The etiological bacterium
The aim of this study was to obtain fundamental data for identification of
Strains of
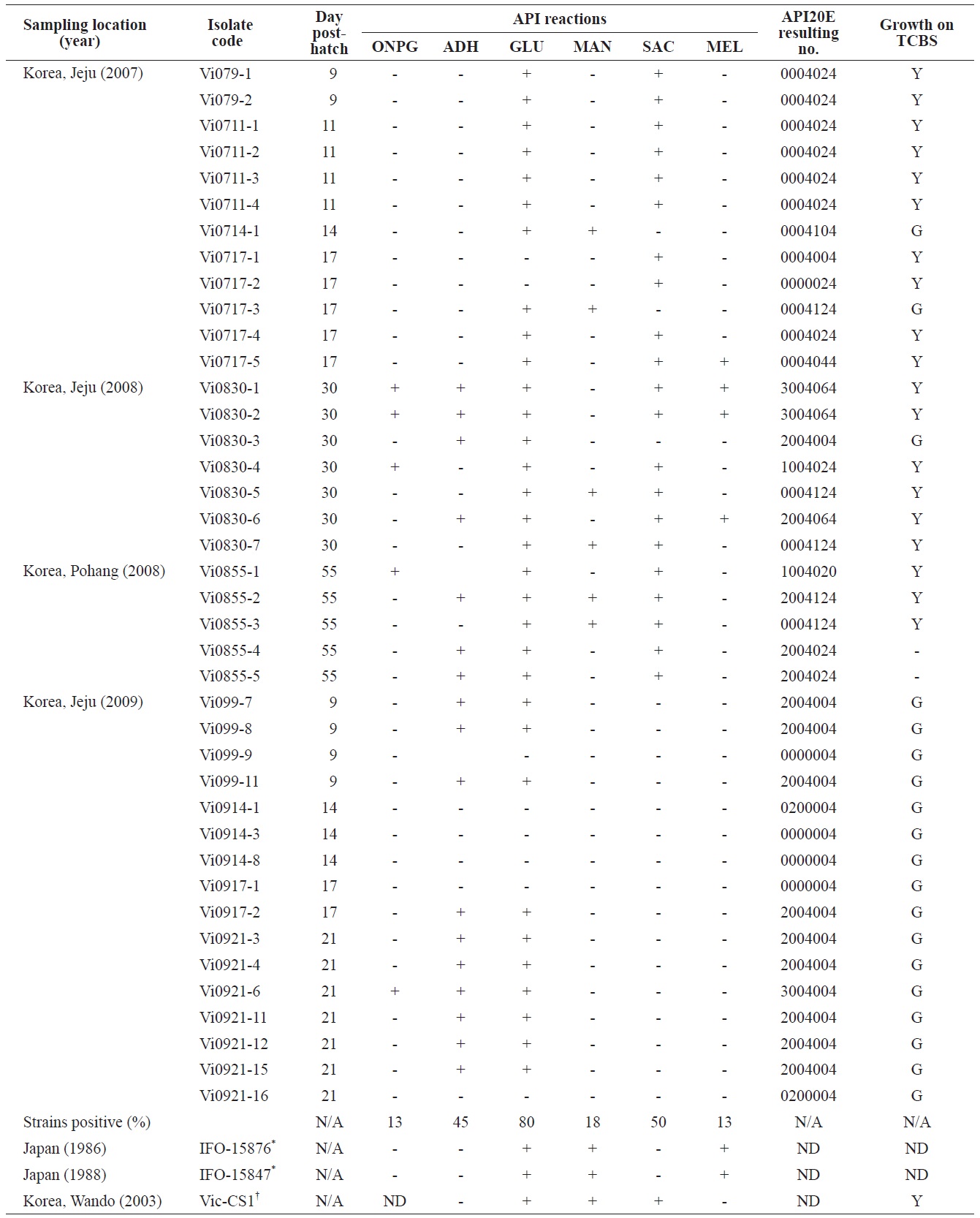
Phenotypic profiles of the Vibrio ichthyoenteri isolated from diseased Olive flounder as determined by the API 20E system
tracts of 9-30 days post-hatching (dph) Olive flounder larvae (approximately 30 at each sampling point) with symptoms of bacterial enteritis in the Jeju hatchery from 2007 to 2009. To acquire more isolates of
Fish were initially disinfected with 0.1% (w/v) benzalkonium chloride before being rinsed with sterile distilled water. The intestine was then aseptically removed and homogenized in 10 mL of sterile 0.85% (w/v) saline. Volumes (0.1 mL) of 10-fold dilutions of the homogenate were individually spread onto tryptic soy agar (Difco, Sparks, MD, USA) containing 1% (w/v) sodium chloride (TNA). The plates were incubated at 25℃ for 48 h, after which representative colonies, normally in pure culture, were selected and streaked for isolation. Forty isolates of
>
Biochemical characterization
All isolates were tested for biochemical characteristics using the API20E system (bioMerieux, Marcy I’Etoile, France). Enzymatic activities were detected using APIZYM commercial kits (bioMerieux) according to the manufacturer’s protocol. API20E and APIZYM tests were conducted three times for each isolate. Growth was observed on TNA at 35℃ after 48 h and on thiosulfate citrate bile salt sucrose (TCBS) 25℃ for 48 h.
