At the turn of the 20th century, a remarkable development, which cannot be compared with anything in the previous century, has emerged in the field of biology, partly because of the completion of the Human Genome Project which identified three billion chromosomes of human deoxyribonucleic acid (DNA). Expectations were that humans might be able to learn the secrets and to understand the mechanisms of life and death as a result of the knowledge gained from the project. A sequence or a function of a gene, however, could not explain everything about human beings; therefore, interest in the interactions of genes and proteins, the physiological functions, and the complex networks in living systems has increased [1].
Recently, the number of attempts to interpret diverse information on complex biological phenomena at the systems level has increased because of the development of high-throughput technologies and data analysis methods for processing large amounts of data [2]. Omics technologies such as genomics, proteomics, metabolomics and transcriptomics have made it possible to obtain large amounts of information and to understand biological phenomena as a whole. Computational studies such as bioinformatics, data mining, machine learning, and so on have enabled us to understand and predict interactions and patterns of biological systems [1].
These recent developments in systematic understanding biological phenomena (human physiological functions) can be incorporated into systems biology. The emergence of systems biology is partly because of the availability of huge amounts of quantitative data because of advancements in high-throughput technologies and partly because of the development of computational methodologies. Systems biology is a quite new term in the field of life sciences, and it focuses on not the living thing itself but the emerging properties of the whole network by looking at the interactions of the essential factors of organisms [3]. Systems biology, therefore, has integrative and multi-scale characteristics.
Traditional medicine has mainly been used by Asian countries such as Korea, China, and Japan for thousands of years to maintain the health of their populations. The use of traditional medicine in the world has markedly increased over the last decades. Traditional medicine is based on clinical experiences acquired over a long time and is guided by East Asian traditional philosophy such as Yin-yang and five phases which underline the balance of the functional system. Traditional medicine also adheres to a personalized and holistic approach to describe health and disease [4].
The viewpoint of systems biology is consistent with the holistic perspective of traditional medicine. The concepts of systems biology and some aspects of traditional medicine are quite similar. For example, whilst systems biology is associated with the integration of the part into the whole, holistic approaches of traditional medicine consider the body as a whole [5]. Therefore, interest in a systems biology approach investigating traditional medicine is greatly increasing, with the main focus being on drug discovery and development. In other words, systems biology is becoming a cutting edge research field in current drug discovery and development.
In this study, therefore, recent advances in systems biology and its applicability to traditional medicine research are investigated. The systems biology approach is discussed from three points of view: data and databases, network analysis and inference, and modeling and systems prediction. Also, the application of systems-biology methodologies to explain the mechanism of herbal medicine is investigated.
These days, the ability to differentiate what information is valuable is becoming more important because of the flood of data and documents. In particular, as the development of high-throughput technologies has made possible the production of large amounts of data in the field of life sciences, the interpretation of those data has become more important [6, 7]. Accordingly, the demand for new and efficient data mining methods has greatly increased. Data mining aims at extracting structured information or discovering novel knowledge from a massive amount of data. For this reason, one of the most important prerequisites for data mining and systems biology is data availability. If the amount of data is not sufficient, data research should not be undertaken.
