The hemotoxic venoms of rattlesnakes and vipers usually affect the circulation system1). Some of these toxins, both enzymattic and non-enzymatic, lead to the spectacular changes in haemostasis and to the frequent hemorrhage seen after snake bite. The enzyme proteins are acetylcholinesterases, ADPases, phospholipases, hialuronidases, and haemostatic proteases. The non-enzymic proteins are disintegrin and bradykinin-potentiating peptides. The activities of these toxins include coagulation and anticoagulation of blood, platelet-activation, anti-platelet function, fibrinolytic activation, and hemorrhage2- 6).
Fibrinolytic and fibrinogenolytic activities have been described in venom from the following snake families:
The fibrin(ogen)olytic proteases have drawn particular attentions as potential therapeutic agents to remove plasma fibrinogen from the circulation for treatment of acute isochaemic stroke and to dissolve blood clot for treatment of occlusive thrombi7-9). Fibrolase from copperhead snake venom degrades both α and β chains of fibrin and showed some promise as a thrombolytic agent. Alfimeprase, a recombinant fibrinolytic enzyme derived from fibrolase has been being developed as a clinical agent.
Terada and his colleagues12-14) purified and characterized several metalloproteases from Chinese mamushi (
The molecular weight of halystase, a serine protease, purified from Japanese common viper (
The objectives of this study were to identify fibinolytic proteases from Yangtze mamushi (
The snake venom from
2. Isolation of fibrinolytic enzyme
The venom powder (2 g) was dissolved and dialyzed in 50 mM Tris-HCl, pH 7.6, which was then centrifuged at 10,000 g for 30 min using the refrigerated centrifuge of model VS-15CF (Vision Scientific, Korea) to remove insoluble substances. The venom solution was applied to a column (2.5 cm × 8 cm) of Q-Sepharose (GE, USA) equilibrated with 50 mM Tris-HCl, pH 7.6. The column was eluted with 200 ml of 50 mM Tris-HCl, pH 7.6 and then with a concentration gradient from 150ml of 50 mM Tris-HCl, pH 7.6 to 150ml of 50 mM Tris-HCl, pH 7.6, 0.3 M NaCl. The column was subsequently eluted with 50ml of 50 mM Tris-HCl, pH 7.6, 0.3 M NaCl. The flow rate was 21 ml per hour and the fraction volume was 7 ml. The fractions with fibrinolytic activity were combined and concentrated by using polyethylene glycol and then applied to a column (2.5 cm × 109 cm) of Sephadex G-75 (GE, USA). The column was eluted with 50 mM Tris-HCl, pH 7.6, 0.15 M NaCl. The flow rate was 14 ml per hour and the fraction volume was 7 ml. All the isolation steps were performed at refrigeration temperature. Absorbance at 280 nm and fibrinolytic activity in the fibrin plate assay of the fractions from the chromatography were determined. The fractions with fibrinolytic activity were subjected to SDS-PAGE analysis.
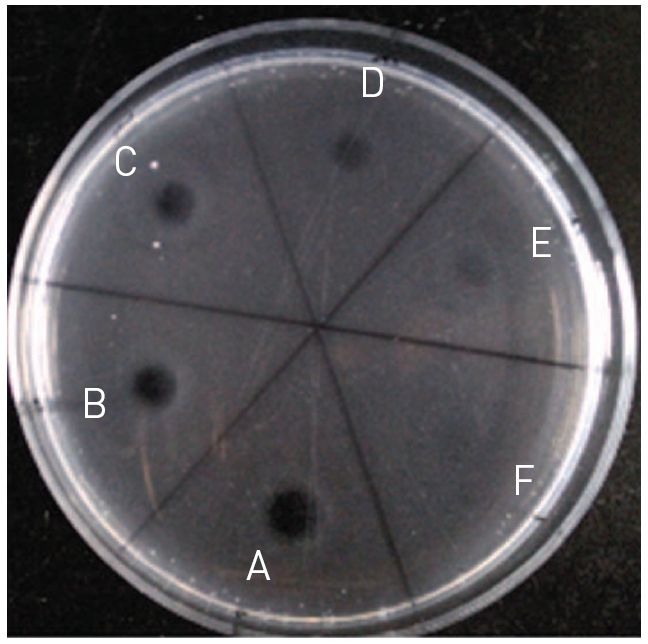
Fibrinolytic activity was determined on fibrin plates according to the following procedure17). An aliquot (200 μl) of thrombin solution (10 U/ml) in 100 mM Tris-HCl, pH 7.8 was added to 9 ml of 0.1% fibrinogen in the same buffer. The mixed solution was added into petri dish with the diameter of 9 cm and incubated at 37?C for 1 hr, until it was converted into fibrin gel. An aliquot (10 μl) of the venom fraction was placed on the fibrin gel and then incubated at 37?C for 18 hr. Two diameters (r1 and r2) at a right angle of hydrolysis zone were measured. The fibrinolysis area was calculated following the formula of 0.785r1 × r2. Relative fibrinolysis (%) was percentage of the fibrinolysis area of treated fibrinolytic protease to that of control without treatment.
Sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) was performed according to the procedure described by Laemmli18). The molecular weight markers (Bio-Rad, USA) for SDS-PAGE consisted of myosin (200 kDa),
An aliquote (200 μl) of 2% proteins in 50 mM Tris-HCl, pH 7.5, 0.15 M NaCl, 10mM CaCl2 was added with 50 μl of the fibrinolytic protease (2 μg/ml) in the same buffer and then the mixture was incubated at 37?C for 6 hour. An aliquote (10 μl) of the mixture was taken at specified times during the incubation and added with equal amount of the sample buffers for SDS-PAGE each time. The mixtures were heated at 100?C for 5 min and analyzed in SDS-PAGE.
Protein concentration was determined using BCA Protein Assay Reagent (Pierce, USA). Bovine serum albumin was used for calibration curve.
1. Isolation of fibrinolytic enzymes from the snake venom
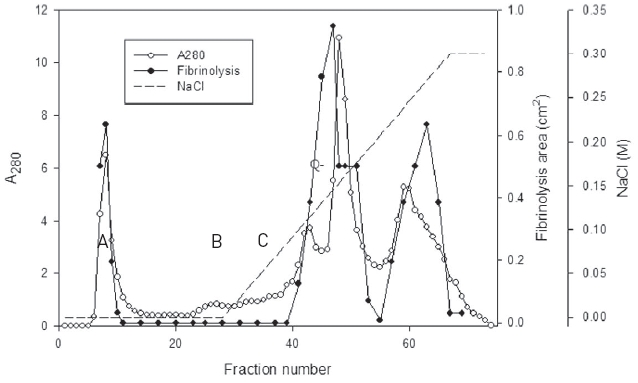
The snake venom was separated into three major peaks of A, B, and C by using chromatography on the column of Q-Sepharose which was eluted using 50 mM Tris-HCl, pH 7.6 and then the same buffer with the concentration gradient up to 0.3 M sodium chloride (Fig. 1).
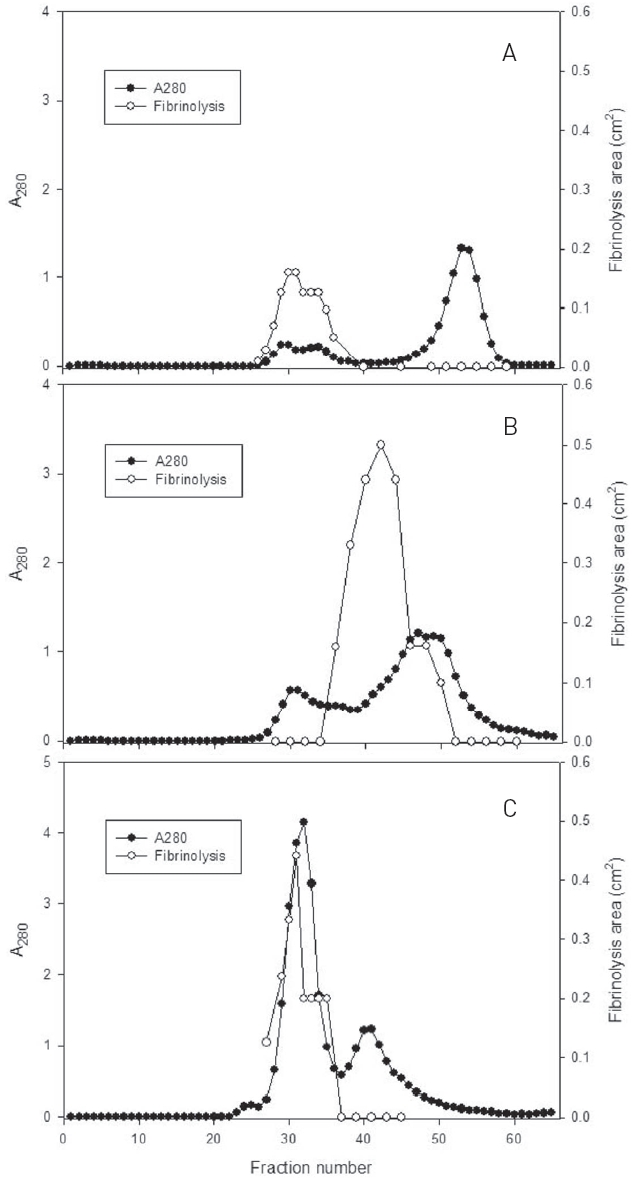
Fibrinolytic activities of the fractions of the three peaks were closely related with protein concentration. The fractions of three major peaks showing fibrinolytic activity were combined, concentrated, and subjected to chromatography using a column of Sephadex G-75 (Fig. 2).
