Species of Tetraselmis in Prasinophyceae are well known as the basic food organisms in aquaculture (Kim and Hur 1998; Park and Hur 2000; Cabrera et al. 2005), as an important source for antioxidative substances in pharmacological studies (Laguna et al. 1993; Kim et al. 2002), and for their inportance in marine ecotoxicological testing (Park et al. 2005).
Since the first description of Tetraselmis by F. Stein in 1878 (Norris et al. 1980), many taxonomical studies on this green motile unicellular algae have been reported. The genus of Tetraselmis, first formalized according to the Christensen Criterion (Christensen 1962) has been identified by not only characteristics such as scale, flagellar hair, and basal bodies (Moestrup and Throndsen 1988; Marin and Melkonian 1994; Throndsen 1997), but also by types of pigments (Egeland et al. 1997; Latasa et al. 2004).
Although, by using light and electron microscope, morphological classification is possible at the species level, taxonomic decisions within the Tetraselmis are difficult to make because of the complex process of cellular characterization.
Genetic data based upon the amplification and sequencing of genes are also accumulated to use as a powerful tools for analysis of diverse microalgae. Thus, it is necessary to collect molecular data on Tetraselmis to determine whether they are identical strains or not. However, it is still difficult to find molecular studies on Tetraselmis.
In this study, 18S rDNA sequence of 41 strains of Tetraselmis was analysed to discriminate the genomic variations. The strains were constructed as a phylogenetic tree by comparing them with other known sequences from the National Center for Biotechnology Information (NCBI) database. These variations were also compared with the differences in the mean size of each strain of Tetraselmis.