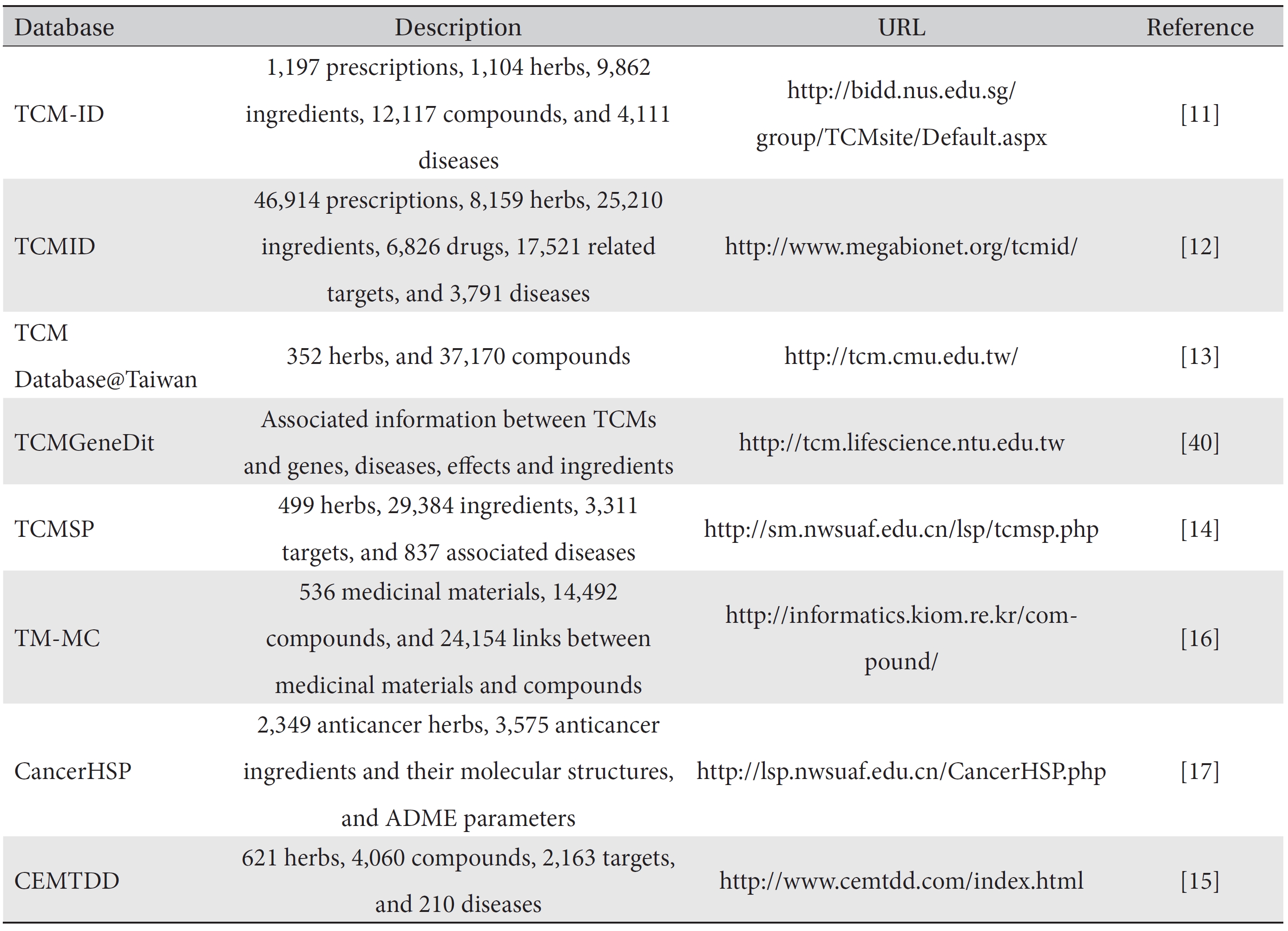
Data mining methods can be conducted by using various kinds of information obtained from sources such as bibliographic literature, experimental data, clinical data, etc. In particular, the bibliographic literature on research conducted in the field of basic Korean medical sciences should be included in data mining by using original texts as data sources. In recent research, more complex and elaborate analyses have become available due to the use of as much data as possible, which is due to the development of computational technologies. Because structured databases are essential to productive data mining, many researchers have focused on the construction of databases for traditional medicine. These efforts have resulted in the constructions of various databases such as the traditional Chinese medical literature analysis and retrieval system (TCMLARS) [8], the Korean traditional knowledge portal [9], and the knowledge of Oriental medicine web service [10], all of which are mainly related to original texts and research papers in the field of traditional medicine. Moreover, sources of traditional medicine information for proper data mining and systems biology have been introduced recently. Especially, information on herbal medicines, including herbal prescriptions, individual herbs, and compounds, could be a very good source for systems biology research because such information can provide increased knowledge for understanding the underlying mechanisms of herbal medicines and for developing drugs. A list of several traditional medicine databases for systems biology research is presented in (Table 1).