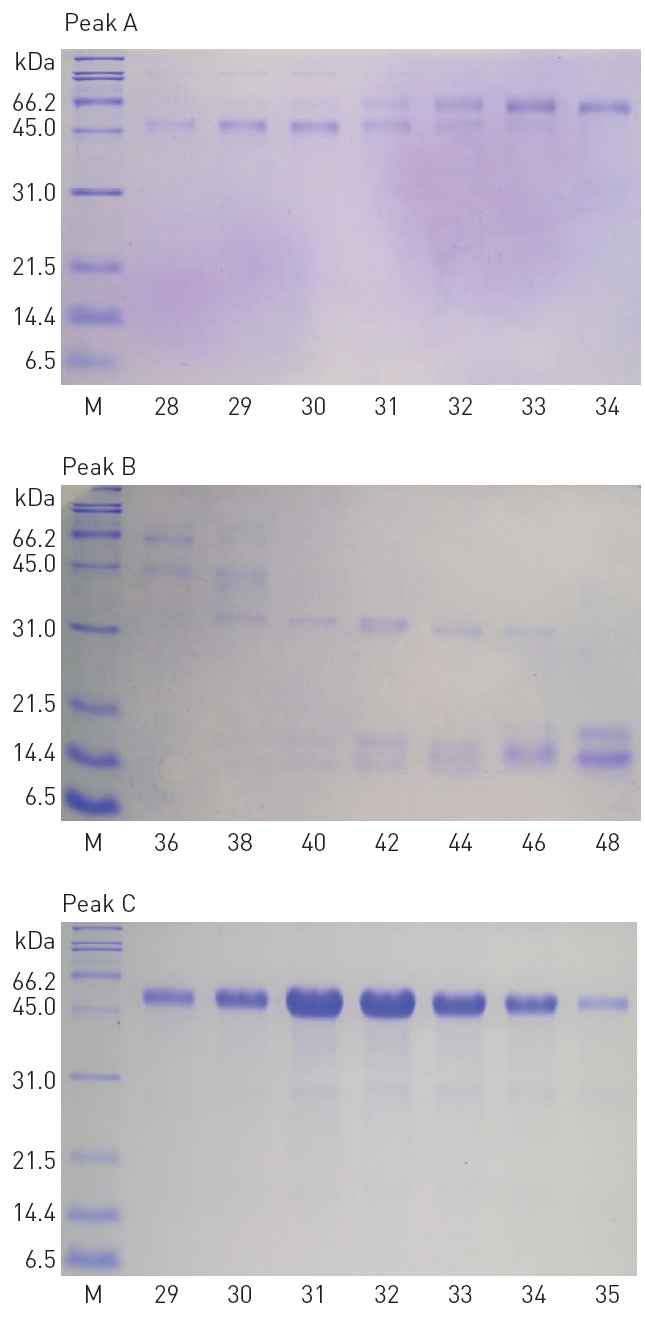
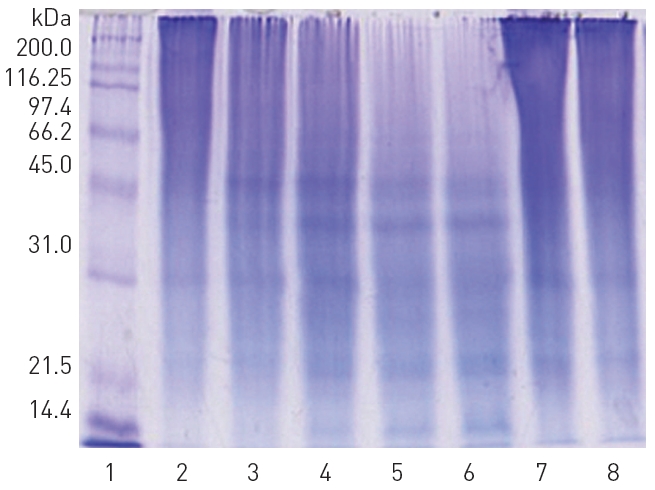
The fractions 28-34 from the peak A, 36-48 from the peak B, and 29-35 from the peak C, showed significant fibrinolytic activities. These fibrinolytic fractions were subjected to SDS-PAGE analysis (Fig. 3).