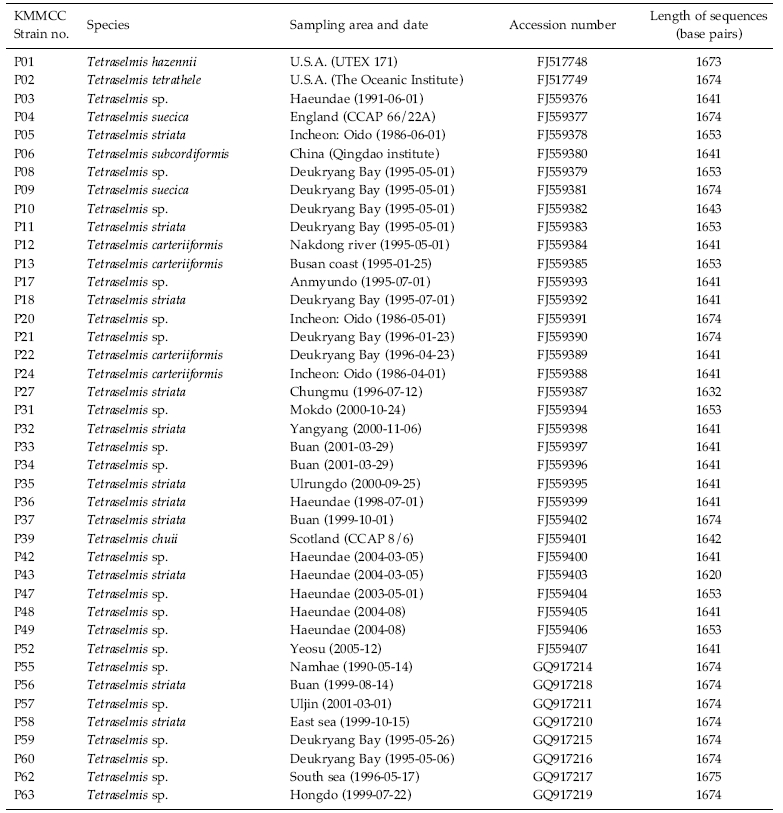
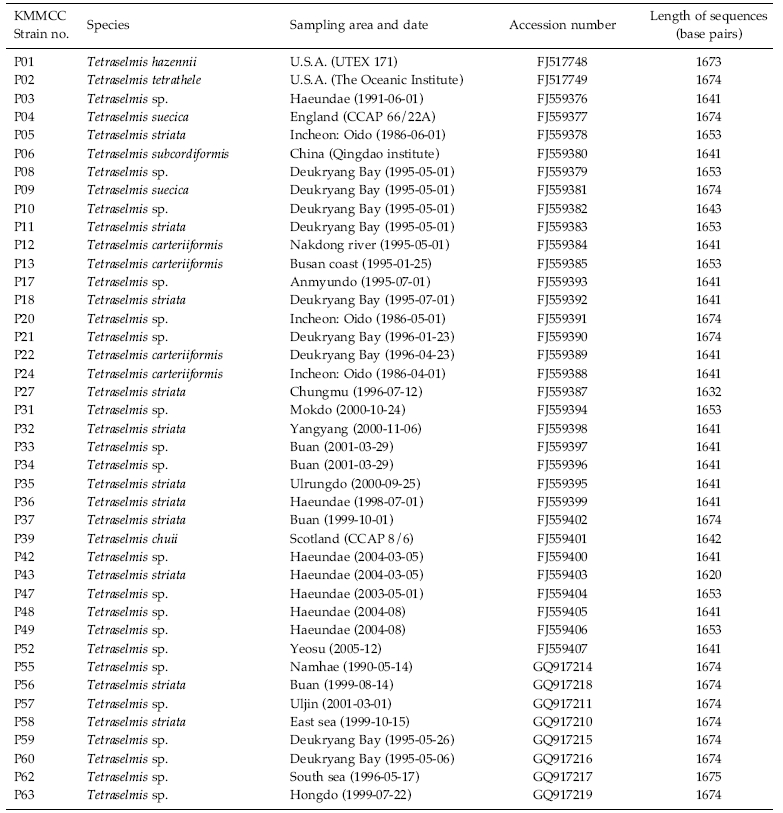
Forty-one strains of Tetraselmis were received from the Korea Marine Microalgae Culture Center (KMMCC) (Hur 2008). Information about the strains is given in Table 1. The strains were grown in 100 mL of f/2 culture medium (Guillard and Ryther 1962) at 22℃ with continuous light with 60 μmol photons m-2 s-1 for 10 days.
The Mean length of 30 specimens of the Tetraselmis strain was measured using a light microscope (×400). Significant differences in mean size of the strains were analysed using ANOVA and Ducan tests (SPSS software version 10.1; SPSS Inc., Chicago, IL, USA) at the level of 5%.
The LiCl method was used to extract the genomic DNA of the Tetraselmis (Hong et al. 1995). The quality and quantity of the extracted genomic DNA were measured by electrophoresis (MupidTM; Advance, Tokyo, Japan) and spectrometry (NanoDrop® ND-1000; NanoDrop Technologies, Wilmington, DE, USA), respectively.
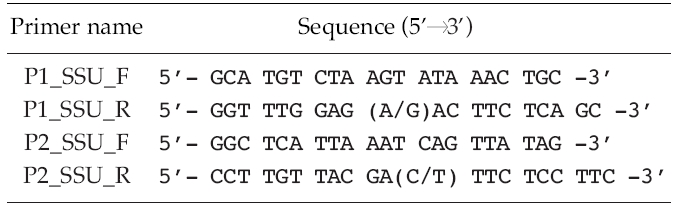
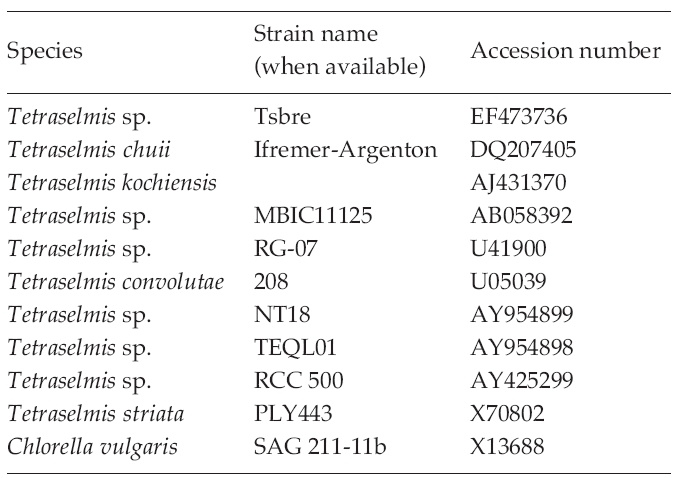
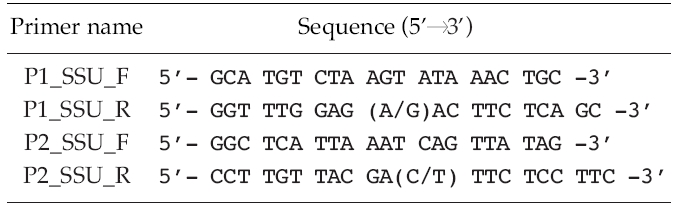
Polymerase chain reactions (PCR) (Mullis and Faloona 1987) performed using 10-100 ng of genomic DNA as a template and 0.5 μM of degenerated primers (Table 2) derived from a conserved region of 10 species of other known microalgae, listed in Table 3. Amplification conditions consisted of one cycle of denaturation at 95°C for 5 min, 30-35 cycles of denaturation at 95°C for 30 s, annealing at 50-55°C for 30 s and extension at 72°C for 1 min 45 s, followed by the final extension at 72°C for 7 min, and then stored at 4°C. About a 1.7 Kb size of PCR product was confirmed at 1% gel electrophoresis and extracted using the AccuPrep® Gel Purification Kit (Bioneer, Daejeon, Korea). Purified PCR products were ligated with the pGEM®-T Easy Vector Systems (Promega, San Luis Obisco, CA, USA) and then were transformed into the Eschrichia colil XL-1 blue. The recombinant plasmids were purified using the AccuPrep® Plasmid Extraction Kit (Bioneer, Korea). Selected clones were confirmed by colony PCR and EcoR restriction enzyme digestion (Fermentas, Burlington, Ontario, Canada) followed by DNA sequencing (Genotech, Daejeon, Korea).
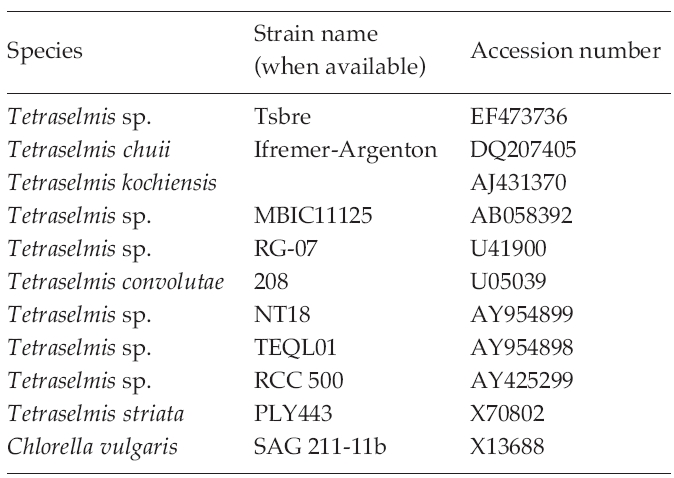
The identities of the acquired partial 18S rDNA sequence from Tetraselmis strains were confirmed by using the Blast N program (http://blast.ncbi.nlm.nih. gov) and were aligned with the other known microalgae 18S rDNA sequences using ClustalW2 (Thompson et al. 1994). The accession numbers of the microalgae sequence found in the NCBI GenBank are given in Table 3.
To confirm the phylogenetic relationships, a total of 1615 positions in the final dataset were subjected to neighbor-joining (NJ) and maximum-parsimony (MP) analysis using the MEGA v.4.0 (MEGA, Tempe, AZ, USA) (Tamura et al. 2007). The evolutionary distance method to construct the NJ tree was calculated with the Kimura 2-parameter (Kimura 1980). The rate variation among sites was modeled with a gamma distribution (0.25-parameter) and all positions containing gaps and missing data were eliminated from the dataset. The MP tree was obtained using the close-neighbor-interchange algorithm (Nei and Kumar 2000) in which the initial trees were obtained with the random additional 100 replications of sequences. All positions containing gaps and missing data were eliminated from the dataset. The reliability of both trees was tested by 2000 replication bootstraps. The 18S rDNA sequence of Chlorella vulgaris was used as the outgroup sequence.