The traditional Chinese medicine-information database (TCM-ID) was developed as a convenient and integrated source for information on all aspects of traditional medicine. TCM-ID, containing 1,197 prescriptions covering 1,104 herbs, 9,862 ingredients and 4,111 diseases, provides a 3D structure of ingredients and is searchable by prescription, medicinal herb or ingredient name [11]. The traditional Chinese medicines integrated database (TCMID), the largest database for traditional medicine, comprises six data fields: 46,914 prescriptions, 8,159 herbs, 25,210 ingredients, 17,521 related targets, and information associated with modern medicines including information on drugs (6,286) and diseases (3,791) [12]. It provides a virtual display of connections between herbs and herbal ingredients and their targets by means of network-display tools. TCM Database@Taiwan has been developed to provide a free 3D small molecular structure database for traditional medicine [13]. The traditional Chinese medicine systems pharmacology (TCMSP) database and analysis platform plays a role not only as a data repository but also as a platform for the visualization and analysis of traditional medicine on a network level [14]. The Chinese ethnic minority traditional drugs database (CEMTDD), containing 621 herbs, 4,060 compounds, 2,163 targets, and 210 diseases, displays networks for relations between herbs and diseases as well as between active compounds and active targets [15]. The database of medicinal materials and chemical compounds in northeast Asian traditional medicine (TMMC), the most recently developed database, has tried to overcome the shortcomings of earlier databases, such as redundancy of herbs in a system, and contains medicinal materials from Korean, Chinese and Japanese pharmacopoeias [16]. The anticancer herbs database of systems pharmacology (CancerHSP) is a specialized database on anticancer herbs. This database, which contains anticancer activities from 492 cancer cell lines, helps to define the molecular mechanisms of anticancer activities [17].
Data availability is the first consideration of data mining and systems biology because the more data that exist; the more the information that can be obtained. The recently- developed traditional medicine databases contain several categories, such as prescriptions, herbs, compounds, targets and diseases. Databases are evolving these days. Whilst some databases provide herb-related information, other databases provide analysis tools to evaluate the relations between information [14]. Recently, specific disease- related databases have been developed, and these specialized databases offer more detailed information for understanding the associations between disease- and treatment-related factors [17].
Nevertheless, existing databases have several drawbacks. First, most databases do not state where their information came from, which will detract from the reliability of the databases. Second, some databases include a large amount of information but they do not discriminate between synonyms of the same medicinal herb. Last, language barriers exist because some databases do not provide multi-language platforms. Nevertheless, recent trends are amazing in the ways that researchers have overcome the shortcomings of databases and have generated many studies based on databases. Overall, databases need to be established to allow systems biology to be defined and evaluated.
As traditional medicine has been practiced for thousands of years because of its clinical efficacy, new treatment methods often arise from clinical situations. Moreover, traditional medicine has recently been shown to be very effective in treating various chronic diseases such as cancer, rheumatoid arthritis, leukemia, and migraine. Therefore, identifying medical knowledge from clinical data and collating quantitative biomarkers could become a significant step to bridge the gap between clinical practice and traditional medical theory [18]. Also, platforms to extract, visualize and analyze multi-dimensional data easily must be built.
3. Network analysis and Inference
The most active research field in systems biology-based traditional medicine studies is the network analysis of the relations among herbal prescriptions, individual herbs, compounds, targets, and diseases. Systems biology, by using computational methods on high-throughput experimental data, can offer a fast way to achieve the goal of identifying the networks by which components interact to carry out their regulatory functions. The major technologies here have become popular tools for understanding the network relations between key factors and for discovering drugs. The recent application of forefront technologies using medicinal herbs and formulae has provided an in depth understanding of their targets and ingredients, which has facilitated an understanding of their mechanisms and actions. However, because herbal formulae and herbs contain a large number of ingredients and their synergistic and antagonistic effects cannot be easily classified, the detailed mechanisms of medicinal herbs and formulae still remain unclear. Although complex ingredients and complex actions of herbs/herbal prescriptions are unique characteristics, they are also ambiguous points that hinder our understanding of their actions. However, recent developments of computational methodologies have allowed the analyses of huge amounts of data simultaneously, and these data mining technologies hold the limelight as a novel approach to the discovery of new drugs.
Network pharmacology, a key technology of systems biology, determines how various ingredients interact by constructing drug-target-disease networks and analyzing them. It has attracted much attention by researching the molecular mechanisms of herbal formulae and herbs with complicated targets and diseases. The rise of network pharmacology accentuates a paradigm shift from a “single- component, single-target” strategy to a “multi-component, multi-target” feature. Traditional medicine takes an initiating role in the “multi-component, multi-target” therapies because hundreds of different components in a prescription can cure a disease or even one herb shows dynamic effects because of its multiple actions. Therefore, explicating the underlying mechanism of a herbal medicine is very difficult because of its complex composition, though its efficacy is not in doubt.