Bacterial genomic DNA was isolated using a Genomic DNA extraction kit (Bioneer, Daejeon, Korea) and following the manufacturer’s protocol. The DNA templates were amplified using PCR in a Gene Amp PCR system 2400 (Perkin Elmer, Wellesley, MA, USA) using universal primers amplifying a 1,343-bp region of the 16S rRNA gene (63F: 5-CAGGCCTAACACATGCAAGTC-3′; 1406R: 5′-ACGGGCGGTGTGTRC-3′) obtained from Bioneer. The DNA templates were amplified by initial denaturation at 94℃ for 5 min, followed by 35 cycles of denaturation at 94℃ for 30 s, annealing at 55℃ for 30 s, extension at 72℃ for 1.5 min, and a final extension at 72℃ for 5 min. Direct sequencing of the amplified DNA fragment was performed using a 3730xl DNA automatic sequencer (Applied Biosystems, Foster City, CA, USA). All sequences were submitted for similarity searches with BLAST (Altschul et al., 1990).
A total of 40 representative isolates of
>
Biochemical characterization
All isolates were negative for lysine decarboxylase, ornithine decarboxylase, citrate utilization, H2S production, urease, tryptophan deaminase, indole production, acetoin production, gelatinase, fermentation/oxidation inositol, fermentation/ oxidation sorbitol, fermentation/oxidation rhamnose, fermentation/ oxidation amygdalin, and fermentation/oxidation arabinose (data not shown). Overall, β-galactosidase (ONPG), arginine dihydrolase, citrate utilization, and/or acid production in four different carbohydrates, including glucose, mannitol, sucrose, and/or melibiose were observed in various isolates (Table 1). No strain grew at 35℃. Almost half of the isolates (19 of 40) produced yellow colonies on TCBS (Table 1).