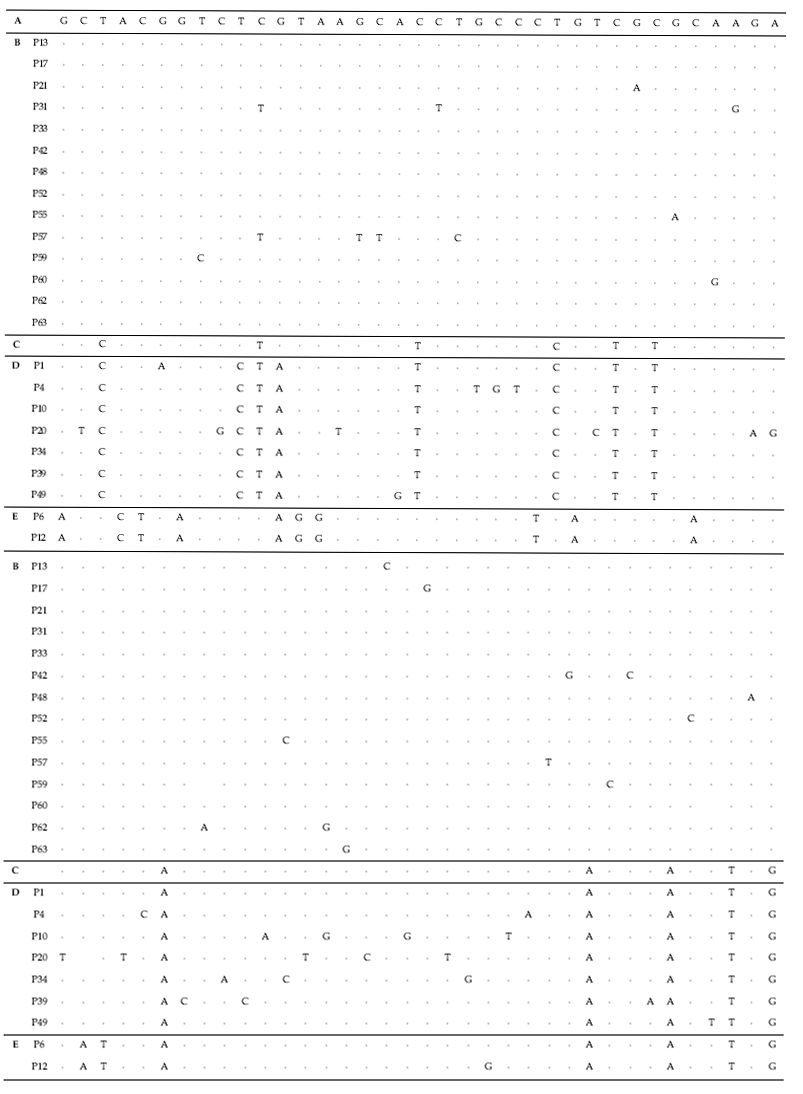
To test their generic similarity, the almost complete 18S rDNA sequences were determined for the 36 strains collected from Korean coastal water and 5 strains from foreign areas. Their accession numbers registered at GenBank are given in Table 1 and the lengths of sequence were varied on the sense and antisense primer used in gene amplification (Table 2). Forty-one strains showed high levels of identity (98-99%) to the correspoding regions of known 18S rDNA sequences such as Tetraselmis striata PLY 443 or Tetraselmis sp. Tsbre using the Blast N program. No differences were found between 13 strains (KMMCC P-5, 11, 18, 22, 24, 27, 32, 35, 36, 37, 43, 56 and 58: group A) and Tetraselmis striata PLY 443 or among 5 strains (KMMCC P-2, 3, 8, 9 and 47: group C) and Tetraselmis sp. Tsbre in their acquired rDNA sequences. Thirteen sequence positions were different between group A and C (Fig. 1). Four strains in group C were collected in May and June from the southern coast of Korea, and 13 strains in group A were gathered from diverse Korean coastal waters during all four seasons. In terms of the algal length, those of group A ranged from 8.75 μm to 13.75 μm were shorter than those of group C, which ranged from 12.5 μm to 17.5 μm.