2. SDS-PAGE analysis of the fibrinolytic fractions
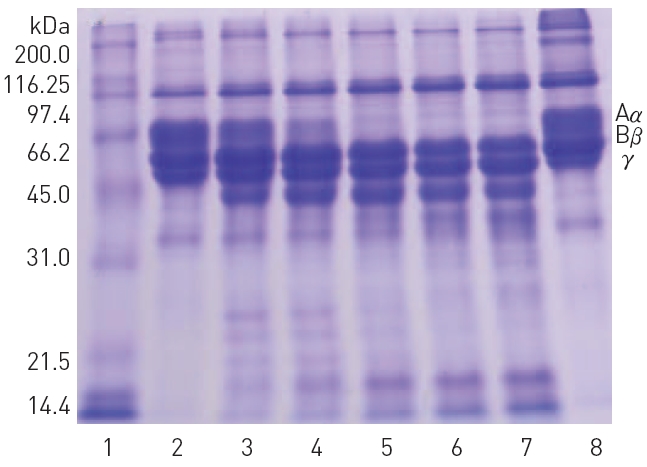
The polypeptide patterns of the fibrinolytic fractions which showed strong fibrinolytic activity are shown in (Fig. 3). The fractions 28-30 and 33-34 contained two polypeptides of 46 and 59 kDa, respectively. The fraction 42 showing the strongest fibrinolytic activity in the peak B contained three polypeptides of 32, 18, and 15 kDa. All the frac-
tions 29-35 from the peak C contained a major polypeptide of 54 kDa. These results suggested that these polypeptides were potential fibrinolytic proteases and the fibrinolytic protease of 54 kDa was a predominant fibrinolytic protease in the venom.
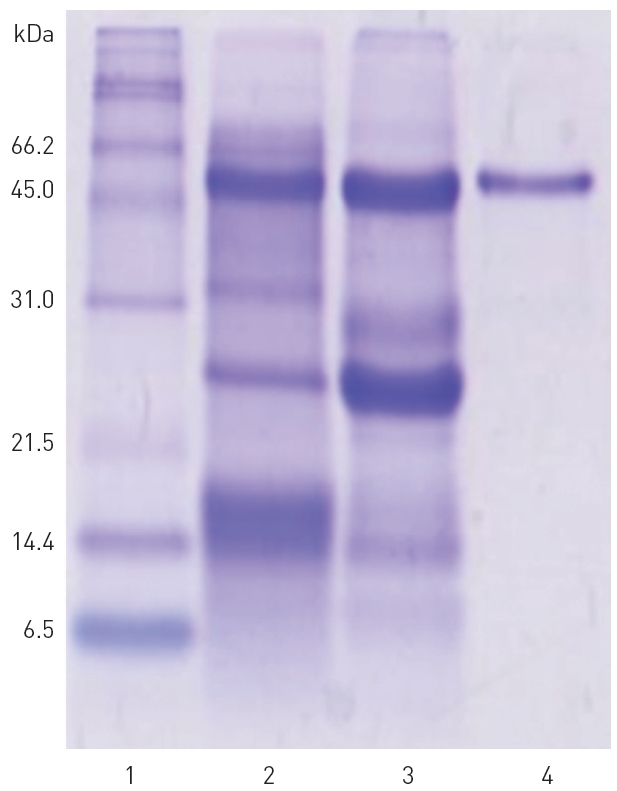
The fibrinolytic fractions 29-35 from the peak C obtained from the Sephadex G-75 column chromatography (Fig. 2) were combined together, concentrated in dialysis tubing using polyethylene glycol, and then subjected to chromatography on the same column of Sephadex G-75 again to purify the
major fibrinolytic protease (results not shown). SDS-PAGE in Fig. 4 confirmed that the purified fibrinolytic protease of 54 kDa (lane 4) was one of the major polypeptides in the snake venom (lane 2) as well as in the combined fibrinolytic fraction of
peak C (lane 3) obtained after Q-Sepharose column chromatography. The amount of the purified fibrinolytic protease obtained from 2 g of the snake venom was 81 mg.