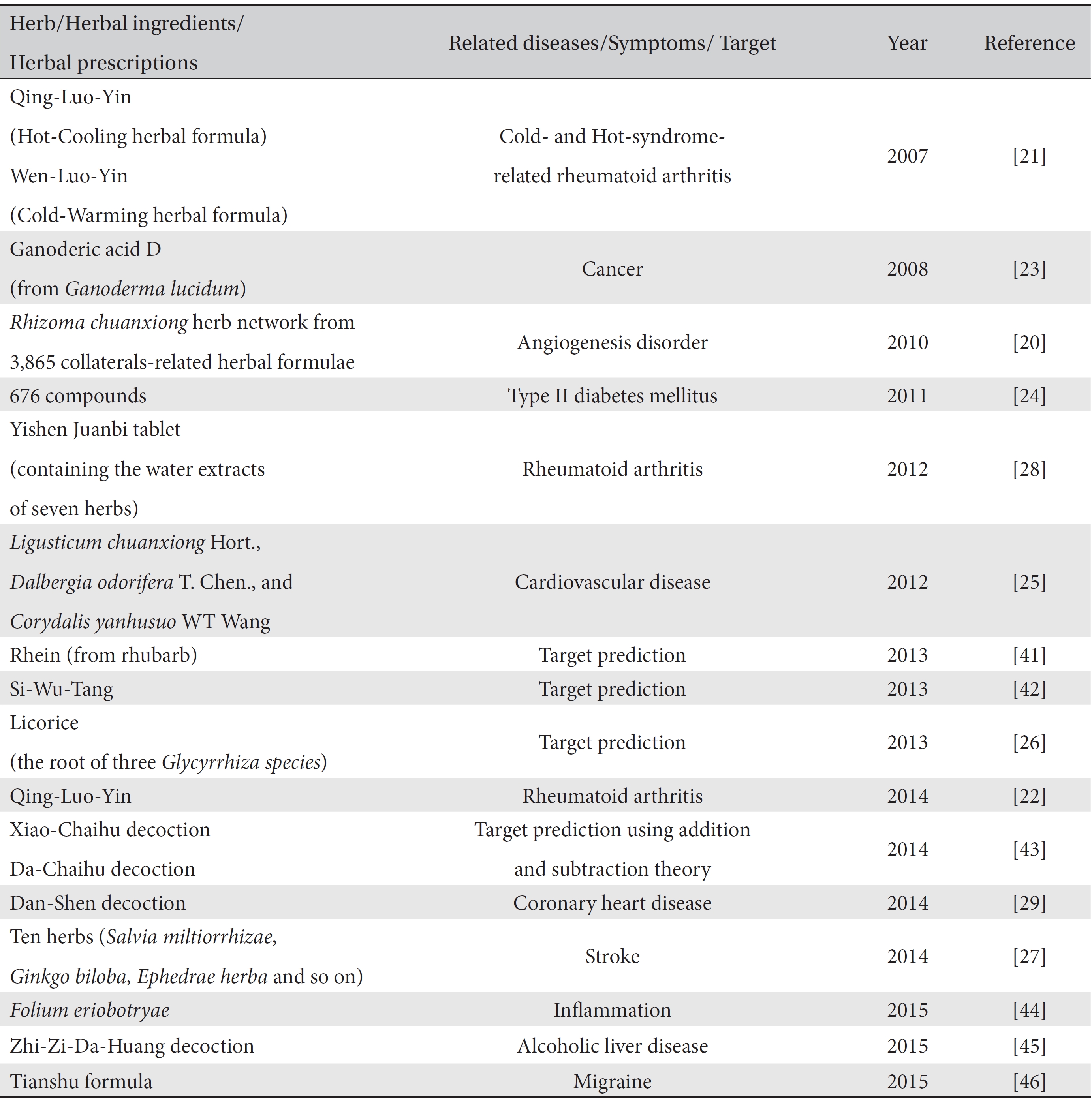
Many network-based computational studies have been conducted to find effective components and to detect the mechanisms of the actions of herbal prescriptions/herbs against many diseases/targets (Table 2). Since the mid- 2000s, interdisciplinary research between network pharmacology and herbs/herbal prescriptions has been undertaken mainly by the Li Shao group [19-22]. That group proposed “network target” theory, which is a disease-specific bio-molecular network, as a core in the “Herb network (Herbal formula) ─ Biological network (Network target) ─ Phenotype network (Disease)” [19]. The close relations among those three networks make it possible to explain the holistic approach of traditional medicine and to provide a new paradigm for drug development.