The sequences of the other 35 strains were seperated into 3 groups; B, D and E. Of these, fourteen strains belonged to group B, which was very close to group A. Only 1-2 sequence positions differentiated between group A and B, excepting KMMCC P-57. Group D contained seven strains which were close to group C but the level of similarity between groups C and D was less so than between groups A and B. Different sequence positions of six strains of group D were 1-4 base pairs compared with group C, whereas KMMCC P-20 was the furtherest from it with 14 sequence differences. The different sequence position of KMMCC P-6 and P-12 in group E showed only one base pair. But group E differed slightly from both of group A and group C. It had 17 different sequence positions in KMMCC P-6 or 18 differences in KMMCC P-12 compare with group A. Eighteen or 19 base pairs differed between group C and KMMCC P-6 or P-12, respectively.
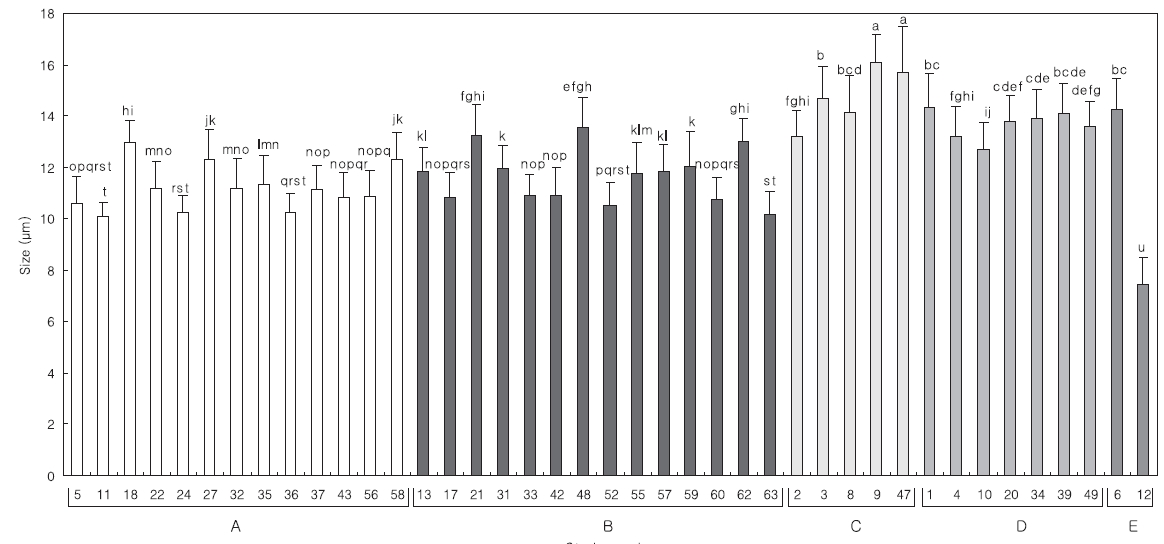
Comparing the algal size with the 18S rDNA sequence analysis in each group, the relationship exhibited more clearly in the algal length than in the width (data not shown). The distribution of mean length is divided into; groups A and B, and groups C and D. The length of the strains from group B was similar to that of group A, Tetraselmis striata, but KMMCC P-21, P-48 and P-62 were larger than the others (Fig. 2.). These three strains were closer to group C than group A. Although KMMCC P-6 and P-12 were considered as the same group E at the point of the gene sequence, their size was significantly different. Among the strains, KMMCC P-6 was the largest with 14.25 μm, and P-12 was the smallest with 7.42 μm. This result means that relationships between 18S rDNA sequences and the size of the strains do not always coincide.