3. Characterization of the fibrinolytic protease
The fibrinolytic protease was diluted to the protein concentrations of 0.5, 0.4, 0.3, 0.2, 0.1, and 0.05 mg/ml and 10 μl of the diluted fibrinolytic protease was used in fibrin plate assay as shown in (Fig. 5). The fibrinolysis zone sizes which indicated fibrinolytic activity were very weak at the protein concentration of 0.05 and 0.1 mg/ml. As the protein concentration increased from 0.2 to 0.5 mg/ml, the size of the fibrinolysis zone gradually increased. The protein concentration of 0.45 mg/ml was used
in the following study to determine the effects of protease inhibitors and salts on the fibrinolytic activity.
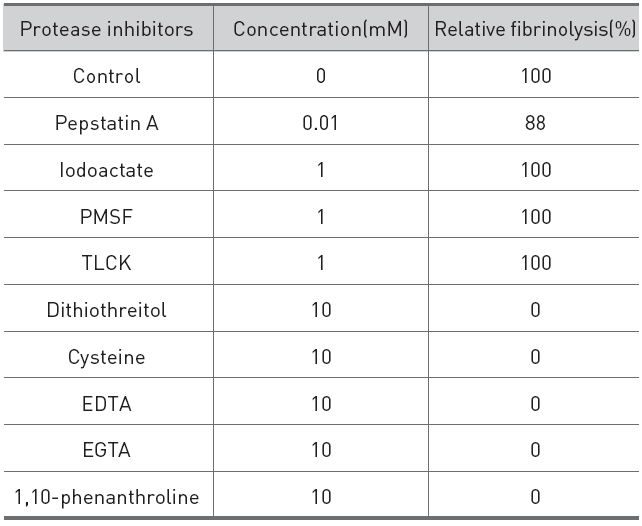
The effects of protease inhibitors on the fibrinolytic activity were determined as shown in (Table 1).
The reducing compounds, such as dithiothreitol and cysteine, which reduce disulfide linkage to sulfhydryl group of proteins completely destroyed the fibrinolytic activity, which suggested that the fibrinolytic protease contained disulfide linkage which is necessary for the activity. Metal chelators, such as EDTA, EGTA, and 1,10-phenanthroline, also destroyed the fibrinolytic activity, which indicated that the fibrinolytic protease was a metalloprotease which required metal ion for the activity. Serine protease inhibitors, PMSF and TLCK, a cysteine protease inhibitor, iodoacetate, and an acid protease inhibitor, pepstatin A showed no inhibition on the activity. These results indicated that the fibrinolytic protease was a metalloprotease which contained disulfide linkage.
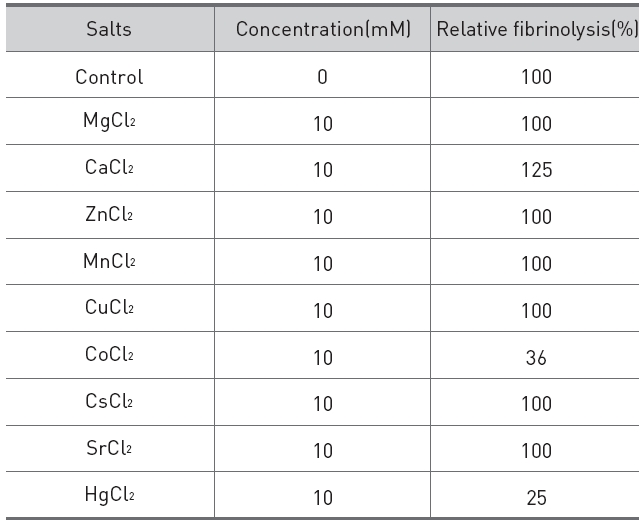
The effects of salts on the fibrinolytic activity of the protease were shown in (Table 2).
