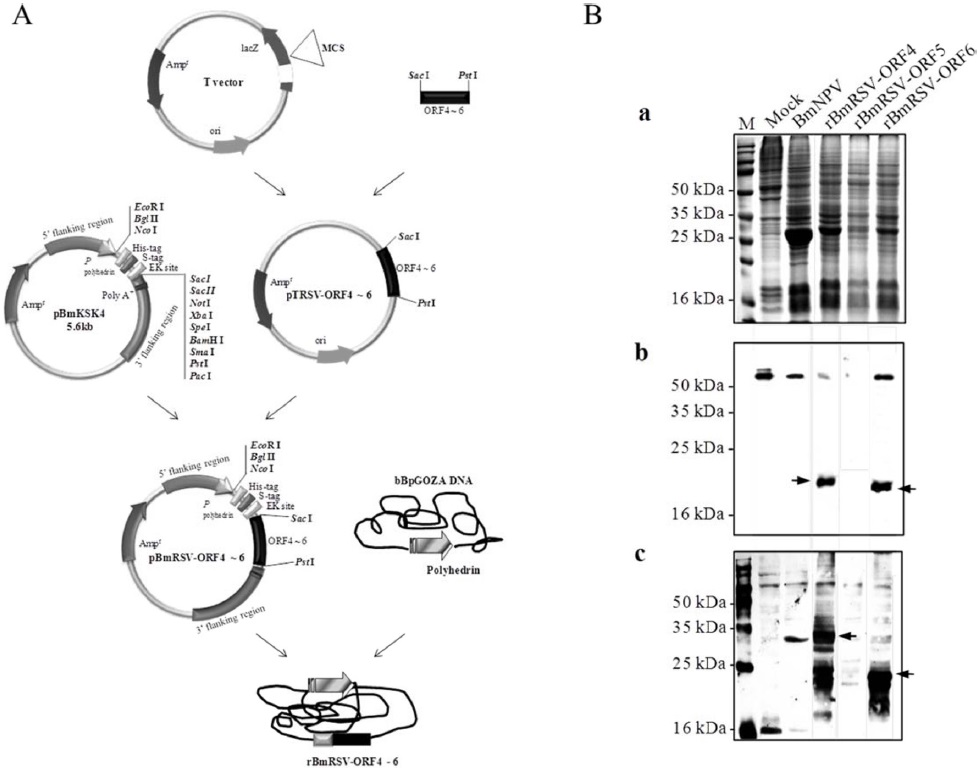
To generate the recombinant baculoviruses, Bm5 cells were co-transfected with a mixture of bBpGOZA and pBmRSV-ORFs 4 to 6 as described in Materials and methods. The recombinant viruses were called rBmRSV-ORF4, rBmRSV-ORF5, and rBmRSV-ORF6 (Fig. 1A). Bm5 cells were infected at a MOI of 5 with each recombinant baculovirus and incubated at 27℃ until harvesting of the cultures. Expression of the recombinant protein in Bm5 cells was analyzed by SDS-PAGE and Western blotting. A negative control consisting of Bm5 cells infected with wild-type BmNPV did not show any reactivity, as expected. The detection of GP4 (ORF4) and M (ORF6) protein band was difficult on SDS-PAGE analysis, but was sufficient to produce detectable signals on Western blot analysis with 6xHis tag or porcine anti-PRRSV antibodies (Fig. 1B). For GP4, several bands were observed that would correspond to non-glycosylated and glycosylated and different glycosylated forms, respectively. A possible explanation for the various patterns of PRRSV proteins expression might be due to incomplete glycosylation in insect cells. However, we could not detect any GP5 (ORF5) signal, even under Western blot analysis. This result could be explained by the high hydrophobicity of this protein, which contains several transmembrane domains (Indik
>
Enhanced expression of the PRRSV structural proteins by SUMO fusion in Bm5 cells
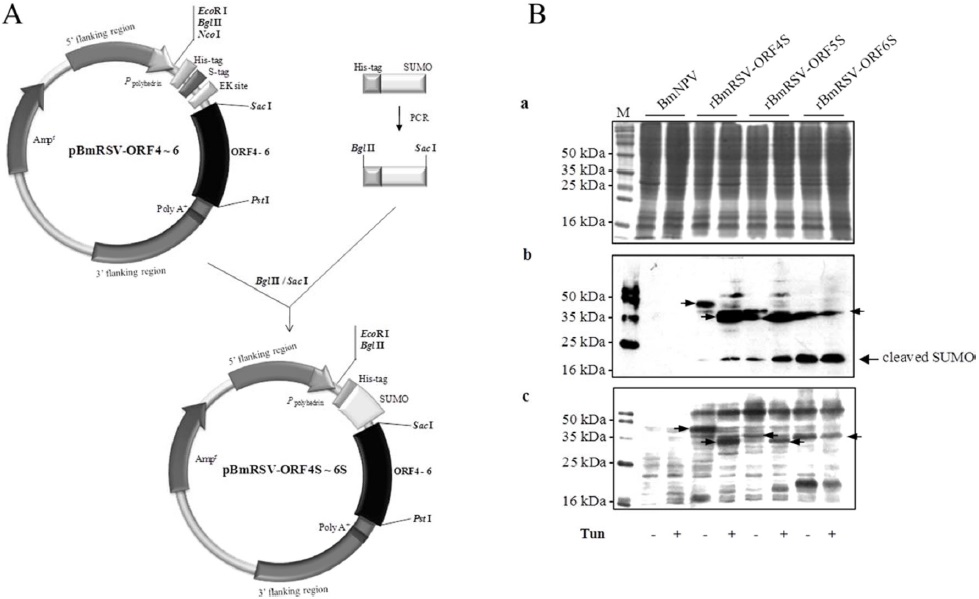
In order to enhance the protein expression, we constructed a plasmid for the expression of ORF4, ORF5, and ORF6 fused to SUMO, respectively (Fig. 2A). There was a sequence of 6xHis tag for detection and purification fused upstream of the SUMO-ORF4, -ORF5, and -ORF6. The new