The domain of network pharmacology research embraces herbal prescriptions, individual herbs, and compounds. The related diseases are also very broad, ranging from immunological disorders such as rheumatoid arthritis to chronic diseases such as diabetes mellitus to other diseases such as cancer, alcohol-related liver diseases, and migraine headaches. Most explorations have focused on drug (herbal prescriptions, herbs, compounds, or ingredients) ─ target networks [23-26], but some have focused on the herb network in herbal prescriptions [20] and the target signaling pathway network [27]. Some studies have spotlighted pattern identification following a disease and have developed a disease-pattern-target network to understand the mechanisms of herbal medicines in treating diseases [21, 28, 29]. The information gained from the comparisons among these networks may provide new knowledge to understand mechanisms, detect new network targets, and propose new combinations of herb/compounds. Overall, for the study of network pharmacology, traditional medicine theories, omics technologies and bioinformatics technologies, including computational mining methods and machine learning methods, are combined to interpret the mechanisms of herbs/prescriptions-target-diseases effectively.
4. Modeling and Systems prediction
The use of mathematical modelling and analysis and prediction of systems has a long history since the first mathematical cardiac model developed in the 1960s [30]. For several decades since then, various kinds of mathematical models have been developed to verify hypotheses and predict systems. A model is a set of structured assertions that state the interactions among entities of a system. The entities in a model can be various, such as the specific biological elements or the specific characteristics of a system. In addition, the interactions can be due to various processes in a system, such as molecular reactions or the movement of a molecule. Changes of entities over time and space can be inferred by using a mathematical model. In other words, computational models allow us to understand a structured system and its functional properties at the systems level [6].
The
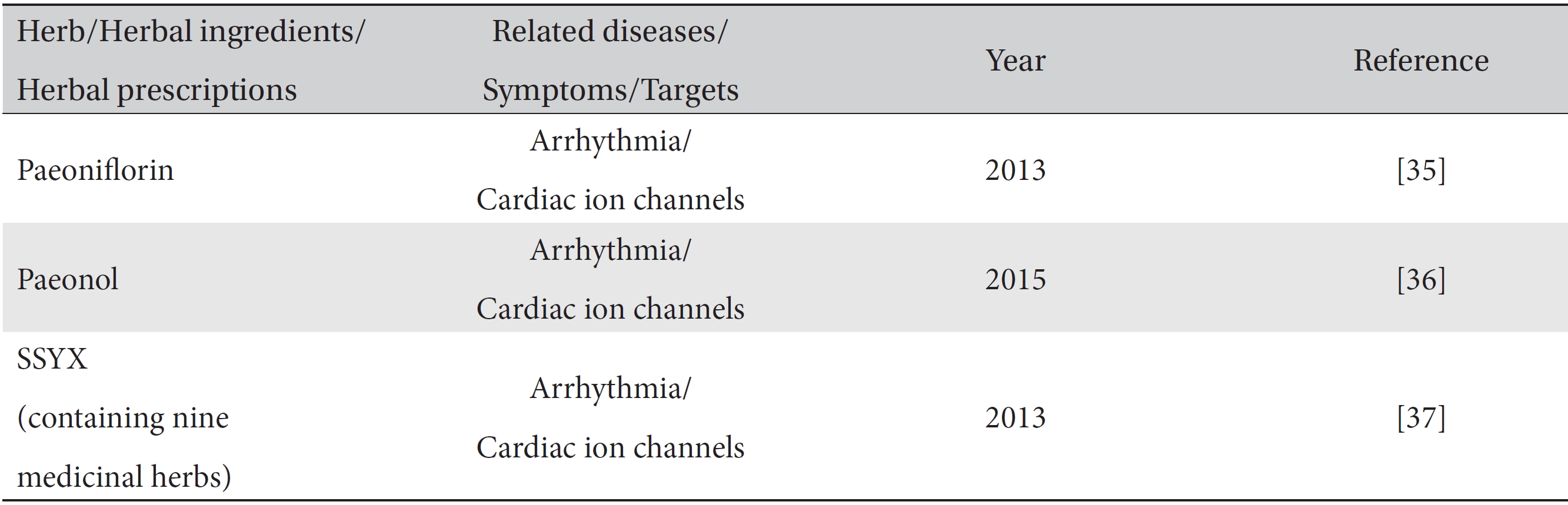
Only a few simulation studies have demonstrated mathematical modeling with herbs and herbal prescriptions (Table 3). Paeoniflorin and paeonol extracted from the paeoniaceae family were shown in
The paradigm shift from a “single-component, single-target” strategy to a “multi-component, multi-target” strategy can be applied in mathematical modelling because a simulation shows the comprehensive actions of herbs and herbal formulae at the systems level. As stated, experimental data can be translated into how it will work in a system and
However, a more widespread integration of modeling and experimental programs needs to be developed to understand the biological function of a whole system. Therefore, the virtual physiological human (VPH) project was initiated in 2006. The VPH is intended to provide a unifying framework to facilitate integration and prediction, eventually creating the human body as a single complex system
Although simulation studies using traditional medicine are just an opening stage, their applicability is growing. For future studies, developing a systematic computational model to predict the complex effects of herbal medicines via computational disease models based on traditional medicine would be a challenge. Such an approach would help us to understand the complicated actions of herbal medicines at the systems level.