Addition of calcium chloride to the fibrinolytic protease increased the fibrinolytic activity and
[Table 1.] Effects of protease inhibitors on the fibrinolytic protease from G. b. siniticus.
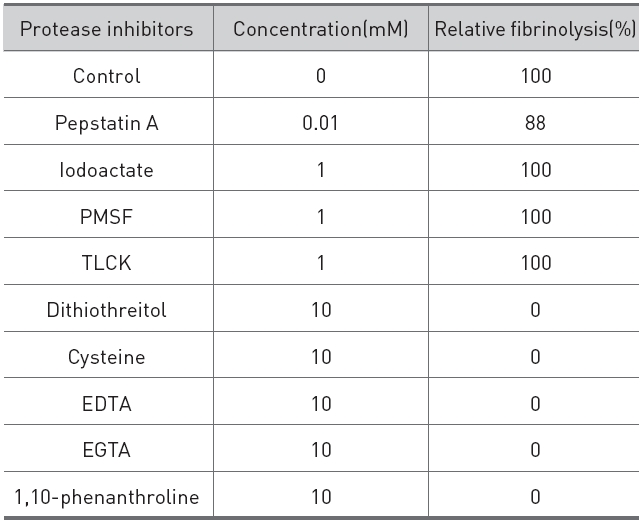
Effects of protease inhibitors on the fibrinolytic protease from G. b. siniticus.
[Table 2.] Effects of salts on the fibrinolytic protease from G. b. siniticus.
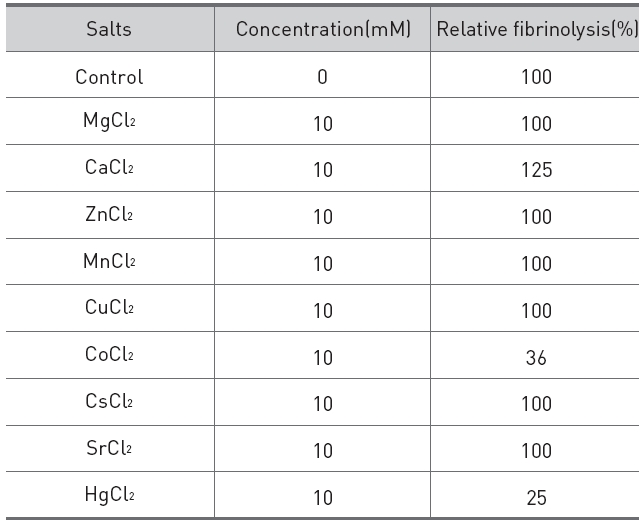
Effects of salts on the fibrinolytic protease from G. b. siniticus.
made the fibrinolysis zone clearer. Addition of cobalt(II) chloride and mercuric chloride partially inhibited the fibrinolytic activity.
4. Hydrolysis of proteins by the fibrinolytic protease
The fibrinolytic protease cleaved preferentially A
When 10 mM calcium choride was omitted from the reaction mixture, hydrolysis of Aα-chain was retarded and Bβ-chain and γ- chain were unaffected (data not shown). The fibrinolytic protease hydrolyzed high molecular weight polypeptides of gelatin into low molecular weight peptides. However, plasmin did not hydrolyze gelatin (Fig. 7). The fibrinolytic protease did not hydrolyze bovine serum albumin (results not shown). Since gelatin is heat-denatured collagen, the fibrinolytic protease may also hydrolyze collagen which is a major protein in extracellular basal membrane and connective tissue.
These results showed that the fibrinolytic protease purified from
Fibrin(ogen)olytic enzymes have been purified from venoms of Chinese mamush (
The molecular weights of polypeptides from the fibrinolytic fractions obtained from
The molecular weight of brevilysine H5 from Chinese mamushi (
However, the protease purified in this study hydrolyzed gelatin, suggesting that it might destroy the connective tissue and induce hemorrhage, if it would be introduced into the body. Snake venom metalloproteases are enzymes with multiple domains whose main toxic effects are due to disruption of the hemostatic system8). They exert their hemorrhagic effects by degradation of proteins such as laminin, fibronectin, collagen type IV, and proteoglycansfrom the endothelial basal membrane20, 21) and by cleavage of fibrinogen, thereby making the protein unclottable and so enhancing the hemorrhagic state4). These effects are thought to be the major mechanism of hemorrhage.
Fibrin(ogen)olytic non-hemorrhagic snake venom metalloproteases, such as fibrolase from