Intensities of enzymatic activities were established using the APIZYM system. Strong alkaline phosphatase, leucine arylamidase, and N-acetyl-β-glucosaminidase activities were induced by
Presently, all isolates derived from the intestine of diseased olive flounder larvae were identified as
Biochemical characterization is still used for identification of bacteria such as
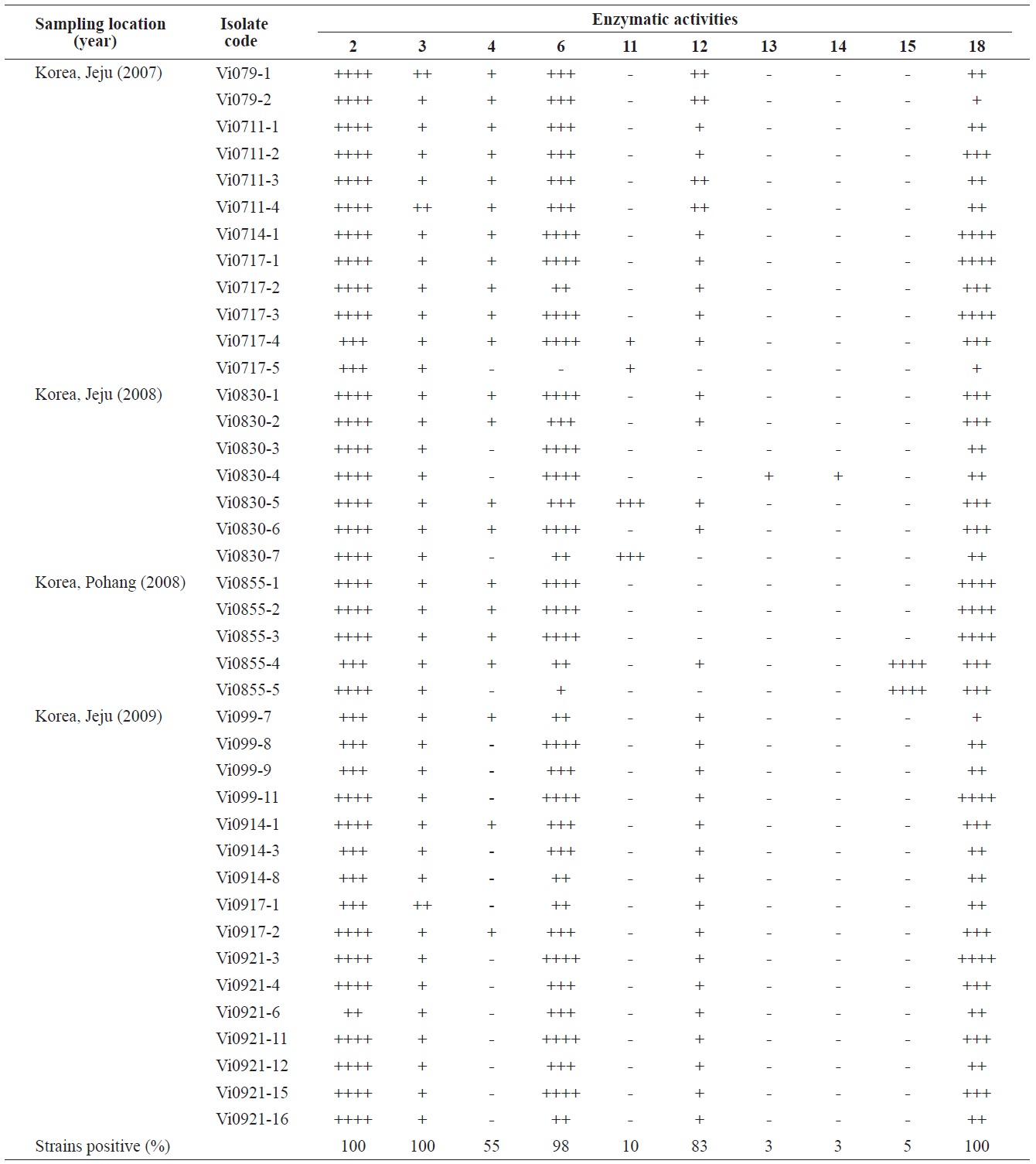
Enzymatic profiles of the Vibrio ichthyoenteri isolated from diseased Olive flounder as determined by the API ZYM system
time exhibited diverse biochemical and enzymatic phenotypes. Some isolates in this study showed arginine dihydrolase activity, ONPG hydrolysis, and acid production from melibiose and utilization of citrate, all of which were not detected in the six strains of
In conclusion, our data suggest that most isolates of the same identity as determined by the 16S rRNA gene sequence exhibited diverse enzymatic and biochemical phenotypes. These results suggest that pathogenic