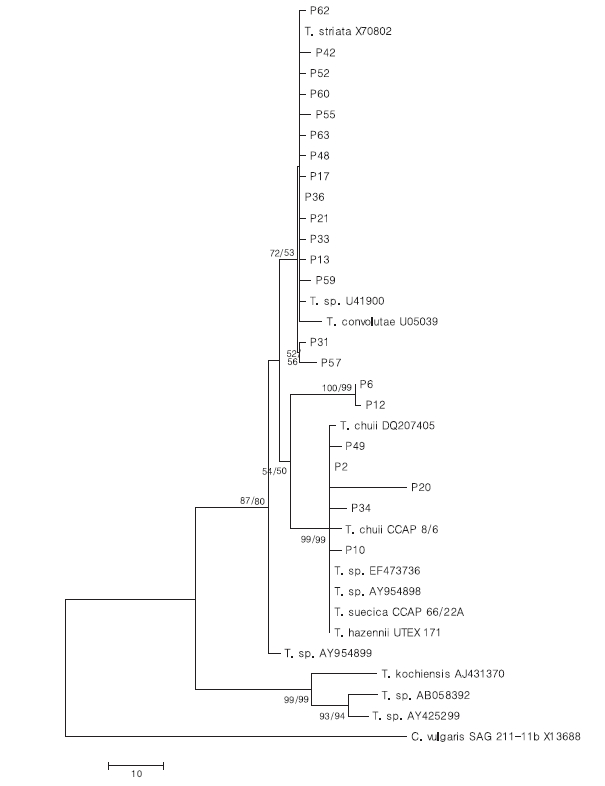
To analyze the relationship of 18S rDNA sequences among the strains of Tetraselmis, a phylogenetic tree was constructed using the NJ and MP methods with 2000 bootstrap values (Fig. 3). KMMCC P-36 and P-2 were selected as representative strains from identical sequences of groups A and C, respectively. Most strains were contained within two clades although separation was supported with less than 50%; 38% and 43% bootstrap values from NJ and MP, respectively. Groups A and B were clustered with Tetraselmis striata and T. convolutae excepting the branch order for KMMCC P-31 and P- 57 strains from the clade with a 72% bootstrap value in NJ tree. They had two and five different sequence positions respectively. Other clades were groups C and D, which were comprised of Tetraselmis chuii and Tetraselmis sp. Tsbre. Group E, comprised of KMMCC P-6 and P-12 was separated from clades of groups C and D with 54% and 50% bootstrap support in NJ and MP tree, respectively. This corresponded with the results on different sequence positions.
Tetraselmis currently contains about 30 species identified by morphological inspection from public microalgal culture collections (e.g., UTEX, CCMP, CCAP). However, only four species: Tetraselmis striata (Steinkotter et al. 1994), T. convolutae (de Jesus et al. 1995), T. kochiensis and T. chuii reported an 18S rDNA sequence. Other strains have not yet been identified as to the level of species.
One of the objectives of this research on their rDNA sequences was to identify strains of Tetraselmis among culture collections. Variation of life-history characteristics such as the flagellate phase, non-motile phase and cyst stage of Tetraselmis depends on environmental factors (Norris et al. 1980). In our experience, morphological discernment of Tetraselmis under a light microscope was often confused with the genera Colacium and Chloromonas. Therefore, analysis of the 18S rDNA sequence is evaluated as a simple tool for distinguishing the genus Tetraselmis. In this study, 13 strains were identical with Tetraselmis striata PLY 443 and 5 strains with Tetraselmis sp. Tsbre. This was more than 43% of the total strains examined. Grouping of Tetraselmis strains based on the result of the 18S rDNA sequence showed a partially corresponding tendency with the mean size variation of the strains. To understand the characterization and discrimination among various strains of Tetraselmis, further detailed molecular studies on the ITS regions or rbc L gene reflecting higher evolutionary rate, which was used in the genus Nannochloropsis (Suda et al. 2002) and DNA hybridization analysis or amplified fragment length polymorphism used in Chlorella vulgaris (Despres et al. 2003; Muller et al. 2005), are needed.