Systems biology has been applied in research on traditional medicine, especially research on herbal medicines. The efficacy of a herbal medicine is considered to be due to the combined effects of its various ingredients, the so called multi-component effect. Therefore, a systems biology approach based on a holistic viewpoint may provide a way to understand the mechanisms of herbal medicines, which is the reason the systems biology approach is highlighted as a meaningful drug development method [31].
The systems biology approach to traditional medicine has special features that are different from those of previous research approaches originating from reductionist’s aspects [47]. However, not many meaningful outcomes are connected to or reflected in clinical situations partly because the linear and reductive perspectives used in modern scientific research are not sufficient to reflect the holistic perspectives of traditional medicine. Accordingly, the use of a systems biology approach should allow progress to be made in understanding the mechanisms of traditional medicine, though shortcomings, such as the lack of databases and computational models, still exist. Therefore, much work still needs to be done if the holistic perspectives of traditional medicine and the multiple actions of herbal medicines are to be understood through the mathematical tools provided by systems biology.
In conclusion, interactions between systems biology and traditional medicine are currently taking place. The applicability of systems biology to traditional medicine depends on establishing the infrastructure to collaborate different disciplines, finding better network pharmacology methods and mathematical models, and integrating computational technologies into holistic perspectives. Clearly, these new mathematical research methodologies will be useful and valuable tools for closing the gap between basic and clinical sciences and for understanding the link between Western and traditional medicine.
[Table. 1] Traditional medicine databases
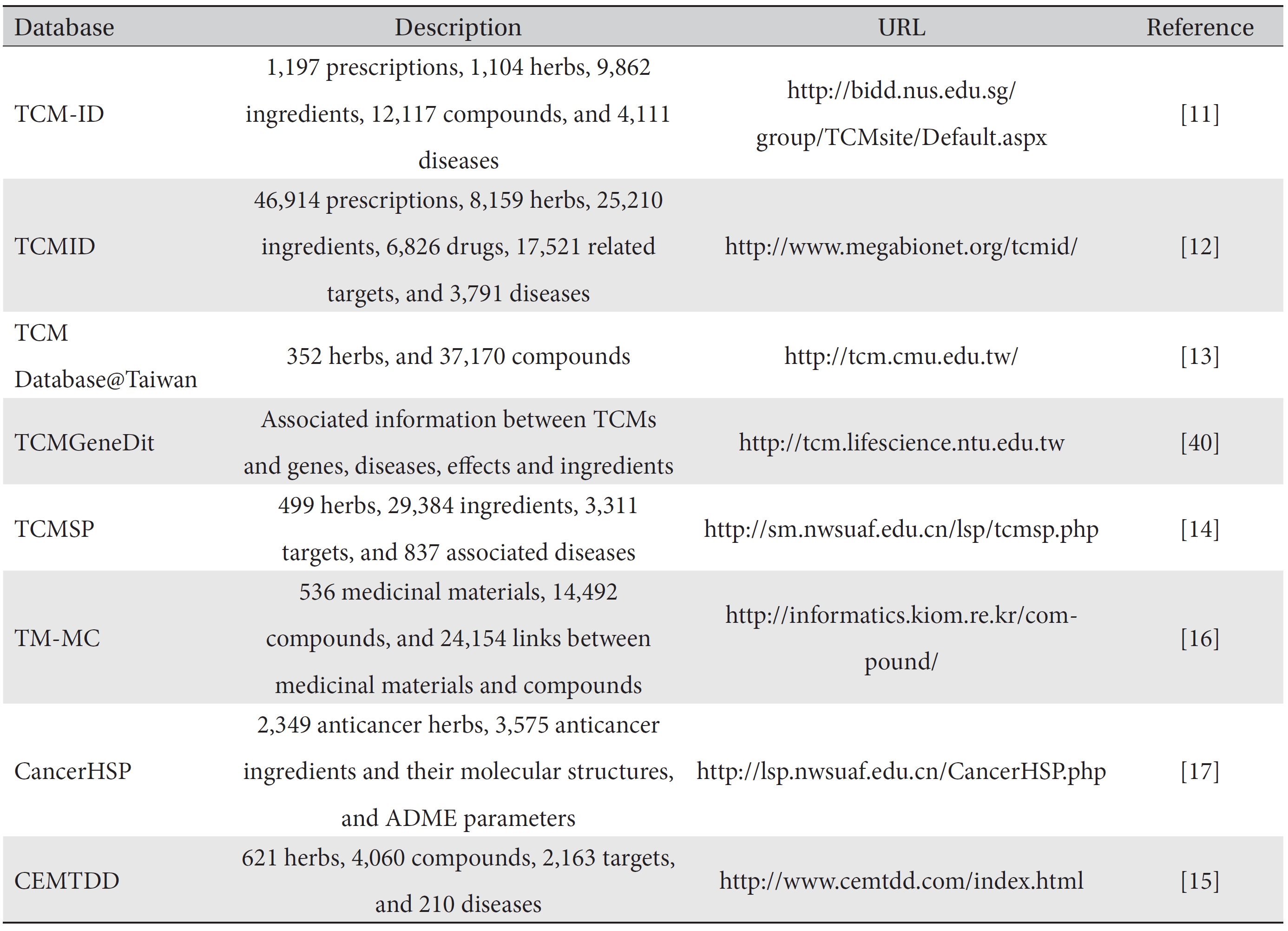

Traditional medicine databases
[Table. 2] Selected network pharmacology studies of traditional medicine
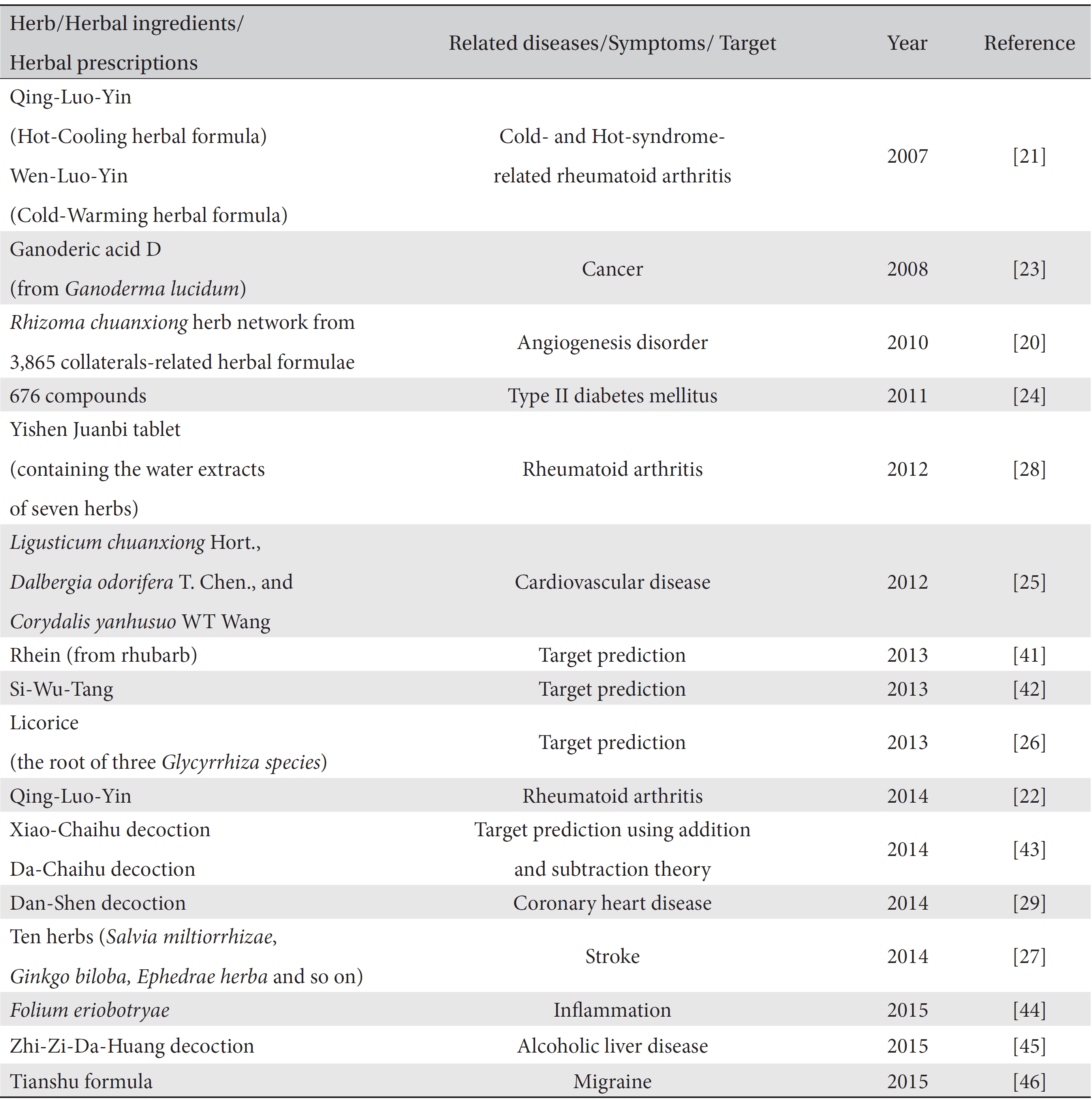
Selected network pharmacology studies of traditional medicine
[Table. 3] Modeling prediction studies of traditional medicine
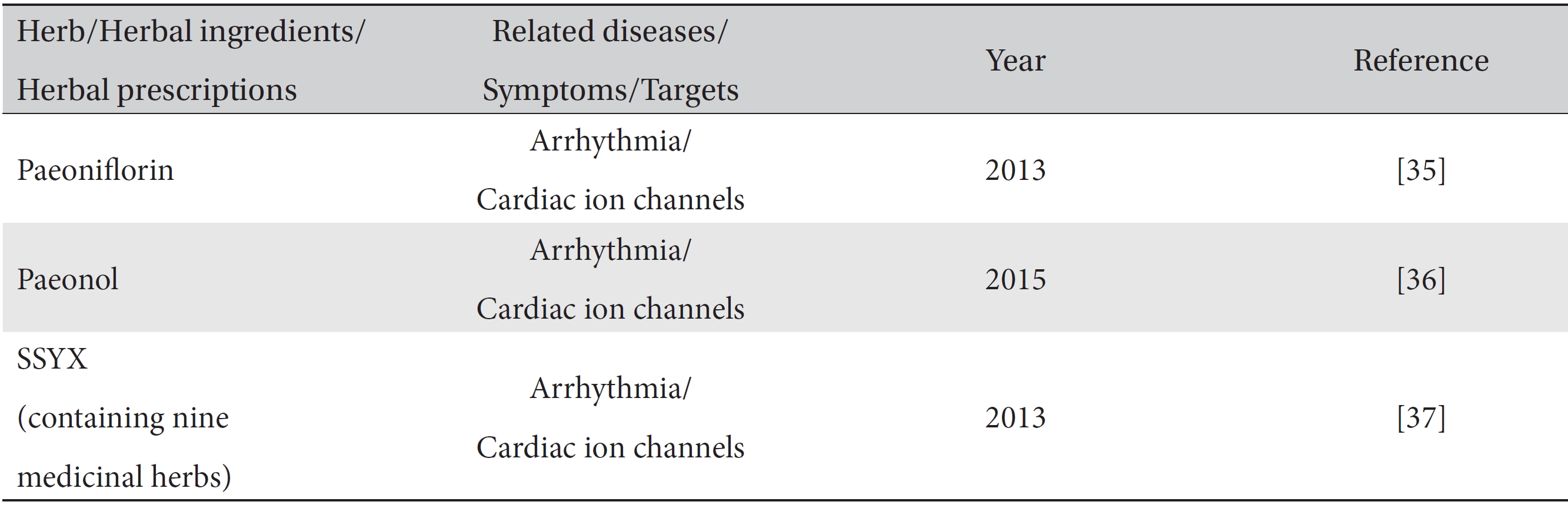
Modeling prediction studies of traditional medicine