viruses were called rBmRSV-ORF4S, rBmRSV-ORF5S, and rBmRSVORF6S. As shown in Fig. 2B (a), the detection of three major protein bands was also difficult on SDS-PAGE analysis; however, ORF5 enhanced the level of expression dramatically, as compared to the un-fused control (Fig. 2B and 1B). Previous reports demonstrated that SUMO fusions may increase the expression of recombinant proteins and enhance the solubility of partially insoluble proteins (Malakhov
>
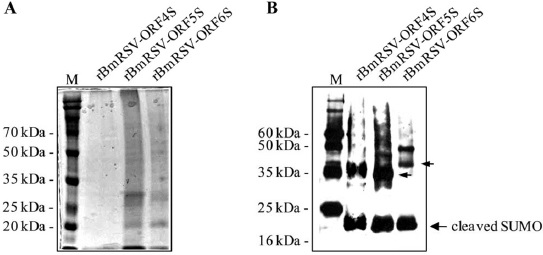
Expression and purification of the PRRSV structural proteins in B. mori larvae
Insect cells infected with recombinant baculovirus have been used for the high-level expression of various eukaryotic gene products (Hummel
>
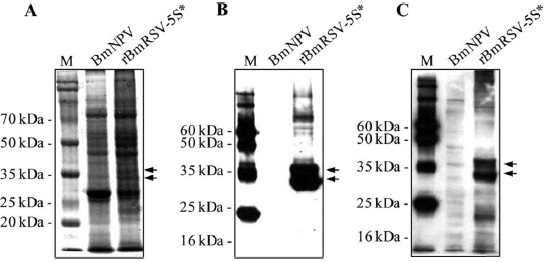
Expression of GP5 by SUMOstar fusion in B. mori larvae
A fusion tag that is not only a tool for detection and purification, but also enhances expression and/or solubility would greatly facilitate both structure/function studies and therapeutic protein production. In eukaryotic expression systems, however, the SUMO tag could be cleaved by endogenous desumoylase. In order to prevent SUMO cleavage and to obtain one band of target protein, we adapted an alternative SUMO-derived tag, designated SUMOstar, which is not processed by native SUMO proteases (Liu
In recent years, silkworm has become an ideal multicellular eukaryotic model system for basic research and practical purpose. Silkworm, as a “biofactory” for producing targeted recombinant protein, has advantages such as low feeding cost, high production, easy purification, and high safety for biohazard. The SUMO or SUMOstar fusion system using silkworm would be very attractive for future expression of target proteins.